[English] 日本語
 Yorodumi
Yorodumi- EMDB-6569: The cryo-EM structure of yeast spliceosomal U4/U6.U5 tri-snRNP (i... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-6569 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | The cryo-EM structure of yeast spliceosomal U4/U6.U5 tri-snRNP (improved map for U5 RNA region at 3.70 A) | |||||||||
 Map data Map data | Reconstruction of tri-snRNP | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | tri-snRNP / pre-mRNA splicing | |||||||||
| Function / homology |  Function and homology information Function and homology informationmRNA decay by 5' to 3' exoribonuclease / spliceosomal conformational changes to generate catalytic conformation / snoRNA splicing / Lsm1-7-Pat1 complex / U6 snRNP / snoRNA guided rRNA 2'-O-methylation / deadenylation-dependent decapping of nuclear-transcribed mRNA / generation of catalytic spliceosome for first transesterification step / box C/D sno(s)RNA 3'-end processing / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) ...mRNA decay by 5' to 3' exoribonuclease / spliceosomal conformational changes to generate catalytic conformation / snoRNA splicing / Lsm1-7-Pat1 complex / U6 snRNP / snoRNA guided rRNA 2'-O-methylation / deadenylation-dependent decapping of nuclear-transcribed mRNA / generation of catalytic spliceosome for first transesterification step / box C/D sno(s)RNA 3'-end processing / spliceosome conformational change to release U4 (or U4atac) and U1 (or U11) / box C/D methylation guide snoRNP complex / U4/U6 snRNP / 7-methylguanosine cap hypermethylation / pICln-Sm protein complex / sno(s)RNA-containing ribonucleoprotein complex / small nuclear ribonucleoprotein complex / splicing factor binding / SMN-Sm protein complex / spliceosomal tri-snRNP complex / snRNP binding / commitment complex / U2-type prespliceosome assembly / U2-type catalytic step 2 spliceosome / U4 snRNA binding / U2 snRNP / U1 snRNP / P-body assembly / U4 snRNP / U2-type prespliceosome / tRNA processing / U3 snoRNA binding / poly(U) RNA binding / precatalytic spliceosome / generation of catalytic spliceosome for second transesterification step / Major pathway of rRNA processing in the nucleolus and cytosol / mRNA 5'-splice site recognition / mRNA 3'-splice site recognition / spliceosomal complex assembly / spliceosomal tri-snRNP complex assembly / Prp19 complex / U5 snRNP / nuclear-transcribed mRNA catabolic process / U5 snRNA binding / pre-mRNA intronic binding / spliceosomal snRNP assembly / U2 snRNA binding / U6 snRNA binding / U1 snRNA binding / U4/U6 x U5 tri-snRNP complex / cellular response to glucose starvation / catalytic step 2 spliceosome / maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / spliceosomal complex / maturation of SSU-rRNA / P-body / small-subunit processome / mRNA splicing, via spliceosome / metallopeptidase activity / rRNA processing / nucleic acid binding / RNA helicase activity / RNA helicase / mRNA binding / GTPase activity / GTP binding / nucleolus / ATP hydrolysis activity / mitochondrion / RNA binding / nucleoplasm / ATP binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.77 Å | |||||||||
 Authors Authors | Wan R / Yan C / Bai R / Wang L / Huang M / Wong C / Shi Y | |||||||||
 Citation Citation |  Journal: Science / Year: 2016 Journal: Science / Year: 2016Title: The 3.8 Å structure of the U4/U6.U5 tri-snRNP: Insights into spliceosome assembly and catalysis. Authors: Ruixue Wan / Chuangye Yan / Rui Bai / Lin Wang / Min Huang / Catherine C L Wong / Yigong Shi /  Abstract: Splicing of precursor messenger RNA is accomplished by a dynamic megacomplex known as the spliceosome. Assembly of a functional spliceosome requires a preassembled U4/U6.U5 tri-snRNP complex, which ...Splicing of precursor messenger RNA is accomplished by a dynamic megacomplex known as the spliceosome. Assembly of a functional spliceosome requires a preassembled U4/U6.U5 tri-snRNP complex, which comprises the U5 small nuclear ribonucleoprotein (snRNP), the U4 and U6 small nuclear RNA (snRNA) duplex, and a number of protein factors. Here we report the three-dimensional structure of a Saccharomyces cerevisiae U4/U6.U5 tri-snRNP at an overall resolution of 3.8 angstroms by single-particle electron cryomicroscopy. The local resolution for the core regions of the tri-snRNP reaches 3.0 to 3.5 angstroms, allowing construction of a refined atomic model. Our structure contains U5 snRNA, the extensively base-paired U4/U6 snRNA, and 30 proteins including Prp8 and Snu114, which amount to 8495 amino acids and 263 nucleotides with a combined molecular mass of ~1 megadalton. The catalytic nucleotide U80 from U6 snRNA exists in an inactive conformation, stabilized by its base-pairing interactions with U4 snRNA and protected by Prp3. Pre-messenger RNA is bound in the tri-snRNP through base-pairing interactions with U6 snRNA and loop I of U5 snRNA. This structure, together with that of the spliceosome, reveals the molecular choreography of the snRNAs in the activation process of the spliceosomal ribozyme. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_6569.map.gz emd_6569.map.gz | 116.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-6569-v30.xml emd-6569-v30.xml emd-6569.xml emd-6569.xml | 10.1 KB 10.1 KB | Display Display |  EMDB header EMDB header |
| Images |  400_6569.gif 400_6569.gif 80_6569.gif 80_6569.gif | 55.5 KB 3.5 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-6569 http://ftp.pdbj.org/pub/emdb/structures/EMD-6569 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6569 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-6569 | HTTPS FTP |
-Related structure data
| Related structure data |  3jcmMC  6561C  6562C  6563C  6564C  6565C  6566C  6567C  6568C  6570C  6571C  6572C  6573C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_6569.map.gz / Format: CCP4 / Size: 122.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_6569.map.gz / Format: CCP4 / Size: 122.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of tri-snRNP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
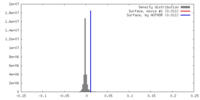
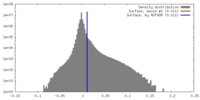
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : U4/U6.U5 tri-snRNP from Saccharomyces cerevisiae
| Entire | Name: U4/U6.U5 tri-snRNP from Saccharomyces cerevisiae |
|---|---|
| Components |
|
-Supramolecule #1000: U4/U6.U5 tri-snRNP from Saccharomyces cerevisiae
| Supramolecule | Name: U4/U6.U5 tri-snRNP from Saccharomyces cerevisiae / type: sample / ID: 1000 / Number unique components: 34 |
|---|
-Supramolecule #1: U4/U6.U5 tri-snRNP
| Supramolecule | Name: U4/U6.U5 tri-snRNP / type: organelle_or_cellular_component / ID: 1 / Recombinant expression: No / Database: NCBI |
|---|---|
| Source (natural) | Organism:  |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Vitrification | Cryogen name: ETHANE / Instrument: FEI VITROBOT MARK IV |
|---|
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Date | Aug 8, 2015 |
| Image recording | Category: FILM / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Digitization - Scanner: TEMSCAN / Number real images: 3141 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 3.77 Å / Resolution method: OTHER / Software - Name: RELION / Number images used: 172134 |
|---|
 Movie
Movie Controller
Controller


























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)