[English] 日本語
 Yorodumi
Yorodumi- EMDB-22790: Nucleosome in interphase chromosome formed in Xenopus egg extract... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22790 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Nucleosome in interphase chromosome formed in Xenopus egg extract (oligo fraction) | |||||||||
 Map data Map data | Fliped and rescaled map for atomic model creation | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | nucleosome / interphase / cell cycle / chromatin / DNA BINDING PROTEIN / STRUCTURAL PROTEIN-DNA complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationstructural constituent of chromatin / nucleosome / heterochromatin formation / nucleosome assembly / protein heterodimerization activity / DNA binding / nucleoplasm / nucleus Similarity search - Function | |||||||||
| Biological species | ||||||||||
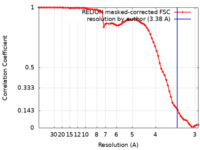
| Method | single particle reconstruction / cryo EM / Resolution: 3.38 Å | |||||||||
 Authors Authors | Arimura Y / Funabiki H | |||||||||
| Funding support |  United States, United States,  Japan, 2 items Japan, 2 items
| |||||||||
 Citation Citation |  Journal: Mol Cell / Year: 2021 Journal: Mol Cell / Year: 2021Title: Structural features of nucleosomes in interphase and metaphase chromosomes. Authors: Yasuhiro Arimura / Rochelle M Shih / Ruby Froom / Hironori Funabiki /  Abstract: Structural heterogeneity of nucleosomes in functional chromosomes is unknown. Here, we devise the template-, reference- and selection-free (TRSF) cryo-EM pipeline to simultaneously reconstruct cryo- ...Structural heterogeneity of nucleosomes in functional chromosomes is unknown. Here, we devise the template-, reference- and selection-free (TRSF) cryo-EM pipeline to simultaneously reconstruct cryo-EM structures of protein complexes from interphase or metaphase chromosomes. The reconstructed interphase and metaphase nucleosome structures are on average indistinguishable from canonical nucleosome structures, despite DNA sequence heterogeneity, cell-cycle-specific posttranslational modifications, and interacting proteins. Nucleosome structures determined by a decoy-classifying method and structure variability analyses reveal the nucleosome structural variations in linker DNA, histone tails, and nucleosome core particle configurations, suggesting that the opening of linker DNA, which is correlated with H2A C-terminal tail positioning, is suppressed in chromosomes. High-resolution (3.4-3.5 Å) nucleosome structures indicate DNA-sequence-independent stabilization of superhelical locations ±0-1 and ±3.5-4.5. The linker histone H1.8 preferentially binds to metaphase chromatin, from which chromatosome cryo-EM structures with H1.8 at the on-dyad position are reconstituted. This study presents the structural characteristics of nucleosomes in chromosomes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22790.map.gz emd_22790.map.gz | 28.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22790-v30.xml emd-22790-v30.xml emd-22790.xml emd-22790.xml | 39.4 KB 39.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22790_fsc.xml emd_22790_fsc.xml | 7.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_22790.png emd_22790.png | 59.4 KB | ||
| Filedesc metadata |  emd-22790.cif.gz emd-22790.cif.gz | 6.6 KB | ||
| Others |  emd_22790_additional_1.map.gz emd_22790_additional_1.map.gz emd_22790_additional_2.map.gz emd_22790_additional_2.map.gz emd_22790_additional_3.map.gz emd_22790_additional_3.map.gz emd_22790_additional_4.map.gz emd_22790_additional_4.map.gz emd_22790_additional_5.map.gz emd_22790_additional_5.map.gz emd_22790_additional_6.map.gz emd_22790_additional_6.map.gz emd_22790_additional_7.map.gz emd_22790_additional_7.map.gz emd_22790_additional_8.map.gz emd_22790_additional_8.map.gz emd_22790_half_map_1.map.gz emd_22790_half_map_1.map.gz emd_22790_half_map_2.map.gz emd_22790_half_map_2.map.gz | 15.3 MB 15.4 MB 15.3 MB 15.3 MB 15.1 MB 15.3 MB 15.1 MB 15.2 MB 23.5 MB 23.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22790 http://ftp.pdbj.org/pub/emdb/structures/EMD-22790 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22790 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22790 | HTTPS FTP |
-Related structure data
| Related structure data |  7kbdMC  7kbeC  7kbfC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10692 (Title: Oligo nucleosome fraction from interphase chromosome in Xenopus egg extract lot 1 EMPIAR-10692 (Title: Oligo nucleosome fraction from interphase chromosome in Xenopus egg extract lot 1Data size: 634.3 Data #1: oligo nucleosome fraction from the interphase chromosome in Xenopus egg extract lot1 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22790.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22790.map.gz / Format: CCP4 / Size: 30.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Fliped and rescaled map for atomic model creation | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.47 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Additional map: 3D classification map for structural variation analysis
+Half map: Nucleosome in interphase chromosome formed in Xenopus egg...
