+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22806 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
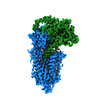
| Title | Human WLS in complex with WNT8A | |||||||||
 Map data Map data | Density modified map | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology information: / : / Wnt protein secretion / neural crest cell fate commitment / positive regulation of Wnt protein secretion / WNT ligand biogenesis and trafficking / secondary palate development / cementum mineralization / beta-catenin destruction complex disassembly / hindbrain development ...: / : / Wnt protein secretion / neural crest cell fate commitment / positive regulation of Wnt protein secretion / WNT ligand biogenesis and trafficking / secondary palate development / cementum mineralization / beta-catenin destruction complex disassembly / hindbrain development / Wnt-protein binding / exocrine pancreas development / frizzled binding / Class B/2 (Secretin family receptors) / Disassembly of the destruction complex and recruitment of AXIN to the membrane / anterior/posterior axis specification / midbrain development / organelle membrane / cell fate commitment / mesoderm formation / positive regulation of Wnt signaling pathway / canonical Wnt signaling pathway / endomembrane system / response to retinoic acid / TCF dependent signaling in response to WNT / cytokine activity / intracellular protein transport / trans-Golgi network / neuron differentiation / Wnt signaling pathway / positive regulation of canonical Wnt signaling pathway / endocytic vesicle membrane / early endosome membrane / positive regulation of canonical NF-kappaB signal transduction / cytoplasmic vesicle / collagen-containing extracellular matrix / early endosome / receptor ligand activity / Golgi membrane / endoplasmic reticulum membrane / Golgi apparatus / endoplasmic reticulum / positive regulation of transcription by RNA polymerase II / extracellular space / extracellular exosome / extracellular region / identical protein binding / plasma membrane / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.19 Å | |||||||||
 Authors Authors | Nygaard R / Jia Y / Kim J / Ross D / Parisi G / Clarke OB / Virshup DM / Mancia F | |||||||||
| Funding support |  United States, United States,  Singapore, 2 items Singapore, 2 items
| |||||||||
 Citation Citation |  Journal: Cell / Year: 2021 Journal: Cell / Year: 2021Title: Structural Basis of WLS/Evi-Mediated Wnt Transport and Secretion. Authors: Rie Nygaard / Jia Yu / Jonathan Kim / Daniel R Ross / Giacomo Parisi / Oliver B Clarke / David M Virshup / Filippo Mancia /   Abstract: Wnts are evolutionarily conserved ligands that signal at short range to regulate morphogenesis, cell fate, and stem cell renewal. The first and essential steps in Wnt secretion are their O- ...Wnts are evolutionarily conserved ligands that signal at short range to regulate morphogenesis, cell fate, and stem cell renewal. The first and essential steps in Wnt secretion are their O-palmitoleation and subsequent loading onto the dedicated transporter Wntless/evenness interrupted (WLS/Evi). We report the 3.2 Å resolution cryogenic electron microscopy (cryo-EM) structure of palmitoleated human WNT8A in complex with WLS. This is accompanied by biochemical experiments to probe the physiological implications of the observed association. The WLS membrane domain has close structural homology to G protein-coupled receptors (GPCRs). A Wnt hairpin inserts into a conserved hydrophobic cavity in the GPCR-like domain, and the palmitoleate protrudes between two helices into the bilayer. A conformational switch of highly conserved residues on a separate Wnt hairpin might contribute to its transfer to receiving cells. This work provides molecular-level insights into a central mechanism in animal body plan development and stem cell biology. