[English] 日本語
 Yorodumi
Yorodumi- EMDB-20546: Poliovirus (Type 1 Mahoney), receptor-catalysed 135S particle inc... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20546 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Poliovirus (Type 1 Mahoney), receptor-catalysed 135S particle incubated with anti-VP1 mAb at RT for 1 hr | |||||||||
 Map data Map data | Type-1 Mahoney Poliovirus, pre-expanded with PVR and incubated with anti-VP1 monoclonal Ab at RT for 1hr | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationsymbiont-mediated suppression of host translation initiation / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane ...symbiont-mediated suppression of host translation initiation / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of RIG-I activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MAVS activity / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / host cell cytoplasmic vesicle membrane / ribonucleoside triphosphate phosphatase activity / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / RNA helicase activity / endocytosis involved in viral entry into host cell / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / proteolysis / RNA binding / zinc ion binding / ATP binding / membrane Similarity search - Function | |||||||||
| Biological species |   Poliovirus type 1 (strain Mahoney) / Poliovirus type 1 (strain Mahoney) /  | |||||||||
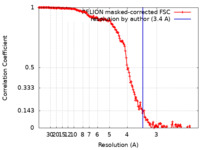
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Hogle JM / Filman DJ / Shah PNM | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: PLoS Pathog / Year: 2020 Journal: PLoS Pathog / Year: 2020Title: Cryo-EM structures reveal two distinct conformational states in a picornavirus cell entry intermediate. Authors: Pranav N M Shah / David J Filman / Krishanthi S Karunatilaka / Emma L Hesketh / Elisabetta Groppelli / Mike Strauss / James M Hogle /    Abstract: The virions of enteroviruses such as poliovirus undergo a global conformational change after binding to the cellular receptor, characterized by a 4% expansion, and by the opening of holes at the two ...The virions of enteroviruses such as poliovirus undergo a global conformational change after binding to the cellular receptor, characterized by a 4% expansion, and by the opening of holes at the two and quasi-three-fold symmetry axes of the capsid. The resultant particle is called a 135S particle or A-particle and is thought to be on the pathway to a productive infection. Previously published studies have concluded that the membrane-interactive peptides, namely VP4 and the N-terminus of VP1, are irreversibly externalized in the 135S particle. However, using established protocols to produce the 135S particle, and single particle cryo-electron microscopy methods, we have identified at least two unique states that we call the early and late 135S particle. Surprisingly, only in the "late" 135S particles have detectable levels of the VP1 N-terminus been trapped outside the capsid. Moreover, we observe a distinct density inside the capsid that can be accounted for by VP4 that remains associated with the genome. Taken together our results conclusively demonstrate that the 135S particle is not a unique conformation, but rather a family of conformations that could exist simultaneously. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20546.map.gz emd_20546.map.gz | 253.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20546-v30.xml emd-20546-v30.xml emd-20546.xml emd-20546.xml | 22.8 KB 22.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_20546_fsc.xml emd_20546_fsc.xml | 15.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_20546.png emd_20546.png | 166.6 KB | ||
| Masks |  emd_20546_msk_1.map emd_20546_msk_1.map | 343 MB |  Mask map Mask map | |
| Others |  emd_20546_half_map_1.map.gz emd_20546_half_map_1.map.gz emd_20546_half_map_2.map.gz emd_20546_half_map_2.map.gz | 273.4 MB 273.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20546 http://ftp.pdbj.org/pub/emdb/structures/EMD-20546 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20546 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20546 | HTTPS FTP |
-Related structure data
| Related structure data |  6q0bMC  6p9oC  6p9wC  6pszC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20546.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20546.map.gz / Format: CCP4 / Size: 343 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Type-1 Mahoney Poliovirus, pre-expanded with PVR and incubated with anti-VP1 monoclonal Ab at RT for 1hr | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.096 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
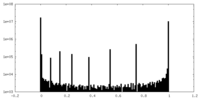
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_20546_msk_1.map emd_20546_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
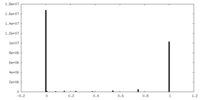
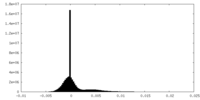
| Density Histograms |
-Half map: Type-1 Mahoney Poliovirus, pre-expanded with PVR and incubated...
| File | emd_20546_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Type-1 Mahoney Poliovirus, pre-expanded with PVR and incubated with anti-VP1 monoclonal Ab at RT for 1hr - half 1 | ||||||||||||
| Projections & Slices |
| ||||||||||||
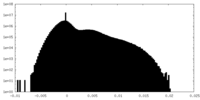
| Density Histograms |
-Half map: Type-1 Mahoney Poliovirus, pre-expanded with PVR and incubated...