+Half map: Nucleosome in interphase chromosome formed in Xenopus egg...
- Sample components
Sample components
-Entire : Nucleosome in interphase chromosome formed in Xenopus egg extract...
| Entire | Name: Nucleosome in interphase chromosome formed in Xenopus egg extract (oligo fraction) |
|---|---|
| Components |
|
-Supramolecule #1: Nucleosome in interphase chromosome formed in Xenopus egg extract...
| Supramolecule | Name: Nucleosome in interphase chromosome formed in Xenopus egg extract (oligo fraction) type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: |
-Macromolecule #1: Histone H3.2
| Macromolecule | Name: Histone H3.2 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 15.421101 KDa |
| Sequence | String: MARTKQTARK STGGKAPRKQ LATKAARKSA PATGGVKKPH RYRPGTVALR EIRRYQKSTE LLIRKLPFQR LVREIAQDFK TDLRFQSSA VMALQEASEA YLVGLFEDTN LCAIHAKRVT IMPKDIQLAR RIRGERA UniProtKB: Histone H3.2 |
-Macromolecule #2: Histone H4
| Macromolecule | Name: Histone H4 / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 11.394426 KDa |
| Sequence | String: MSGRGKGGKG LGKGGAKRHR KVLRDNIQGI TKPAIRRLAR RGGVKRISGL IYEETRGVLK VFLENVIRDA VTYTEHAKRK TVTAMDVVY ALKRQGRTLY GFGG UniProtKB: Histone H4 |
-Macromolecule #3: Histone H2A
| Macromolecule | Name: Histone H2A / type: protein_or_peptide / ID: 3 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 14.883174 KDa |
| Sequence | String: MSGRGKKVQK AASGKASRSA KAGLQFPVGR IHRLLRKGNY AVRIGSGSAI YLAATLEYLC AEVLELAGNA ARDNKKLRIM PRHIQLAVR NDDELAKLFE GVTIADGGVL PNIQSALLPK KTVKGSSSSQ EPTAVESQEF UniProtKB: Histone H2A |
-Macromolecule #4: Histone H2B 1.1
| Macromolecule | Name: Histone H2B 1.1 / type: protein_or_peptide / ID: 4 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 13.965265 KDa |
| Sequence | String: MPEPAKSAPA PKKGSKKAVT KTQKKDGKKR RKSRKESYAI YVYKVLKQVH PDTGISSKAM SIMNSFVNDV FERIAGEASR LAHYNKRST ITSREIQTAV RLLLPGELAK HAVSEGTKAV TKYTSAK UniProtKB: Histone H2B 1.1 |
-Macromolecule #5: DNA (151-MER)
| Macromolecule | Name: DNA (151-MER) / type: dna / ID: 5 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 46.847863 KDa |
