[English] 日本語
 Yorodumi
Yorodumi- EMDB-12262: Lateral-open conformation of the lid-locked BAM complex (BamA E43... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12262 | ||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Lateral-open conformation of the lid-locked BAM complex (BamA E435C S665C, BamBDCE) by cryoEM | ||||||||||||||||||||||||
 Map data Map data | Postprocessed masked map of the lateral-open conformation of a lid-locked BAM complex | ||||||||||||||||||||||||
 Sample Sample |
| ||||||||||||||||||||||||
 Keywords Keywords | Outer membrane protein assembly / beta-barrel / Gram negative bacteria / protein foldase / MEMBRANE PROTEIN | ||||||||||||||||||||||||
| Function / homology |  Function and homology information Function and homology informationBam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / protein insertion into membrane / Secretion of toxins / cell outer membrane / protein-folding chaperone binding / protein-macromolecule adaptor activity / cell adhesion / response to antibiotic / cell surface ...Bam protein complex / Gram-negative-bacterium-type cell outer membrane assembly / protein insertion into membrane / Secretion of toxins / cell outer membrane / protein-folding chaperone binding / protein-macromolecule adaptor activity / cell adhesion / response to antibiotic / cell surface / membrane / identical protein binding Similarity search - Function | ||||||||||||||||||||||||
| Biological species |   | ||||||||||||||||||||||||
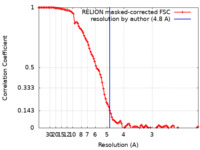
| Method | single particle reconstruction / cryo EM / Resolution: 4.8 Å | ||||||||||||||||||||||||
 Authors Authors | Haysom SF | ||||||||||||||||||||||||
| Funding support |  United Kingdom, United Kingdom,  Belgium, 7 items Belgium, 7 items
| ||||||||||||||||||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: The role of membrane destabilisation and protein dynamics in BAM catalysed OMP folding. Authors: Paul White / Samuel F Haysom / Matthew G Iadanza / Anna J Higgins / Jonathan M Machin / James M Whitehouse / Jim E Horne / Bob Schiffrin / Charlotte Carpenter-Platt / Antonio N Calabrese / ...Authors: Paul White / Samuel F Haysom / Matthew G Iadanza / Anna J Higgins / Jonathan M Machin / James M Whitehouse / Jim E Horne / Bob Schiffrin / Charlotte Carpenter-Platt / Antonio N Calabrese / Kelly M Storek / Steven T Rutherford / David J Brockwell / Neil A Ranson / Sheena E Radford /   Abstract: The folding of β-barrel outer membrane proteins (OMPs) in Gram-negative bacteria is catalysed by the β-barrel assembly machinery (BAM). How lateral opening in the β-barrel of the major subunit ...The folding of β-barrel outer membrane proteins (OMPs) in Gram-negative bacteria is catalysed by the β-barrel assembly machinery (BAM). How lateral opening in the β-barrel of the major subunit BamA assists in OMP folding, and the contribution of membrane disruption to BAM catalysis remain unresolved. Here, we use an anti-BamA monoclonal antibody fragment (Fab1) and two disulphide-crosslinked BAM variants (lid-locked (LL), and POTRA-5-locked (P5L)) to dissect these roles. Despite being lethal in vivo, we show that all complexes catalyse folding in vitro, albeit less efficiently than wild-type BAM. CryoEM reveals that while Fab1 and BAM-P5L trap an open-barrel state, BAM-LL contains a mixture of closed and contorted, partially-open structures. Finally, all three complexes globally destabilise the lipid bilayer, while BamA does not, revealing that the BAM lipoproteins are required for this function. Together the results provide insights into the role of BAM structure and lipid dynamics in OMP folding. | ||||||||||||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12262.map.gz emd_12262.map.gz | 7.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12262-v30.xml emd-12262-v30.xml emd-12262.xml emd-12262.xml | 30 KB 30 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12262_fsc.xml emd_12262_fsc.xml | 10.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_12262.png emd_12262.png | 106 KB | ||
| Masks |  emd_12262_msk_1.map emd_12262_msk_1.map | 103 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-12262.cif.gz emd-12262.cif.gz | 8.4 KB | ||
| Others |  emd_12262_half_map_1.map.gz emd_12262_half_map_1.map.gz emd_12262_half_map_2.map.gz emd_12262_half_map_2.map.gz | 95.7 MB 95.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12262 http://ftp.pdbj.org/pub/emdb/structures/EMD-12262 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12262 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12262 | HTTPS FTP |
-Related structure data
| Related structure data |  7nbxMC  7bm5C  7bnqC  7ncsC  7nd0C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_12262.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12262.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Postprocessed masked map of the lateral-open conformation of a lid-locked BAM complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12262_msk_1.map emd_12262_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 2 from non-uniform refinement (cryoSPARC 2.2.0) of...
