[English] 日本語
 Yorodumi
Yorodumi- EMDB-22281: CryoEM structure of human presequence protease in partial closed ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22281 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of human presequence protease in partial closed state 1 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Partial open state / HYDROLASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationMitochondrial protein import / Hydrolases; Acting on peptide bonds (peptidases); Metalloendopeptidases / amyloid-beta complex / growth cone lamellipodium / cellular response to norepinephrine stimulus / collateral sprouting in absence of injury / growth cone filopodium / microglia development / Formyl peptide receptors bind formyl peptides and many other ligands / axo-dendritic transport ...Mitochondrial protein import / Hydrolases; Acting on peptide bonds (peptidases); Metalloendopeptidases / amyloid-beta complex / growth cone lamellipodium / cellular response to norepinephrine stimulus / collateral sprouting in absence of injury / growth cone filopodium / microglia development / Formyl peptide receptors bind formyl peptides and many other ligands / axo-dendritic transport / regulation of Wnt signaling pathway / axon midline choice point recognition / regulation of synapse structure or activity / hippocampal neuron apoptotic process / astrocyte activation involved in immune response / NMDA selective glutamate receptor signaling pathway / regulation of spontaneous synaptic transmission / mating behavior / : / growth factor receptor binding / peptidase activator activity / Insertion of tail-anchored proteins into the endoplasmic reticulum membrane / PTB domain binding / positive regulation of amyloid fibril formation / Golgi-associated vesicle / astrocyte projection / Lysosome Vesicle Biogenesis / Deregulated CDK5 triggers multiple neurodegenerative pathways in Alzheimer's disease models / neuron remodeling / nuclear envelope lumen / dendrite development / TRAF6 mediated NF-kB activation / positive regulation of protein metabolic process / signaling receptor activator activity / negative regulation of long-term synaptic potentiation / Advanced glycosylation endproduct receptor signaling / transition metal ion binding / The NLRP3 inflammasome / modulation of excitatory postsynaptic potential / main axon / intracellular copper ion homeostasis / regulation of multicellular organism growth / ECM proteoglycans / response to insulin-like growth factor stimulus / positive regulation of T cell migration / regulation of presynapse assembly / neuronal dense core vesicle / Purinergic signaling in leishmaniasis infection / cellular response to manganese ion / positive regulation of chemokine production / Notch signaling pathway / swimming behavior / neuron projection maintenance / extracellular matrix organization / clathrin-coated pit / positive regulation of mitotic cell cycle / axonogenesis / Mitochondrial protein degradation / positive regulation of calcium-mediated signaling / ionotropic glutamate receptor signaling pathway / platelet alpha granule lumen / astrocyte activation / response to interleukin-1 / regulation of neuron apoptotic process / cellular response to cAMP / cellular response to copper ion / positive regulation of glycolytic process / endosome lumen / trans-Golgi network membrane / positive regulation of interleukin-1 beta production / protein serine/threonine kinase binding / dendritic shaft / positive regulation of long-term synaptic potentiation / learning / central nervous system development / Post-translational protein phosphorylation / adult locomotory behavior / serine-type endopeptidase inhibitor activity / locomotory behavior / microglial cell activation / cellular response to nerve growth factor stimulus / protein processing / metalloendopeptidase activity / TAK1-dependent IKK and NF-kappa-B activation / positive regulation of non-canonical NF-kappaB signal transduction / synapse organization / visual learning / recycling endosome / positive regulation of interleukin-6 production / positive regulation of JNK cascade / Golgi lumen / regulation of long-term neuronal synaptic plasticity / response to lead ion / cognition / Regulation of Insulin-like Growth Factor (IGF) transport and uptake by Insulin-like Growth Factor Binding Proteins (IGFBPs) / enzyme activator activity / cellular response to amyloid-beta / endocytosis / metallopeptidase activity / neuron projection development Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
