[English] 日本語
 Yorodumi
Yorodumi- EMDB-16390: Transmembrane domain of active state homomeric GluA1 AMPA recepto... -
+ Open data
Open data
- Basic information
Basic information
| Entry |  | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Transmembrane domain of active state homomeric GluA1 AMPA receptor in tandem with TARP gamma 3 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | AMPAR / ion channels / neurotransmission / MEMBRANE PROTEIN | |||||||||
| Function / homology |  Function and homology information Function and homology informationCargo concentration in the ER / cellular response to amine stimulus / axonal spine / COPII-mediated vesicle transport / positive regulation of membrane potential / cellular response to ammonium ion / neurotransmitter receptor transport, postsynaptic endosome to lysosome / LGI-ADAM interactions / myosin V binding / neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration ...Cargo concentration in the ER / cellular response to amine stimulus / axonal spine / COPII-mediated vesicle transport / positive regulation of membrane potential / cellular response to ammonium ion / neurotransmitter receptor transport, postsynaptic endosome to lysosome / LGI-ADAM interactions / myosin V binding / neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration / Trafficking of AMPA receptors / neuron spine / regulation of AMPA receptor activity / neurotransmitter receptor internalization / response to arsenic-containing substance / cellular response to dsRNA / postsynaptic neurotransmitter receptor diffusion trapping / dendritic spine membrane / Synaptic adhesion-like molecules / long-term synaptic depression / cellular response to peptide hormone stimulus / neurotransmitter receptor localization to postsynaptic specialization membrane / protein kinase A binding / spinal cord development / neuronal cell body membrane / Activation of AMPA receptors / perisynaptic space / transmission of nerve impulse / AMPA glutamate receptor activity / channel regulator activity / Trafficking of GluR2-containing AMPA receptors / response to lithium ion / immunoglobulin binding / AMPA glutamate receptor complex / adenylate cyclase binding / cellular response to organic cyclic compound / ionotropic glutamate receptor complex / excitatory synapse / neuronal action potential / asymmetric synapse / G-protein alpha-subunit binding / regulation of receptor recycling / voltage-gated calcium channel activity / regulation of postsynaptic membrane potential / Unblocking of NMDA receptors, glutamate binding and activation / postsynaptic density, intracellular component / protein targeting / positive regulation of synaptic transmission / glutamate receptor binding / long-term memory / beta-2 adrenergic receptor binding / response to electrical stimulus / glutamate-gated calcium ion channel activity / glutamate-gated receptor activity / presynaptic active zone membrane / response to fungicide / somatodendritic compartment / synapse assembly / cellular response to brain-derived neurotrophic factor stimulus / dendrite membrane / ligand-gated monoatomic ion channel activity involved in regulation of presynaptic membrane potential / positive regulation of synaptic transmission, glutamatergic / response to cocaine / dendritic shaft / synaptic membrane / synaptic transmission, glutamatergic / PDZ domain binding / transmitter-gated monoatomic ion channel activity involved in regulation of postsynaptic membrane potential / long-term synaptic potentiation / cellular response to amino acid stimulus / postsynaptic density membrane / ionotropic glutamate receptor binding / modulation of chemical synaptic transmission / neuromuscular junction / regulation of synaptic plasticity / Schaffer collateral - CA1 synapse / response to organic cyclic compound / receptor internalization / recycling endosome / response to toxic substance / cellular response to growth factor stimulus / cerebral cortex development / response to peptide hormone / small GTPase binding / synaptic vesicle membrane / recycling endosome membrane / G-protein beta-subunit binding / protein localization / cell-cell junction / synaptic vesicle / response to estradiol / presynapse / presynaptic membrane / amyloid-beta binding / early endosome membrane / cell body / scaffold protein binding / chemical synaptic transmission / postsynaptic membrane / postsynapse Similarity search - Function | |||||||||