| File | emd_12262_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 2 from non-uniform refinement (cryoSPARC 2.2.0) of the polished particle stack | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Halfmap 1 from non-uniform refinement (cryoSPARC 2.2.0) of...
| File | emd_12262_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Halfmap 1 from non-uniform refinement (cryoSPARC 2.2.0) of the polished particle stack | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : beta-barrel assembly machinery (BAM) complex (BamABCDE)
| Entire | Name: beta-barrel assembly machinery (BAM) complex (BamABCDE) |
|---|---|
| Components |
|
-Supramolecule #1: beta-barrel assembly machinery (BAM) complex (BamABCDE)
| Supramolecule | Name: beta-barrel assembly machinery (BAM) complex (BamABCDE) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 203 KDa |
-Macromolecule #1: Outer membrane protein assembly factor BamA
| Macromolecule | Name: Outer membrane protein assembly factor BamA / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 90.601352 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAMKKLLIAS LLFSSATVYG AEGFVVKDIH FEGLQRVAVG AALLSMPVRT GDTVNDEDIS NTIRALFATG NFEDVRVLRD GDTLLVQVK ERPTIASITF SGNKSVKDDM LKQNLEASGV RVGESLDRTT IADIEKGLED FYYSVGKYSA SVKAVVTPLP R NRVDLKLV ...String: MAMKKLLIAS LLFSSATVYG AEGFVVKDIH FEGLQRVAVG AALLSMPVRT GDTVNDEDIS NTIRALFATG NFEDVRVLRD GDTLLVQVK ERPTIASITF SGNKSVKDDM LKQNLEASGV RVGESLDRTT IADIEKGLED FYYSVGKYSA SVKAVVTPLP R NRVDLKLV FQEGVSAEIQ QINIVGNHAF TTDELISHFQ LRDEVPWWNV VGDRKYQKQK LAGDLETLRS YYLDRGYARF NI DSTQVSL TPDKKGIYVT VNITEGDQYK LSGVEVSGNL AGHSAEIEQL TKIEPGELYN GTKVTKMEDD IKKLLGRYGY AYP RVQSMP EINDADKTVK LRVNVDAGNR FYVRKIRFEG NDTSKDAVLR REMRQMEGAW LGSDLVDQGK ERLNRLGFFE TVDT DTQRV PGSPDQVDVV YKVKERNTGS FNFGIGYGTC SGVSFQAGVQ QDNWLGTGYA VGINGTKNDY QTYAELSVTN PYFTV DGVS LGGRLFYNDF QADDADLSDY TNKSYGTDVT LGFPINEYNS LRAGLGYVHN SLSNMQPQVA MWRYLYSMGE HPSTSD QDN SFKTDDFTFN YGWTYNKLDR GYFPTDGSRV NLTGKVTIPG SDNEYYKVTL DTATYVPIDD DHKWVVLGRT RWGYGDG LG GKEMPFYENF YAGGSSTVRG FQCNTIGPKA VYFPHQASNY DPDYDYESAT QDGAKDLSKS DDAVGGNAMA VASLEFIT P TPFISDKYAN SVRTSFFWDM GTVWDTNWDS SQYSGYPDYS DPSNIRMSAG IALQWMSPLG PLVFSYAQPF KKYDGDKAE QFQFNIGKTW UniProtKB: Outer membrane protein assembly factor BamA |
-Macromolecule #2: Outer membrane protein assembly factor BamB
| Macromolecule | Name: Outer membrane protein assembly factor BamB / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 41.918945 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MQLRKLLLPG LLSVTLLSGC SLFNSEEDVV KMSPLPTVEN QFTPTTAWST SVGSGIGNFY SNLHPALADN VVYAADRAGL VKALNADDG KEIWSVSLAE KDGWFSKEPA LLSGGVTVSG GHVYIGSEKA QVYALNTSDG TVAWQTKVAG EALSRPVVSD G LVLIHTSN ...String: MQLRKLLLPG LLSVTLLSGC SLFNSEEDVV KMSPLPTVEN QFTPTTAWST SVGSGIGNFY SNLHPALADN VVYAADRAGL VKALNADDG KEIWSVSLAE KDGWFSKEPA LLSGGVTVSG GHVYIGSEKA QVYALNTSDG TVAWQTKVAG EALSRPVVSD G LVLIHTSN GQLQALNEAD GAVKWTVNLD MPSLSLRGES APTTAFGAAV VGGDNGRVSA VLMEQGQMIW QQRISQATGS TE IDRLSDV DTTPVVVNGV VFALAYNGNL TALDLRSGQI MWKRELGSVN DFIVDGNRIY LVDQNDRVMA LTIDGGVTLW TQS DLLHRL LTSPVLYNGN LVVGDSEGYL HWINVEDGRF VAQQKVDSSG FQTEPVAADG KLLIQAKDGT VYSITR UniProtKB: Outer membrane protein assembly factor BamB |
