[English] 日本語
 Yorodumi
Yorodumi- EMDB-21653: Cryo-EM Structure of Hepatitis B virus T=4 capsid in complex with... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21653 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of Hepatitis B virus T=4 capsid in complex with the antiviral molecule DBT1 | |||||||||
 Map data Map data | HBV capsid in complex with the antiviral molecule DBT1. Icosahedral averaging imposed. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | capsid / antiviral / HBV / VIRUS LIKE PARTICLE | |||||||||
| Function / homology |  Function and homology information Function and homology informationmicrotubule-dependent intracellular transport of viral material towards nucleus / T=4 icosahedral viral capsid / viral penetration into host nucleus / host cell / host cell cytoplasm / symbiont entry into host cell / structural molecule activity / DNA binding / RNA binding / identical protein binding Similarity search - Function | |||||||||
| Biological species |  Hepatitis B virus genotype D subtype adw Hepatitis B virus genotype D subtype adw | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.6 Å | |||||||||
 Authors Authors | Schlicksup C / Laughlin P | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: ACS Chem Biol / Year: 2020 Journal: ACS Chem Biol / Year: 2020Title: Local Stabilization of Subunit-Subunit Contacts Causes Global Destabilization of Hepatitis B Virus Capsids. Authors: Christopher John Schlicksup / Patrick Laughlin / Steven Dunkelbarger / Joseph Che-Yen Wang / Adam Zlotnick /  Abstract: Development of antiviral molecules that bind virion is a strategy that remains in its infancy, and the details of their mechanisms are poorly understood. Here we investigate the behavior of DBT1, a ...Development of antiviral molecules that bind virion is a strategy that remains in its infancy, and the details of their mechanisms are poorly understood. Here we investigate the behavior of DBT1, a dibenzothiazepine that specifically interacts with the capsid protein of hepatitis B virus (HBV). We found that DBT1 stabilizes protein-protein interaction, accelerates capsid assembly, and can induce formation of aberrant particles. Paradoxically, DBT1 can cause preformed capsids to dissociate. These activities may lead to (i) assembly of empty and defective capsids, inhibiting formation of new virus, and (ii) disruption of mature viruses, which are metastable, to inhibit new infection. Using cryo-electron microscopy, we observed that DBT1 led to asymmetric capsids where well-defined DBT1 density was bound at all intersubunit contacts. These results suggest that DBT1 can support assembly by increasing buried surface area but induce disassembly of metastable capsids by favoring asymmetry to induce structural defects. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21653.map.gz emd_21653.map.gz | 190.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21653-v30.xml emd-21653-v30.xml emd-21653.xml emd-21653.xml | 13 KB 13 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21653_fsc.xml emd_21653_fsc.xml | 22.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_21653.png emd_21653.png | 100.6 KB | ||
| Masks |  emd_21653_msk_1.map emd_21653_msk_1.map | 1000 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-21653.cif.gz emd-21653.cif.gz | 5.8 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21653 http://ftp.pdbj.org/pub/emdb/structures/EMD-21653 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21653 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21653 | HTTPS FTP |
-Related structure data
| Related structure data |  6wfsMC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21653.map.gz / Format: CCP4 / Size: 1000 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21653.map.gz / Format: CCP4 / Size: 1000 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | HBV capsid in complex with the antiviral molecule DBT1. Icosahedral averaging imposed. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.65 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_21653_msk_1.map emd_21653_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Hepatitis B virus genotype D subtype adw
| Entire | Name:  Hepatitis B virus genotype D subtype adw Hepatitis B virus genotype D subtype adw |
|---|---|
| Components |
|
-Supramolecule #1: Hepatitis B virus genotype D subtype adw
| Supramolecule | Name: Hepatitis B virus genotype D subtype adw / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1 / NCBI-ID: 10419 / Sci species name: Hepatitis B virus genotype D subtype adw / Sci species strain: isolate United Kingdom/adyw/1979 / Virus type: VIRUS-LIKE PARTICLE / Virus isolate: OTHER / Virus enveloped: No / Virus empty: Yes |
|---|
-Macromolecule #1: Capsid protein
| Macromolecule | Name: Capsid protein / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Hepatitis B virus genotype D subtype adw Hepatitis B virus genotype D subtype adw |
| Molecular weight | Theoretical: 16.791104 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: MDIDPYKEFG ATVELLSFLP SDFFPSVRDL LDTAAALYRD ALESPEHASP HHTALRQAIL AWGDLMTLAT WVGTNLEDPA SRDLVVSYV NTNVGLKFRQ LLWFHISALT FGRETVLEYL VSFGVWIRTP PAYRPPNAPI LSTLPETTVV C UniProtKB: Capsid protein |
-Macromolecule #2: 11-oxo-N-[2-(4-sulfamoylphenyl)ethyl]-10,11-dihydrodibenzo[b,f][1...
| Macromolecule | Name: 11-oxo-N-[2-(4-sulfamoylphenyl)ethyl]-10,11-dihydrodibenzo[b,f][1,4]thiazepine-8-carboxamide type: ligand / ID: 2 / Number of copies: 4 / Formula: U0A |
|---|---|
| Molecular weight | Theoretical: 453.534 Da |
| Chemical component information | 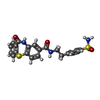 ChemComp-U0A: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 10 mg/mL | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| |||||||||
| Grid | Details: unspecified | |||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295.15 K / Instrument: FEI VITROBOT MARK III |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Number grids imaged: 1 / Number real images: 679 / Average electron dose: 33.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Initial model | PDB ID: Chain - Chain ID: A / Chain - Residue range: 1-143 / Chain - Source name: PDB / Chain - Initial model type: experimental model |
|---|---|
| Refinement | Space: REAL / Protocol: FLEXIBLE FIT / Target criteria: Correlation Coefficient |
| Output model |  PDB-6wfs: |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)