登録情報 データベース : EMDB / ID : EMD-20551タイトル Structure of a MAPK pathway complex Structure of a MAPK pathway complex 複合体 : ERK pathway complexタンパク質・ペプチド : Serine/threonine-protein kinase B-rafタンパク質・ペプチド : 14-3-3 protein zeta/delta / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Homo sapiens (ヒト)手法 / / 解像度 : 6.8 Å Park E / Rawson S 資金援助 Organization Grant number 国 National Institutes of Health/National Human Genome Research Institute (NIH/NHGRI) P50CA165962 National Institutes of Health/National Cancer Institute (NIH/NCI) R50CA221830
ジャーナル : Nature / 年 : 2019タイトル : Architecture of autoinhibited and active BRAF-MEK1-14-3-3 complexes.著者 : Eunyoung Park / Shaun Rawson / Kunhua Li / Byeong-Won Kim / Scott B Ficarro / Gonzalo Gonzalez-Del Pino / Humayun Sharif / Jarrod A Marto / Hyesung Jeon / Michael J Eck / 要旨 : RAF family kinases are RAS-activated switches that initiate signalling through the MAP kinase cascade to control cellular proliferation, differentiation and survival. RAF activity is tightly ... RAF family kinases are RAS-activated switches that initiate signalling through the MAP kinase cascade to control cellular proliferation, differentiation and survival. RAF activity is tightly regulated and inappropriate activation is a frequent cause of cancer; however, the structural basis for RAF regulation is poorly understood at present. Here we use cryo-electron microscopy to determine autoinhibited and active-state structures of full-length BRAF in complexes with MEK1 and a 14-3-3 dimer. The reconstruction reveals an inactive BRAF-MEK1 complex restrained in a cradle formed by the 14-3-3 dimer, which binds the phosphorylated S365 and S729 sites that flank the BRAF kinase domain. The BRAF cysteine-rich domain occupies a central position that stabilizes this assembly, but the adjacent RAS-binding domain is poorly ordered and peripheral. The 14-3-3 cradle maintains autoinhibition by sequestering the membrane-binding cysteine-rich domain and blocking dimerization of the BRAF kinase domain. In the active state, these inhibitory interactions are released and a single 14-3-3 dimer rearranges to bridge the C-terminal pS729 binding sites of two BRAFs, which drives the formation of an active, back-to-back BRAF dimer. Our structural snapshots provide a foundation for understanding normal RAF regulation and its mutational disruption in cancer and developmental syndromes. 履歴 登録 2019年8月1日 - ヘッダ(付随情報) 公開 2019年10月2日 - マップ公開 2019年10月9日 - 更新 2024年11月6日 - 現状 2024年11月6日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
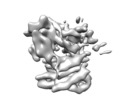
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Homo sapiens (ヒト)
Homo sapiens (ヒト) データ登録者
データ登録者 米国, 2件
米国, 2件  引用
引用 ジャーナル: Nature / 年: 2019
ジャーナル: Nature / 年: 2019
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_20551.map.gz
emd_20551.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-20551-v30.xml
emd-20551-v30.xml emd-20551.xml
emd-20551.xml EMDBヘッダ
EMDBヘッダ emd_20551.png
emd_20551.png emd-20551.cif.gz
emd-20551.cif.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-20551
http://ftp.pdbj.org/pub/emdb/structures/EMD-20551 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20551
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20551 emd_20551_validation.pdf.gz
emd_20551_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_20551_full_validation.pdf.gz
emd_20551_full_validation.pdf.gz emd_20551_validation.xml.gz
emd_20551_validation.xml.gz emd_20551_validation.cif.gz
emd_20551_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20551
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20551 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20551
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-20551 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_20551.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_20551.map.gz / 形式: CCP4 / 大きさ: 64 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) Homo sapiens (ヒト)
Homo sapiens (ヒト) 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 画像解析
画像解析
 ムービー
ムービー コントローラー
コントローラー





























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)