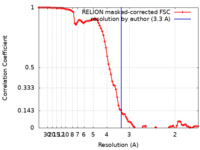
| Method | single particle reconstruction / cryo EM / Resolution: 3.3 Å | |||||||||
 Authors Authors | Liang WG / Zhao M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural basis for the mechanisms of human presequence protease conformational switch and substrate recognition. Authors: Wenguang G Liang / Juwina Wijaya / Hui Wei / Alex J Noble / Jordan M Mancl / Swansea Mo / David Lee / John V Lin King / Man Pan / Chang Liu / Carla M Koehler / Minglei Zhao / Clinton S ...Authors: Wenguang G Liang / Juwina Wijaya / Hui Wei / Alex J Noble / Jordan M Mancl / Swansea Mo / David Lee / John V Lin King / Man Pan / Chang Liu / Carla M Koehler / Minglei Zhao / Clinton S Potter / Bridget Carragher / Sheng Li / Wei-Jen Tang /  Abstract: Presequence protease (PreP), a 117 kDa mitochondrial M16C metalloprotease vital for mitochondrial proteostasis, degrades presequence peptides cleaved off from nuclear-encoded proteins and other ...Presequence protease (PreP), a 117 kDa mitochondrial M16C metalloprotease vital for mitochondrial proteostasis, degrades presequence peptides cleaved off from nuclear-encoded proteins and other aggregation-prone peptides, such as amyloid β (Aβ). PreP structures have only been determined in a closed conformation; thus, the mechanisms of substrate binding and selectivity remain elusive. Here, we leverage advanced vitrification techniques to overcome the preferential denaturation of one of two ~55 kDa homologous domains of PreP caused by air-water interface adsorption. Thereby, we elucidate cryoEM structures of three apo-PreP open states along with Aβ- and citrate synthase presequence-bound PreP at 3.3-4.6 Å resolution. Together with integrative biophysical and pharmacological approaches, these structures reveal the key stages of the PreP catalytic cycle and how the binding of substrates or PreP inhibitor drives a rigid body motion of the protein for substrate binding and catalysis. Together, our studies provide key mechanistic insights into M16C metalloproteases for future therapeutic innovations. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22281.map.gz emd_22281.map.gz | 49.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22281-v30.xml emd-22281-v30.xml emd-22281.xml emd-22281.xml | 24.6 KB 24.6 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_22281_fsc.xml emd_22281_fsc.xml | 9.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_22281.png emd_22281.png | 164.8 KB | ||
| Masks |  emd_22281_msk_1.map emd_22281_msk_1.map | 64 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-22281.cif.gz emd-22281.cif.gz | 7.1 KB | ||
| Others |  emd_22281_additional_1.map.gz emd_22281_additional_1.map.gz emd_22281_half_map_1.map.gz emd_22281_half_map_1.map.gz emd_22281_half_map_2.map.gz emd_22281_half_map_2.map.gz | 59.9 MB 49.6 MB 49.6 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22281 http://ftp.pdbj.org/pub/emdb/structures/EMD-22281 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22281 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22281 | HTTPS FTP |
-Related structure data