| File | emd_20546_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Type-1 Mahoney Poliovirus, pre-expanded with PVR and incubated with anti-VP1 monoclonal Ab at RT for 1hr - half 2 | ||||||||||||
| Projections & Slices |
| ||||||||||||
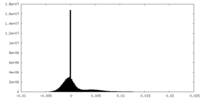
| Density Histograms |
- Sample components
Sample components
-Entire : Poliovirus type 1 (strain Mahoney)
| Entire | Name:   Poliovirus type 1 (strain Mahoney) Poliovirus type 1 (strain Mahoney) |
|---|---|
| Components |
|
-Supramolecule #1: Poliovirus type 1 (strain Mahoney)
| Supramolecule | Name: Poliovirus type 1 (strain Mahoney) / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 / NCBI-ID: 12081 / Sci species name: Poliovirus type 1 (strain Mahoney) / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 10.0 MDa |
| Virus shell | Shell ID: 1 / Name: Capsid / Diameter: 300.0 Å / T number (triangulation number): 1 |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Poliovirus type 1 (strain Mahoney) / Strain: Mahoney Poliovirus type 1 (strain Mahoney) / Strain: Mahoney |
| Molecular weight | Theoretical: 33.488613 KDa |
| Sequence | String: GLGQMLESMI DNTVRETVGA ATSRDALPNT EASGPTHSKE IPALTAVETG ATNPLVPSDT VQTRHVVQHR SRSESSIESF FARGACVTI MTVDNPASTT NKDKLFAVWK ITYKDTVQLR RKLEFFTYSR FDMELTFVVT ANFTETNNGH ALNQVYQIMY V PPGAPVPE ...String: GLGQMLESMI DNTVRETVGA ATSRDALPNT EASGPTHSKE IPALTAVETG ATNPLVPSDT VQTRHVVQHR SRSESSIESF FARGACVTI MTVDNPASTT NKDKLFAVWK ITYKDTVQLR RKLEFFTYSR FDMELTFVVT ANFTETNNGH ALNQVYQIMY V PPGAPVPE KWDDYTWQTS SNPSIFYTYG TAPARISVPY VGISNAYSHF YDGFSKVPLK DQSAALGDSL YGAASLNDFG IL AVRVVND HNPTKVTSKI RVYLKPKHIR VWCPRPPRAV AYYGPGVDYK DGTLTPLSTK DLTTY |
-Macromolecule #2: Capsid protein VP2
| Macromolecule | Name: Capsid protein VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Poliovirus type 1 (strain Mahoney) / Strain: Mahoney Poliovirus type 1 (strain Mahoney) / Strain: Mahoney |
| Molecular weight | Theoretical: 30.075783 KDa |
| Sequence | String: SPNIEACGYS DRVLQLTLGN STITTQEAAN SVVAYGRWPE YLRDSEANPV DQPTEPDVAA CRFYTLDTVS WTKESRGWWW KLPDALRDM GLFGQNMYYH YLGRSGYTVH VQCNASKFHQ GALGVFAVPE MCLAGDSNTT TMHTSYQNAN PGEKGGTFTG T FTPDNNQT ...String: SPNIEACGYS DRVLQLTLGN STITTQEAAN SVVAYGRWPE YLRDSEANPV DQPTEPDVAA CRFYTLDTVS WTKESRGWWW KLPDALRDM GLFGQNMYYH YLGRSGYTVH VQCNASKFHQ GALGVFAVPE MCLAGDSNTT TMHTSYQNAN PGEKGGTFTG T FTPDNNQT SPARRFCPVD YLLGNGTLLG NAFVFPHQII NLRTNNCATL VLPYVNSLSI DSMVKHNNWG IAILPLAPLN FA SESSPEI PITLTIAPMC CEFNGLRNIT LPRLQ |
-Macromolecule #3: Capsid protein VP3
| Macromolecule | Name: Capsid protein VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Poliovirus type 1 (strain Mahoney) / Strain: Mahoney Poliovirus type 1 (strain Mahoney) / Strain: Mahoney |
| Molecular weight | Theoretical: 26.547482 KDa |
| Sequence | String: GLPVMNTPGS NQYLTADNFQ SPCALPEFDV TPPIDIPGEV KNMMELAEID TMIPFDLSAT KKNTMEMYRV RLSDKPHTDD PILCLSLSP ASDPRLSHTM LGEILNYYTH WAGSLKFTFL FCGSMMATGK LLVSYAPPGA DPPKKRKEAM LGTHVIWDIG L QSSCTMVV ...String: GLPVMNTPGS NQYLTADNFQ SPCALPEFDV TPPIDIPGEV KNMMELAEID TMIPFDLSAT KKNTMEMYRV RLSDKPHTDD PILCLSLSP ASDPRLSHTM LGEILNYYTH WAGSLKFTFL FCGSMMATGK LLVSYAPPGA DPPKKRKEAM LGTHVIWDIG L QSSCTMVV PWISNTTYRQ TIDDSFTEGG YISVFYQTRI VVPLSTPREM DILGFVSACN DFSVRLLRDT THIEQKALAQ |
-Macromolecule #4: Capsid protein VP4
| Macromolecule | Name: Capsid protein VP4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Poliovirus type 1 (strain Mahoney) / Strain: Mahoney Poliovirus type 1 (strain Mahoney) / Strain: Mahoney |
| Molecular weight | Theoretical: 7.690482 KDa |
| Sequence | String: (MYR)GAQVSSQKV GAHENSNRAY GGSTINYTTI NYYRDSASNA ASKQDFSQDP SKFTEPIKDV LIKTAPMLNS |
-Macromolecule #5: anti-VP1 mAb
| Macromolecule | Name: anti-VP1 mAb / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 44.452984 KDa |
| Sequence | String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)C(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) ...String: (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)C(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)C(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)C(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) C(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)C(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)C(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)C (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) C(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) C(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)(UNK)(UNK) (UNK)(UNK)(UNK)(UNK)(UNK)(UNK) (UNK)C |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.4 mg/mL |
|---|---|
| Buffer | pH: 7.5 / Details: 20 mM Tris-HCl, pH 7.5, 2 mM CaCl2 |
| Grid | Support film - Material: CARBON / Support film - topology: LACEY / Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
| Details | This sample was monodisperse. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Detector mode: COUNTING / Number grids imaged: 1 / Average electron dose: 1.06 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: SPOT SCAN / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT |
|---|---|
| Output model |  PDB-6q0b: |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)