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22806.map.gz emd_22806.map.gz | 9.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22806-v30.xml emd-22806-v30.xml emd-22806.xml emd-22806.xml | 21.6 KB 21.6 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_22806.png emd_22806.png | 86.7 KB | ||
| Masks |  emd_22806_msk_1.map emd_22806_msk_1.map emd_22806_msk_2.map emd_22806_msk_2.map | 347.6 MB 347.6 MB |  Mask map Mask map | |
| Others |  emd_22806_additional_1.map.gz emd_22806_additional_1.map.gz emd_22806_half_map_1.map.gz emd_22806_half_map_1.map.gz emd_22806_half_map_2.map.gz emd_22806_half_map_2.map.gz | 9.1 MB 322.2 MB 322.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22806 http://ftp.pdbj.org/pub/emdb/structures/EMD-22806 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22806 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22806 | HTTPS FTP |
-Validation report
| Summary document |  emd_22806_validation.pdf.gz emd_22806_validation.pdf.gz | 958.6 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22806_full_validation.pdf.gz emd_22806_full_validation.pdf.gz | 958.2 KB | Display | |
| Data in XML |  emd_22806_validation.xml.gz emd_22806_validation.xml.gz | 14.9 KB | Display | |
| Data in CIF |  emd_22806_validation.cif.gz emd_22806_validation.cif.gz | 16.5 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22806 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22806 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22806 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22806 | HTTPS FTP |
-Related structure data
| Related structure data |  7kc4MC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10699 (Title: Human WLS in complex with WNT8A / Data size: 1.9 TB EMPIAR-10699 (Title: Human WLS in complex with WNT8A / Data size: 1.9 TBData #1: Unaligned multi frame micrographs of WNT8A WLS complex [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22806.map.gz / Format: CCP4 / Size: 9.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22806.map.gz / Format: CCP4 / Size: 9.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Density modified map | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.83 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
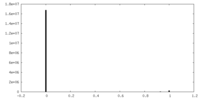
-Mask #1
| File |  emd_22806_msk_1.map emd_22806_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
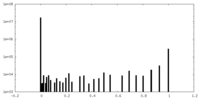
-Mask #2
| File |  emd_22806_msk_2.map emd_22806_msk_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
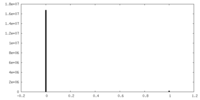
-Additional map: Blurred map (B=100)
| File | emd_22806_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Blurred map (B=100) | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
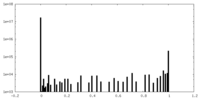
-Half map: Half map
| File | emd_22806_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: Half map
| File | emd_22806_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Half map | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Complex of WLS/Evi and WNT8A in nanodisc
| Entire | Name: Complex of WLS/Evi and WNT8A in nanodisc |
|---|---|
| Components |
|
-Supramolecule #1: Complex of WLS/Evi and WNT8A in nanodisc
