[English] 日本語
 Yorodumi
Yorodumi- EMDB-11471: Extended cryo-EM map of native human uromodulin filament core at ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11471 | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Title | Extended cryo-EM map of native human uromodulin filament core at 4.7 A resolution | |||||||||||||||
 Map data Map data | sharpened map | |||||||||||||||
 Sample Sample |
| |||||||||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.7 Å | |||||||||||||||
 Authors Authors | Stanisich JJ / Zyla D / Afanasyev P / Xu J / Pilhofer M / Boehringer D / Glockshuber R | |||||||||||||||
| Funding support |  Switzerland, 4 items Switzerland, 4 items
| |||||||||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: The cryo-EM structure of the human uromodulin filament core reveals a unique assembly mechanism. Authors: Jessica J Stanisich / Dawid S Zyla / Pavel Afanasyev / Jingwei Xu / Anne Kipp / Eric Olinger / Olivier Devuyst / Martin Pilhofer / Daniel Boehringer / Rudi Glockshuber /    Abstract: The glycoprotein uromodulin (UMOD) is the most abundant protein in human urine and forms filamentous homopolymers that encapsulate and aggregate uropathogens, promoting pathogen clearance by urine ...The glycoprotein uromodulin (UMOD) is the most abundant protein in human urine and forms filamentous homopolymers that encapsulate and aggregate uropathogens, promoting pathogen clearance by urine excretion. Despite its critical role in the innate immune response against urinary tract infections, the structural basis and mechanism of UMOD polymerization remained unknown. Here, we present the cryo-EM structure of the UMOD filament core at 3.5 Å resolution, comprised of the bipartite zona pellucida (ZP) module in a helical arrangement with a rise of ~65 Å and a twist of ~180°. The immunoglobulin-like ZPN and ZPC subdomains of each monomer are separated by a long linker that interacts with the preceding ZPC and following ZPN subdomains by β-sheet complementation. The unique filament architecture suggests an assembly mechanism in which subunit incorporation could be synchronized with proteolytic cleavage of the C-terminal pro-peptide that anchors assembly-incompetent UMOD precursors to the membrane. | |||||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11471.map.gz emd_11471.map.gz | 116.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11471-v30.xml emd-11471-v30.xml emd-11471.xml emd-11471.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11471.png emd_11471.png | 81.6 KB | ||
| Masks |  emd_11471_msk_1.map emd_11471_msk_1.map | 125 MB |  Mask map Mask map | |
| Others |  emd_11471_additional.map.gz emd_11471_additional.map.gz emd_11471_half_map_1.map.gz emd_11471_half_map_1.map.gz emd_11471_half_map_2.map.gz emd_11471_half_map_2.map.gz | 115.9 MB 23.2 MB 23.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11471 http://ftp.pdbj.org/pub/emdb/structures/EMD-11471 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11471 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11471 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_11471.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11471.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | sharpened map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.084 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_11471_msk_1.map emd_11471_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
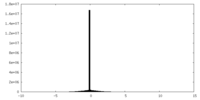
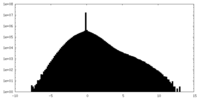
| Density Histograms |
-Additional map: unsharpened map
| File | emd_11471_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | unsharpened map | ||||||||||||
| Projections & Slices |
| ||||||||||||
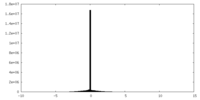
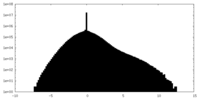
| Density Histograms |
-Half map: Half map A
| File | emd_11471_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map A | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map B
| File | emd_11471_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map B | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Native human uromodulin filament core
| Entire | Name: Native human uromodulin filament core |
|---|---|
| Components |
|
-Supramolecule #1: Native human uromodulin filament core
| Supramolecule | Name: Native human uromodulin filament core / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all Details: Native uromodulin was purified from healthy human urine. |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Experimental: 1.28 kDa/nm |
-Macromolecule #1: Native human uromodulin
| Macromolecule | Name: Native human uromodulin / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LLEHRLECGA NDMKVSLGKC QLKSLGFDKV FMY LSDSRC SGFNDRDNRD WVSVVTPARD GPCGTVLTRN ETHATYSNTL YLADEIIIRD LNIK INFAC SYPLDMKVSL KTALQPMVSA LNIRVGGTGM FTVRMALFQT PSYTQPYQGS SVTLS TEAF LYVGTMLDGG ...String: LLEHRLECGA NDMKVSLGKC QLKSLGFDKV FMY LSDSRC SGFNDRDNRD WVSVVTPARD GPCGTVLTRN ETHATYSNTL YLADEIIIRD LNIK INFAC SYPLDMKVSL KTALQPMVSA LNIRVGGTGM FTVRMALFQT PSYTQPYQGS SVTLS TEAF LYVGTMLDGG DLSRFALLMT NCYATPSSNA TDPLKYFIIQ DRCPHTRDST IQVVEN GES SQGRFSVQMF RFAGNYDLVY LHCEVYLCDT MNEKCKPTCS G |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | filament |
- Sample preparation
Sample preparation
| Concentration | 1.58 mg/mL |
|---|---|
| Buffer | pH: 8.2 / Component - Concentration: 0.5 mM / Component - Formula: EDTA / Component - Name: Ethylenediaminetetraacetic acid |
| Grid | Model: Homemade / Material: COPPER / Support film - Material: CARBON / Support film - topology: LACEY / Support film - Film thickness: 3.0 nm |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 95 % / Chamber temperature: 282 K / Instrument: FEI VITROBOT MARK IV Details: 3.5 ul sample, 30 s wait time, 0.5 s drain time, 13.5 s blotting from the back. |
| Details | individual, isolated fibers |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Number grids imaged: 2 / Number real images: 9543 / Average exposure time: 6.0 sec. / Average electron dose: 45.0 e/Å2 / Details: Data was joined from two sessions |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.3 µm / Nominal defocus min: 0.8 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller









 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)