+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 |  | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Structure of the human CCANdeltaT CENP-A alpha-satellite complex | ||||||||||||
 マップデータ マップデータ | Local resolution filtered map from Relion | ||||||||||||
 試料 試料 |
| ||||||||||||
 キーワード キーワード | Chromosome / kinetochore / cell division / centromere / CELL CYCLE | ||||||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報Mis6-Sim4 complex / positive regulation of protein localization to kinetochore / centromere complex assembly / kinetochore organization / spindle attachment to meiosis I kinetochore / metaphase chromosome alignment / kinetochore binding / sex differentiation / CENP-A containing chromatin assembly / centromeric DNA binding ...Mis6-Sim4 complex / positive regulation of protein localization to kinetochore / centromere complex assembly / kinetochore organization / spindle attachment to meiosis I kinetochore / metaphase chromosome alignment / kinetochore binding / sex differentiation / CENP-A containing chromatin assembly / centromeric DNA binding / chordate embryonic development / negative regulation of epithelial cell apoptotic process / kinetochore assembly / protein localization to chromosome, centromeric region / attachment of mitotic spindle microtubules to kinetochore / condensed chromosome, centromeric region / inner kinetochore / establishment of mitotic spindle orientation / mitotic cytokinesis / mitotic sister chromatid segregation / chromosome, centromeric region / negative regulation of megakaryocyte differentiation / protein localization to CENP-A containing chromatin / pericentric heterochromatin / Replacement of protamines by nucleosomes in the male pronucleus / CENP-A containing nucleosome / Packaging Of Telomere Ends / Recognition and association of DNA glycosylase with site containing an affected purine / Cleavage of the damaged purine / Amplification of signal from unattached kinetochores via a MAD2 inhibitory signal / Recognition and association of DNA glycosylase with site containing an affected pyrimidine / Cleavage of the damaged pyrimidine / Deposition of new CENPA-containing nucleosomes at the centromere / Mitotic Prometaphase / EML4 and NUDC in mitotic spindle formation / telomere organization / Inhibition of DNA recombination at telomere / Meiotic synapsis / RNA Polymerase I Promoter Opening / Assembly of the ORC complex at the origin of replication / Resolution of Sister Chromatid Cohesion / Regulation of endogenous retroelements by the Human Silencing Hub (HUSH) complex / innate immune response in mucosa / NRIF signals cell death from the nucleus / SUMOylation of chromatin organization proteins / DNA methylation / Condensation of Prophase Chromosomes / Chromatin modifications during the maternal to zygotic transition (MZT) / HCMV Late Events / SIRT1 negatively regulates rRNA expression / positive regulation of epithelial cell proliferation / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / PRC2 methylates histones and DNA / mitotic spindle organization / Regulation of endogenous retroelements by KRAB-ZFP proteins / Defective pyroptosis / Regulation of endogenous retroelements by Piwi-interacting RNAs (piRNAs) / HDACs deacetylate histones / chromosome segregation / Nonhomologous End-Joining (NHEJ) / RNA Polymerase I Promoter Escape / Transcriptional regulation by small RNAs / RHO GTPases Activate Formins / Formation of the beta-catenin:TCF transactivating complex / RUNX1 regulates genes involved in megakaryocyte differentiation and platelet function / Activated PKN1 stimulates transcription of AR (androgen receptor) regulated genes KLK2 and KLK3 / G2/M DNA damage checkpoint / HDMs demethylate histones / NoRC negatively regulates rRNA expression / DNA Damage/Telomere Stress Induced Senescence / B-WICH complex positively regulates rRNA expression / kinetochore / PKMTs methylate histone lysines / Meiotic recombination / Pre-NOTCH Transcription and Translation / centriolar satellite / Metalloprotease DUBs / RMTs methylate histone arginines / Activation of anterior HOX genes in hindbrain development during early embryogenesis / Transcriptional regulation of granulopoiesis / HCMV Early Events / antimicrobial humoral immune response mediated by antimicrobial peptide / structural constituent of chromatin / Separation of Sister Chromatids / antibacterial humoral response / UCH proteinases / nucleosome / heterochromatin formation / E3 ubiquitin ligases ubiquitinate target proteins / nucleosome assembly / mitotic cell cycle / actin cytoskeleton / Recruitment and ATM-mediated phosphorylation of repair and signaling proteins at DNA double strand breaks / chromatin organization / HATs acetylate histones / RUNX1 regulates transcription of genes involved in differentiation of HSCs / chromosome / MLL4 and MLL3 complexes regulate expression of PPARG target genes in adipogenesis and hepatic steatosis / Processing of DNA double-strand break ends / midbody 類似検索 - 分子機能 | ||||||||||||