| Supramolecule | Name: Complex of WLS/Evi and WNT8A in nanodisc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 / Details: Nanodisc were formed using MSP1E3D1 and POPG lipid |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Recombinant expression | Organism:  Homo sapiens (human) / Recombinant strain: FreeStyle 293-F Cells Homo sapiens (human) / Recombinant strain: FreeStyle 293-F Cells |
| Molecular weight | Theoretical: 105.6 KDa |
-Macromolecule #1: Protein Wnt-8a
| Macromolecule | Name: Protein Wnt-8a / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 40.052359 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MGNLFMLWAA LGICCAAFSA SAWSVNNFLI TGPKAYLTYT TSVALGAQSG IEECKFQFAW ERWNCPENAL QLSTHNRLRS ATRETSFIH AISSAGVMYI ITKNCSMGDF ENCGCDGSNN GKTGGHGWIW GGCSDNVEFG ERISKLFVDS LEKGKDARAL M NLHNNRAG ...String: MGNLFMLWAA LGICCAAFSA SAWSVNNFLI TGPKAYLTYT TSVALGAQSG IEECKFQFAW ERWNCPENAL QLSTHNRLRS ATRETSFIH AISSAGVMYI ITKNCSMGDF ENCGCDGSNN GKTGGHGWIW GGCSDNVEFG ERISKLFVDS LEKGKDARAL M NLHNNRAG RLAVRATMKR TCKCHGISGS CSIQTCWLQL AEFREMGDYL KAKYDQALKI EMDKRQLRAG NSAEGHWVPA EA FLPSAEA ELIFLEESPD YCTCNSSLGI YGTEGRECLQ NSHNTSRWER RSCGRLCTEC GLQVEERKTE VISSCNCKFQ WCC TVKCDQ CRHVVSKYYC ARSPGSAQSL GKGSAGGWSH PQFEK |
-Macromolecule #2: Protein wntless homolog
| Macromolecule | Name: Protein wntless homolog / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 65.64943 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MAGAIIENMS TKKLCIVGGI LLVFQIIAFL VGGLIAPGPT TAVSYMSVKC VDARKNHHKT KWFVPWGPNH CDKIRDIEEA IPREIEAND IVFSVHIPLP HMEMSPWFQF MLFILQLDIA FKLNNQIREN AEVSMDVSLA YRDDAFAEWT EMAHERVPRK L KCTFTSPK ...String: MAGAIIENMS TKKLCIVGGI LLVFQIIAFL VGGLIAPGPT TAVSYMSVKC VDARKNHHKT KWFVPWGPNH CDKIRDIEEA IPREIEAND IVFSVHIPLP HMEMSPWFQF MLFILQLDIA FKLNNQIREN AEVSMDVSLA YRDDAFAEWT EMAHERVPRK L KCTFTSPK TPEHEGRYYE CDVLPFMEIG SVAHKFYLLN IRLPVNEKKK INVGIGEIKD IRLVGIHQNG GFTKVWFAMK TF LTPSIFI IMVWYWRRIT MMSRPPVLLE KVIFALGISM TFINIPVEWF SIGFDWTWML LFGDIRQGIF YAMLLSFWII FCG EHMMDQ HERNHIAGYW KQVGPIAVGS FCLFIFDMCE RGVQLTNPFY SIWTTDIGTE LAMAFIIVAG ICLCLYFLFL CFMV FQVFR NISGKQSSLP AMSKVRRLHY EGLIFRFKFL MLITLACAAM TVIFFIVSQV TEGHWKWGGV TVQVNSAFFT GIYGM WNLY VFALMFLYAP SHKNYGEDQS NGDLGVHSGE ELQLTTTITH VDGPTEIYKL TRKEAQEAEN LYFQSHHHHH HHHHHD YKD DDDK |
-Macromolecule #5: PALMITIC ACID
| Macromolecule | Name: PALMITIC ACID / type: ligand / ID: 5 / Number of copies: 1 / Formula: PLM |
|---|---|
| Molecular weight | Theoretical: 256.424 Da |
| Chemical component information |  ChemComp-PLM: |
-Macromolecule #6: 1-CIS-9-OCTADECANOYL-2-CIS-9-HEXADECANOYL PHOSPHATIDYL GLYCEROL
| Macromolecule | Name: 1-CIS-9-OCTADECANOYL-2-CIS-9-HEXADECANOYL PHOSPHATIDYL GLYCEROL type: ligand / ID: 6 / Number of copies: 3 / Formula: DR9 |
|---|---|
| Molecular weight | Theoretical: 746.991 Da |
| Chemical component information |  ChemComp-DR9: |
-Macromolecule #7: CHOLESTEROL HEMISUCCINATE
| Macromolecule | Name: CHOLESTEROL HEMISUCCINATE / type: ligand / ID: 7 / Number of copies: 1 / Formula: Y01 |
|---|---|
| Molecular weight | Theoretical: 486.726 Da |
| Chemical component information |  ChemComp-Y01: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 8 / Component:
| |||||||||
| Grid | Model: Quantifoil R0.6/1 / Material: GOLD / Pretreatment - Type: PLASMA CLEANING | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Digitization - Dimensions - Width: 5760 pixel / Digitization - Dimensions - Height: 4092 pixel / Digitization - Sampling interval: 5.0 µm / Number grids imaged: 1 / Number real images: 12747 / Average exposure time: 3.0 sec. / Average electron dose: 58.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
 Movie
Movie Controller
Controller















 Z
Z Y
Y X
X