[English] 日本語
 Yorodumi
Yorodumi- EMDB-25683: CryoEM structure of Methylococcus capsulatus (Bath) pMMO treated ... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of Methylococcus capsulatus (Bath) pMMO treated with potassium cyanide in a native lipid nanodisc at 3.65 Angstrom resolution | |||||||||
 Map data Map data | CryoEM structure of Methylococcus capsulatus (Bath) pMMO treated with potassium cyanide in a native lipid nanodisc at 3.65 angstrom resolution | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | complex / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationmethane monooxygenase (particulate) / methane monooxygenase (soluble) / methane monooxygenase [NAD(P)H] activity / monooxygenase activity / membrane / metal ion binding Similarity search - Function | |||||||||
| Biological species |  Methylococcus capsulatus str. Bath (bacteria) Methylococcus capsulatus str. Bath (bacteria) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.65 Å | |||||||||
 Authors Authors | Koo CW / Rosenzweig AC | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Science / Year: 2022 Journal: Science / Year: 2022Title: Recovery of particulate methane monooxygenase structure and activity in a lipid bilayer. Authors: Christopher W Koo / Frank J Tucci / Yuan He / Amy C Rosenzweig /  Abstract: Bacterial methane oxidation using the enzyme particulate methane monooxygenase (pMMO) contributes to the removal of environmental methane, a potent greenhouse gas. Crystal structures determined using ...Bacterial methane oxidation using the enzyme particulate methane monooxygenase (pMMO) contributes to the removal of environmental methane, a potent greenhouse gas. Crystal structures determined using inactive, detergent-solubilized pMMO lack several conserved regions neighboring the proposed active site. We show that reconstituting pMMO in nanodiscs with lipids extracted from the native organism restores methane oxidation activity. Multiple nanodisc-embedded pMMO structures determined by cryo-electron microscopy to 2.14- to 2.46-angstrom resolution reveal the structure of pMMO in a lipid environment. The resulting model includes stabilizing lipids, regions of the PmoA and PmoC subunits not observed in prior structures, and a previously undetected copper-binding site in the PmoC subunit with an adjacent hydrophobic cavity. These structures provide a revised framework for understanding and engineering pMMO function. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_25683.map.gz emd_25683.map.gz | 96.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-25683-v30.xml emd-25683-v30.xml emd-25683.xml emd-25683.xml | 17.1 KB 17.1 KB | Display Display |  EMDB header EMDB header |
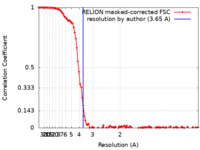
| FSC (resolution estimation) |  emd_25683_fsc.xml emd_25683_fsc.xml | 10.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_25683.png emd_25683.png | 126.9 KB | ||
| Filedesc metadata |  emd-25683.cif.gz emd-25683.cif.gz | 5.8 KB | ||
| Others |  emd_25683_additional_1.map.gz emd_25683_additional_1.map.gz | 90.1 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-25683 http://ftp.pdbj.org/pub/emdb/structures/EMD-25683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-25683 | HTTPS FTP |
-Related structure data
| Related structure data |  7t4oMC  7s4hC  7s4iC  7s4jC  7s4kC  7s4lC  7s4mC  7t4pC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_25683.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_25683.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of Methylococcus capsulatus (Bath) pMMO treated with potassium cyanide in a native lipid nanodisc at 3.65 angstrom resolution | ||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.511 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Additional map: CryoEM structure of Methylococcus capsulatus (Bath) pMMO treated...
| File | emd_25683_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | CryoEM structure of Methylococcus capsulatus (Bath) pMMO treated with potassium cyanide in a native lipid nanodisc at 3.65 angstrom resolution, unsharpened | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : pMMO complex in a native lipid nanodisc
| Entire | Name: pMMO complex in a native lipid nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: pMMO complex in a native lipid nanodisc
| Supramolecule | Name: pMMO complex in a native lipid nanodisc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#3 |
|---|---|
| Source (natural) | Organism:  Methylococcus capsulatus str. Bath (bacteria) Methylococcus capsulatus str. Bath (bacteria) |
-Macromolecule #1: Particulate methane monooxygenase alpha subunit