| Sequence | String: (DA)(DG)(DG)(DA)(DT)(DA)(DT)(DC)(DA)(DC) (DA)(DA)(DT)(DC)(DC)(DA)(DT)(DA)(DT)(DC) (DT)(DG)(DA)(DC)(DA)(DC)(DG)(DT)(DG) (DC)(DC)(DT)(DG)(DG)(DA)(DG)(DA)(DC)(DT) (DA) (DG)(DG)(DG)(DA)(DG)(DT) ...String: (DA)(DG)(DG)(DA)(DT)(DA)(DT)(DC)(DA)(DC) (DA)(DA)(DT)(DC)(DC)(DA)(DT)(DA)(DT)(DC) (DT)(DG)(DA)(DC)(DA)(DC)(DG)(DT)(DG) (DC)(DC)(DT)(DG)(DG)(DA)(DG)(DA)(DC)(DT) (DA) (DG)(DG)(DG)(DA)(DG)(DT)(DA)(DA) (DT)(DC)(DC)(DC)(DC)(DT)(DT)(DG)(DG)(DC) (DG)(DG) (DT)(DT)(DA)(DA)(DA)(DA)(DC) (DG)(DC)(DG)(DG)(DG)(DG)(DG)(DA)(DC)(DA) (DG)(DC)(DG) (DC)(DG)(DT)(DA)(DC)(DG) (DT)(DG)(DC)(DG)(DT)(DT)(DT)(DA)(DA)(DG) (DC)(DG)(DG)(DT) (DG)(DC)(DT)(DA)(DG) (DA)(DG)(DC)(DT)(DG)(DT)(DC)(DT)(DA)(DC) (DG)(DA)(DC)(DC)(DA) (DA)(DT)(DT)(DG) (DA)(DG)(DC)(DG)(DG)(DC)(DC)(DT)(DC)(DG) (DG)(DC)(DA)(DC)(DC)(DG) (DG)(DG)(DA) (DT)(DT)(DG)(DT)(DG)(DA)(DT)(DA) |
-Macromolecule #6: DNA (151-MER)
| Macromolecule | Name: DNA (151-MER) / type: dna / ID: 6 / Number of copies: 1 / Classification: DNA |
|---|---|
| Source (natural) | Organism: |
| Molecular weight | Theoretical: 46.355543 KDa |
| Sequence | String: (DT)(DA)(DT)(DC)(DA)(DC)(DA)(DA)(DT)(DC) (DC)(DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA) (DG)(DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA) (DA)(DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA) (DG) (DA)(DC)(DA)(DG)(DC)(DT) ...String: (DT)(DA)(DT)(DC)(DA)(DC)(DA)(DA)(DT)(DC) (DC)(DC)(DG)(DG)(DT)(DG)(DC)(DC)(DG)(DA) (DG)(DG)(DC)(DC)(DG)(DC)(DT)(DC)(DA) (DA)(DT)(DT)(DG)(DG)(DT)(DC)(DG)(DT)(DA) (DG) (DA)(DC)(DA)(DG)(DC)(DT)(DC)(DT) (DA)(DG)(DC)(DA)(DC)(DC)(DG)(DC)(DT)(DT) (DA)(DA) (DA)(DC)(DG)(DC)(DA)(DC)(DG) (DT)(DA)(DC)(DG)(DC)(DG)(DC)(DT)(DG)(DT) (DC)(DC)(DC) (DC)(DC)(DG)(DC)(DG)(DT) (DT)(DT)(DT)(DA)(DA)(DC)(DC)(DG)(DC)(DC) (DA)(DA)(DG)(DG) (DG)(DG)(DA)(DT)(DT) (DA)(DC)(DT)(DC)(DC)(DC)(DT)(DA)(DG)(DT) (DC)(DT)(DC)(DC)(DA) (DG)(DG)(DC)(DA) (DC)(DG)(DT)(DG)(DT)(DC)(DA)(DG)(DA)(DT) (DA)(DT)(DA)(DG)(DA)(DT) (DT)(DG)(DT) (DG)(DA)(DT)(DA)(DT)(DC)(DC)(DT) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 Component:
| |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: GOLD / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 10 sec. / Pretreatment - Atmosphere: OTHER / Details: 20W | |||||||||||||||||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Average electron dose: 33.11 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)