-Macromolecule #3: Outer membrane protein assembly factor BamC
| Macromolecule | Name: Outer membrane protein assembly factor BamC / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 36.875277 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MAYSVQKSRL AKVAGVSLVL LLAACSSDSR YKRQVSGDEA YLEAAPLAEL HAPAGMILPV TSGDYAIPVT NGSGAVGKAL DIRPPAQPL ALVSGARTQF TGDTASLLVE NGRGNTLWPQ VVSVLQAKNY TITQRDDAGQ TLTTDWVQWN RLDEDEQYRG R YQISVKPQ ...String: MAYSVQKSRL AKVAGVSLVL LLAACSSDSR YKRQVSGDEA YLEAAPLAEL HAPAGMILPV TSGDYAIPVT NGSGAVGKAL DIRPPAQPL ALVSGARTQF TGDTASLLVE NGRGNTLWPQ VVSVLQAKNY TITQRDDAGQ TLTTDWVQWN RLDEDEQYRG R YQISVKPQ GYQQAVTVKL LNLEQAGKPV ADAASMQRYS TEMMNVISAG LDKSATDAAN AAQNRASTTM DVQSAADDTG LP MLVVRGP FNVVWQRLPA ALEKVGMKVT DSTRSQGNMA VTYKPLSDSD WQELGASDPG LASGDYKLQV GDLDNRSSLQ FID PKGHTL TQSQNDALVA VFQAAFSK UniProtKB: Outer membrane protein assembly factor BamC |
-Macromolecule #4: Outer membrane protein assembly factor BamD
| Macromolecule | Name: Outer membrane protein assembly factor BamD / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 27.85835 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MTRMKYLVAA ATLSLFLAGC SGSKEEVPDN PPNEIYATAQ QKLQDGNWRQ AITQLEALDN RYPFGPYSQQ VQLDLIYAYY KNADLPLAQ AAIDRFIRLN PTHPNIDYVM YMRGLTNMAL DDSALQGFFG VDRSDRDPQH ARAAFSDFSK LVRGYPNSQY T TDATKRLV ...String: MTRMKYLVAA ATLSLFLAGC SGSKEEVPDN PPNEIYATAQ QKLQDGNWRQ AITQLEALDN RYPFGPYSQQ VQLDLIYAYY KNADLPLAQ AAIDRFIRLN PTHPNIDYVM YMRGLTNMAL DDSALQGFFG VDRSDRDPQH ARAAFSDFSK LVRGYPNSQY T TDATKRLV FLKDRLAKYE YSVAEYYTER GAWVAVVNRV EGMLRDYPDT QATRDALPLM ENAYRQMQMN AQAEKVAKII AA NSSNT UniProtKB: Outer membrane protein assembly factor BamD |
-Macromolecule #5: Outer membrane protein assembly factor BamE
| Macromolecule | Name: Outer membrane protein assembly factor BamE / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 13.530256 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MRCKTLTAAA AVLLMLTAGC STLERVVYRP DINQGNYLTA NDVSKIRVGM TQQQVAYALG TPLMSDPFGT NTWFYVFRQQ PGHEGVTQQ TLTLTFNSSG VLTNIDNKPA LSGNGGHHHH HHHH UniProtKB: Outer membrane protein assembly factor BamE |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 3.3 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Pretreatment - Type: GLOW DISCHARGE Details: Grids were glow discharged in a GlowQube Plus for 30s at 60 mA | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277.15 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Average electron dose: 49.7 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model |
| ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Details | Initial local fitting accomplished in Chimera. VMD was then used to setup cMDFF simulations that were run using NAMD. Both 7BNQ and 5LJO were used as initial models, with the best fitting parts of each then combined and flexibly fit again to generate the final conformation. This was then Real Space Refined in Phenix to generate the final structure | ||||||||||||||||||
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Overall B value: 211 | ||||||||||||||||||
| Output model |  PDB-7nbx: |
 Movie
Movie Controller
Controller




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)