| Macromolecule | Name: Particulate methane monooxygenase alpha subunit / type: protein_or_peptide / ID: 1 / Number of copies: 3 / Enantiomer: LEVO / EC number: methane monooxygenase (particulate) |
|---|---|
| Source (natural) | Organism:  Methylococcus capsulatus str. Bath (bacteria) / Strain: ATCC 33009 / NCIMB 11132 / Bath Methylococcus capsulatus str. Bath (bacteria) / Strain: ATCC 33009 / NCIMB 11132 / Bath |
| Molecular weight | Theoretical: 46.129746 KDa |
| Sequence | String: MKTIKDRIAK WSAIGLLSAV AATAFYAPSA SAHGEKSQAA FMRMRTIHWY DLSWSKEKVK INETVEIKGK FHVFEGWPET VDEPDVAFL NVGMPGPVFI RKESYIGGQL VPRSVRLEIG KTYDFRVVLK ARRPGDWHVH TMMNVQGGGP IIGPGKWITV E GSMSEFRN ...String: MKTIKDRIAK WSAIGLLSAV AATAFYAPSA SAHGEKSQAA FMRMRTIHWY DLSWSKEKVK INETVEIKGK FHVFEGWPET VDEPDVAFL NVGMPGPVFI RKESYIGGQL VPRSVRLEIG KTYDFRVVLK ARRPGDWHVH TMMNVQGGGP IIGPGKWITV E GSMSEFRN PVTTLTGQTV DLENYNEGNT YFWHAFWFAI GVAWIGYWSR RPIFIPRLLM VDAGRADELV SATDRKVAMG FL AATILIV VMAMSSANSK YPITIPLQAG TMRGMKPLEL PAPTVSVKVE DATYRVPGRA MRMKLTITNH GNSPIRLGEF YTA SVRFLD SDVYKDTTGY PEDLLAEDGL SVSDNSPLAP GETRTVDVTA SDAAWEVYRL SDIIYDPDSR FAGLLFFFDA TGNR QVVQI DAPLIPSFM UniProtKB: Particulate methane monooxygenase alpha subunit |
-Macromolecule #2: Ammonia monooxygenase/methane monooxygenase, subunit C family protein
| Macromolecule | Name: Ammonia monooxygenase/methane monooxygenase, subunit C family protein type: protein_or_peptide / ID: 2 / Number of copies: 3 / Enantiomer: LEVO / EC number: methane monooxygenase (soluble) |
|---|---|
| Source (natural) | Organism:  Methylococcus capsulatus str. Bath (bacteria) / Strain: ATCC 33009 / NCIMB 11132 / Bath Methylococcus capsulatus str. Bath (bacteria) / Strain: ATCC 33009 / NCIMB 11132 / Bath |
| Molecular weight | Theoretical: 29.839309 KDa |
| Sequence | String: MAATTIGGAA AAEAPLLDKK WLTFALAIYT VFYLWVRWYE GVYGWSAGLD SFAPEFETYW MNFLYTEIVL EIVTASILWG YLWKTRDRN LAALTPREEL RRNFTHLVWL VAYAWAIYWG ASYFTEQDGT WHQTIVRDTD FTPSHIIEFY LSYPIYIITG F AAFIYAKT ...String: MAATTIGGAA AAEAPLLDKK WLTFALAIYT VFYLWVRWYE GVYGWSAGLD SFAPEFETYW MNFLYTEIVL EIVTASILWG YLWKTRDRN LAALTPREEL RRNFTHLVWL VAYAWAIYWG ASYFTEQDGT WHQTIVRDTD FTPSHIIEFY LSYPIYIITG F AAFIYAKT RLPFFAKGIS LPYLVLVVGP FMILPNVGLN EWGHTFWFME ELFVAPLHYG FVIFGWLALA VMGTLTQTFY SF AQGGLGQ SLCEAVDEGL IAK UniProtKB: Ammonia monooxygenase/methane monooxygenase, subunit C family protein |
-Macromolecule #3: Particulate methane monooxygenase beta subunit
| Macromolecule | Name: Particulate methane monooxygenase beta subunit / type: protein_or_peptide / ID: 3 / Number of copies: 3 / Enantiomer: LEVO / EC number: methane monooxygenase (particulate) |
|---|---|
| Source (natural) | Organism:  Methylococcus capsulatus str. Bath (bacteria) / Strain: ATCC 33009 / NCIMB 11132 / Bath Methylococcus capsulatus str. Bath (bacteria) / Strain: ATCC 33009 / NCIMB 11132 / Bath |
| Molecular weight | Theoretical: 28.445098 KDa |
| Sequence | String: MSAAQSAVRS HAEAVQVSRT IDWMALFVVF FVIVGSYHIH AMLTMGDWDF WSDWKDRRLW VTVTPIVLVT FPAAVQSYLW ERYRLPWGA TVCVLGLLLG EWINRYFNFW GWTYFPINFV FPASLVPGAI ILDTVLMLSG SYLFTAIVGA MGWGLIFYPG N WPIIAPLH ...String: MSAAQSAVRS HAEAVQVSRT IDWMALFVVF FVIVGSYHIH AMLTMGDWDF WSDWKDRRLW VTVTPIVLVT FPAAVQSYLW ERYRLPWGA TVCVLGLLLG EWINRYFNFW GWTYFPINFV FPASLVPGAI ILDTVLMLSG SYLFTAIVGA MGWGLIFYPG N WPIIAPLH VPVEYNGMLM SIADIQGYNY VRTGTPEYIR MVEKGTLRTF GKDVAPVSAF FSAFMSILIY FMWHFIGRWF SN ERFLQST UniProtKB: Particulate methane monooxygenase beta subunit |
-Macromolecule #4: COPPER (II) ION
| Macromolecule | Name: COPPER (II) ION / type: ligand / ID: 4 / Number of copies: 5 / Formula: CU |
|---|---|
| Molecular weight | Theoretical: 63.546 Da |
| Chemical component information |  ChemComp-CU: |
-Macromolecule #5: 1,2-DIDECANOYL-SN-GLYCERO-3-PHOSPHOCHOLINE
| Macromolecule | Name: 1,2-DIDECANOYL-SN-GLYCERO-3-PHOSPHOCHOLINE / type: ligand / ID: 5 / Number of copies: 9 / Formula: P1O |
|---|---|
| Molecular weight | Theoretical: 566.728 Da |
| Chemical component information |  ChemComp-P1O: |
-Macromolecule #6: DIUNDECYL PHOSPHATIDYL CHOLINE
| Macromolecule | Name: DIUNDECYL PHOSPHATIDYL CHOLINE / type: ligand / ID: 6 / Number of copies: 3 / Formula: PLC |
|---|---|
| Molecular weight | Theoretical: 622.834 Da |
| Chemical component information |  ChemComp-PLC: |
-Macromolecule #7: DECANE
| Macromolecule | Name: DECANE / type: ligand / ID: 7 / Number of copies: 9 / Formula: D10 |
|---|---|
| Molecular weight | Theoretical: 142.282 Da |
| Chemical component information |  ChemComp-D10: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.3 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 1.2 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 2.2 µm / Nominal defocus min: 0.8 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller










 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)