| Related structure data |  6xovMC  6xosC  6xotC  6xouC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10937 (Title: CryoEM of PreP prepared via Chameleon / Data size: 2.2 TB / Data #1: Apo-PreP micrographs [micrographs - multiframe] / Data #2: PreP half particles [micrographs - single frame] EMPIAR-10937 (Title: CryoEM of PreP prepared via Chameleon / Data size: 2.2 TB / Data #1: Apo-PreP micrographs [micrographs - multiframe] / Data #2: PreP half particles [micrographs - single frame]Data #3: Citrate synthase presequence bound PreP [micrographs - multiframe] Data #4: Amyloid beta bound PreP [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22281.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22281.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.855 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_22281_msk_1.map emd_22281_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Additional map: #1
| File | emd_22281_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_22281_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #2
| File | emd_22281_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : CryoEM map of Human Presequence Protease in partial close state 1
| Entire | Name: CryoEM map of Human Presequence Protease in partial close state 1 |
|---|---|
| Components |
|
-Supramolecule #1: CryoEM map of Human Presequence Protease in partial close state 1
| Supramolecule | Name: CryoEM map of Human Presequence Protease in partial close state 1 type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Presequence protease, mitochondrial
| Macromolecule | Name: Presequence protease, mitochondrial / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO EC number: Hydrolases; Acting on peptide bonds (peptidases); Metalloendopeptidases |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 114.901461 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MHHHHHHAAA CERALQYKLG DKIHGFTVNQ VTSVPELFLT AVKLTHDDTG ARYLHLARED TNNLFSVQFR TTPMDSTGVP HILEHTVLC GSQKYPCRDP FFKMLNRSLS TFMNAFTASD YTLYPFSTQN PKDFQNLLSV YLDATFFPCL RELDFWQEGW R LEHENPSD ...String: MHHHHHHAAA CERALQYKLG DKIHGFTVNQ VTSVPELFLT AVKLTHDDTG ARYLHLARED TNNLFSVQFR TTPMDSTGVP HILEHTVLC GSQKYPCRDP FFKMLNRSLS TFMNAFTASD YTLYPFSTQN PKDFQNLLSV YLDATFFPCL RELDFWQEGW R LEHENPSD PQTPLVFKGV VFNEMKGAFT DNERIFSQHL QNRLLPDHTY SVVSGGDPLC IPELTWEQLK QFHATHYHPS NA RFFTYGN FPLEQHLKQI HEEALSKFQK IEPSTVVPAQ TPWDKPREFQ ITCGPDSFAT DPSKQTTVSV SFLLPDITDT FEA FTLSLL SSLLTSGPNS PFYKALIESG LGTDFSPDVG YNGYTREAYF SVGLQGIVEK DIETVRSLID RTIDEVVEKG FEDD RIEAL LHKIEIQMKH QSTSFGLMLT SYIASCWNHD GDPVELLKLG NQLAKFRQCL QENPKFLQEK VKQYFKNNQH KLTLS MRPD DKYHEKQAQV EATKLKQKVE ALSPGDRQQI YEKGLELRSQ QSKPQDASCL PALKVSDIEP TIPVTELDVV LTAGDI PVQ YCAQPTNGMV YFRAFSSLNT LPEELRPYVP LFCSVLTKLG CGLLDYREQA QQIELKTGGM SASPHVLPDD SHMDTYE QG VLFSSLCLDR NLPDMMQLWS EIFNNPCFEE EEHFKVLVKM TAQELANGIP DSGHLYASIR AGRTLTPAGD LQETFSGM D QVRLMKRIAE MTDIKPILRK LPRIKKHLLN GDNMRCSVNA TPQQMPQTEK AVEDFLRSIG RSKKERRPVR PHTVEKPVP SSSGGDAHVP HGSQVIRKLV MEPTFKPWQM KTHFLMPFPV NYVGECIRTV PYTDPDHASL KILARLMTAK FLHTEIREKG GAYGGGAKL SHNGIFTLYS YRDPNTIETL QSFGKAVDWA KSGKFTQQDI DEAKLSVFST VDAPVAPSDK GMDHFLYGLS D EMKQAHRE QLFAVSHDKL LAVSDRYLGT GKSTHGLAIL GPENPKIAKD PSWIIR UniProtKB: Presequence protease, mitochondrial |
-Macromolecule #2: Amyloid-beta precursor protein
| Macromolecule | Name: Amyloid-beta precursor protein / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 4.335852 KDa |
| Sequence | String: DAEFRHDSGY EVHHQKLVFF AEDVGSNKGA IIGLMVGGVV UniProtKB: Amyloid-beta precursor protein |
-Macromolecule #3: Amyloid-beta precursor protein
| Macromolecule | Name: Amyloid-beta precursor protein / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 273.33 Da |
| Sequence | String: (UNK)(UNK)(UNK) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.7 Details: 20 mM Tris, pH 7.7, 150 mM NaCl, 10mM KCl, 20 mM EDTA and 1 mM 2-mercaptoethanol |
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 308 K / Instrument: SPOTITON |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average exposure time: 6.0 sec. / Average electron dose: 66.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.0 µm / Nominal defocus min: 1.2 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)