[English] 日本語
 Yorodumi
Yorodumi- EMDB-11999: M. pneumoniae 70S ribosome in complex with chloramphenicol obtain... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11999 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | M. pneumoniae 70S ribosome in complex with chloramphenicol obtained from in situ data using M, focused refinement of 50S sub-unit | |||||||||||||||
 Map data Map data | ||||||||||||||||
 Sample Sample |
| |||||||||||||||
| Function / homology |  Function and homology information Function and homology informationlarge ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding ...large ribosomal subunit / transferase activity / 5S rRNA binding / ribosomal large subunit assembly / large ribosomal subunit rRNA binding / cytosolic large ribosomal subunit / cytoplasmic translation / tRNA binding / negative regulation of translation / rRNA binding / structural constituent of ribosome / ribosome / translation / ribonucleoprotein complex / response to antibiotic / mRNA binding / cytoplasm Similarity search - Function | |||||||||||||||
| Biological species |  Mycoplasma pneumoniae (Filterable agent of primary atypical pneumonia) Mycoplasma pneumoniae (Filterable agent of primary atypical pneumonia) | |||||||||||||||
| Method | subtomogram averaging / cryo EM / Resolution: 3.4 Å | |||||||||||||||
 Authors Authors | Tegunov D / Xue L / Dienemann C / Cramer P / Mahamid J | |||||||||||||||
| Funding support |  Germany, 4 items Germany, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Nat Methods / Year: 2021 Journal: Nat Methods / Year: 2021Title: Multi-particle cryo-EM refinement with M visualizes ribosome-antibiotic complex at 3.5 Å in cells. Authors: Dimitry Tegunov / Liang Xue / Christian Dienemann / Patrick Cramer / Julia Mahamid /  Abstract: Cryo-electron microscopy (cryo-EM) enables macromolecular structure determination in vitro and inside cells. In addition to aligning individual particles, accurate registration of sample motion and ...Cryo-electron microscopy (cryo-EM) enables macromolecular structure determination in vitro and inside cells. In addition to aligning individual particles, accurate registration of sample motion and three-dimensional deformation during exposures are crucial for achieving high-resolution reconstructions. Here we describe M, a software tool that establishes a reference-based, multi-particle refinement framework for cryo-EM data and couples a comprehensive spatial deformation model to in silico correction of electron-optical aberrations. M provides a unified optimization framework for both frame-series and tomographic tilt-series data. We show that tilt-series data can provide the same resolution as frame-series data on a purified protein specimen, indicating that the alignment step no longer limits the resolution obtainable from tomographic data. In combination with Warp and RELION, M resolves to residue level a 70S ribosome bound to an antibiotic inside intact bacterial cells. Our work provides a computational tool that facilitates structural biology in cells. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11999.map.gz emd_11999.map.gz | 191.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11999-v30.xml emd-11999-v30.xml emd-11999.xml emd-11999.xml | 12.8 KB 12.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_11999_fsc.xml emd_11999_fsc.xml | 13 KB | Display |  FSC data file FSC data file |
| Images |  emd_11999.png emd_11999.png | 85.2 KB | ||
| Masks |  emd_11999_msk_1.map emd_11999_msk_1.map | 166.4 MB |  Mask map Mask map | |
| Others |  emd_11999_half_map_1.map.gz emd_11999_half_map_1.map.gz emd_11999_half_map_2.map.gz emd_11999_half_map_2.map.gz | 156.7 MB 156.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11999 http://ftp.pdbj.org/pub/emdb/structures/EMD-11999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11999 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11999 | HTTPS FTP |
-Related structure data
| Related structure data |  7oodM C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10499 (Title: Tilt series of native M. pneumoniae cells treated with chloramphenicol EMPIAR-10499 (Title: Tilt series of native M. pneumoniae cells treated with chloramphenicolData size: 83.8 Data #1: Unaligned tilt movies of M. pneumoniae [tilt series])  EMPIAR-10731 (Title: Locating Macromolecular Assemblies in Cells by 2D Template Matching with cisTEM EMPIAR-10731 (Title: Locating Macromolecular Assemblies in Cells by 2D Template Matching with cisTEMData size: 67.8 Data #1: 50S templates of Mycoplasma pneumoniae used for 2D and 3D template matching [reconstructed volumes] Data #2: Single-exposure micrographs of untilted Mycoplasma pneumoniae cells [micrographs - single frame] Data #3: Tomographic tilt series of Mycoplasma pneumoniae cells [tilt series] Data #4: Tomographic reconstructions of Mycoplasma pneumoniae cells [reconstructed volumes]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11999.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11999.map.gz / Format: CCP4 / Size: 209.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.8502 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11999_msk_1.map emd_11999_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
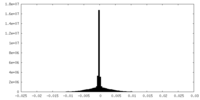
| Density Histograms |
-Half map: #1
| File | emd_11999_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_11999_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : 70S ribosome with chloramphenicol
| Entire | Name: 70S ribosome with chloramphenicol |
|---|---|
| Components |
|
-Supramolecule #1: 70S ribosome with chloramphenicol
| Supramolecule | Name: 70S ribosome with chloramphenicol / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  Mycoplasma pneumoniae (Filterable agent of primary atypical pneumonia) Mycoplasma pneumoniae (Filterable agent of primary atypical pneumonia) |
| Molecular weight | Experimental: 2.7 MDa |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | subtomogram averaging |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 120.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)