| 生物種 |  Homo sapiens (ヒト) Homo sapiens (ヒト) | ||||||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 8.9 Å | ||||||||||||
 データ登録者 データ登録者 | Yatskevich S / Muir KW / Bellini D / Zhang Z / Yang J / Tischer T / Predin M / Dendooven T / McLaughlin SH / Barford D | ||||||||||||
| 資金援助 |  英国, 英国,  ドイツ, 3件 ドイツ, 3件
| ||||||||||||
 引用 引用 |  ジャーナル: Science / 年: 2022 ジャーナル: Science / 年: 2022タイトル: Structure of the human inner kinetochore bound to a centromeric CENP-A nucleosome. 著者: Stanislau Yatskevich / Kyle W Muir / Dom Bellini / Ziguo Zhang / Jing Yang / Thomas Tischer / Masa Predin / Tom Dendooven / Stephen H McLaughlin / David Barford /  要旨: Kinetochores assemble onto specialized centromeric CENP-A (centromere protein A) nucleosomes (CENP-A) to mediate attachments between chromosomes and the mitotic spindle. We describe cryo-electron ...Kinetochores assemble onto specialized centromeric CENP-A (centromere protein A) nucleosomes (CENP-A) to mediate attachments between chromosomes and the mitotic spindle. We describe cryo-electron microscopy structures of the human inner kinetochore constitutive centromere associated network (CCAN) complex bound to CENP-A reconstituted onto α-satellite DNA. CCAN forms edge-on contacts with CENP-A, and a linker DNA segment of the α-satellite repeat emerges from the fully wrapped end of the nucleosome to thread through the central CENP-LN channel that tightly grips the DNA. The CENP-TWSX histone-fold module further augments DNA binding and partially wraps the linker DNA in a manner reminiscent of canonical nucleosomes. Our study suggests that the topological entrapment of the linker DNA by CCAN provides a robust mechanism by which kinetochores withstand both pushing and pulling forces exerted by the mitotic spindle. | ||||||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| 添付画像 |
|---|
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_14375.map.gz emd_14375.map.gz | 15.5 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-14375-v30.xml emd-14375-v30.xml emd-14375.xml emd-14375.xml | 32.2 KB 32.2 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_14375_fsc.xml emd_14375_fsc.xml | 7.2 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_14375.png emd_14375.png | 59.2 KB | ||
| Filedesc metadata |  emd-14375.cif.gz emd-14375.cif.gz | 9.5 KB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-14375 http://ftp.pdbj.org/pub/emdb/structures/EMD-14375 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14375 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-14375 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7yyhMC  7pb4C  7pb8C  7piiC  7pknC 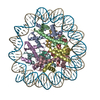 7r5rC  7r5sC  7r5vC  7ywxC M: このマップから作成された原子モデル C: 同じ文献を引用 ( |
|---|---|
| 類似構造データ | 類似検索 - 機能・相同性  F&H 検索 F&H 検索 |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_14375.map.gz / 形式: CCP4 / 大きさ: 25.3 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_14375.map.gz / 形式: CCP4 / 大きさ: 25.3 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Local resolution filtered map from Relion | ||||||||||||||||||||||||||||||||||||
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.662 Å | ||||||||||||||||||||||||||||||||||||
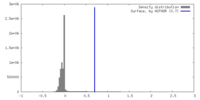
| 密度 |
| ||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
|
-添付データ
- 試料の構成要素
試料の構成要素
+全体 : CCAN-CENP-A inner centromere complex
+超分子 #1: CCAN-CENP-A inner centromere complex
+分子 #1: Histone H3-like centromeric protein A
+分子 #2: Histone H4
+分子 #3: Histone H2A type 1-C
+分子 #4: Histone H2B type 1-C/E/F/G/I
+分子 #5: Centromere protein H
+分子 #6: Centromere protein I
+分子 #8: Centromere protein K
+分子 #9: Centromere protein L
+分子 #10: Centromere protein M
+分子 #11: Centromere protein N
+分子 #12: Centromere protein O
+分子 #13: Centromere protein P
+分子 #14: Centromere protein Q
+分子 #15: Centromere protein R
+分子 #16: Centromere protein U
+分子 #17: Centromere protein C
+分子 #7: DNA (171-MER)
+分子 #18: DNA (171-MER)
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.8 |
|---|---|
| 凍結 | 凍結剤: ETHANE |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 平均電子線量: 50.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / 最大 デフォーカス(公称値): 2.6 µm / 最小 デフォーカス(公称値): 1.2 µm |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)




















 Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ)