| Biological species |   | |||||||||
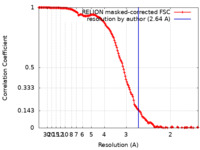
| Method | single particle reconstruction / cryo EM / Resolution: 2.64 Å | |||||||||
 Authors Authors | Zhang D / Ivica J / Krieger JM / Ho H / Yamashita K / Cais O / Greger I | |||||||||
| Funding support |  United Kingdom, 2 items United Kingdom, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2023 Journal: Nature / Year: 2023Title: Structural mobility tunes signalling of the GluA1 AMPA glutamate receptor. Authors: Danyang Zhang / Josip Ivica / James M Krieger / Hinze Ho / Keitaro Yamashita / Imogen Stockwell / Rozbeh Baradaran / Ondrej Cais / Ingo H Greger /   Abstract: AMPA glutamate receptors (AMPARs), the primary mediators of excitatory neurotransmission in the brain, are either GluA2 subunit-containing and thus Ca-impermeable, or GluA2-lacking and Ca-permeable. ...AMPA glutamate receptors (AMPARs), the primary mediators of excitatory neurotransmission in the brain, are either GluA2 subunit-containing and thus Ca-impermeable, or GluA2-lacking and Ca-permeable. Despite their prominent expression throughout interneurons and glia, their role in long-term potentiation and their involvement in a range of neuropathologies, structural information for GluA2-lacking receptors is currently absent. Here we determine and characterize cryo-electron microscopy structures of the GluA1 homotetramer, fully occupied with TARPγ3 auxiliary subunits (GluA1/γ3). The gating core of both resting and open-state GluA1/γ3 closely resembles GluA2-containing receptors. However, the sequence-diverse N-terminal domains (NTDs) give rise to a highly mobile assembly, enabling domain swapping and subunit re-alignments in the ligand-binding domain tier that are pronounced in desensitized states. These transitions underlie the unique kinetic properties of GluA1. A GluA2 mutant (F231A) increasing NTD dynamics phenocopies this behaviour, and exhibits reduced synaptic responses, reflecting the anchoring function of the AMPAR NTD at the synapse. Together, this work underscores how the subunit-diverse NTDs determine subunit arrangement, gating properties and ultimately synaptic signalling efficiency among AMPAR subtypes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_16390.map.gz emd_16390.map.gz | 14.6 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-16390-v30.xml emd-16390-v30.xml emd-16390.xml emd-16390.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_16390_fsc.xml emd_16390_fsc.xml | 14.7 KB | Display |  FSC data file FSC data file |
| Images |  emd_16390.png emd_16390.png | 49.2 KB | ||
| Masks |  emd_16390_msk_1.map emd_16390_msk_1.map | 109.3 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-16390.cif.gz emd-16390.cif.gz | 6.6 KB | ||
| Others |  emd_16390_half_map_1.map.gz emd_16390_half_map_1.map.gz emd_16390_half_map_2.map.gz emd_16390_half_map_2.map.gz | 100 MB 100 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-16390 http://ftp.pdbj.org/pub/emdb/structures/EMD-16390 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16390 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-16390 | HTTPS FTP |
-Related structure data
| Related structure data |  8c2hMC  8c1pC  8c1qC  8c1rC  8c1sC  8c2iC  8p3qC  8p3sC  8p3tC  8p3uC  8p3vC  8p3wC  8p3xC  8p3yC  8p3zC  8pivC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_16390.map.gz / Format: CCP4 / Size: 109.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_16390.map.gz / Format: CCP4 / Size: 109.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.826 Å | ||||||||||||||||||||
| Density |
| ||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Mask #1
| File |  emd_16390_msk_1.map emd_16390_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
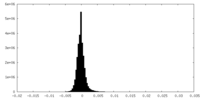
| Density Histograms |
-Half map: #2
| File | emd_16390_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
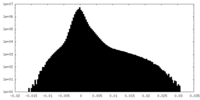
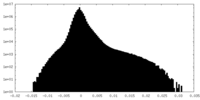
| Density Histograms |
-Half map: #1
| File | emd_16390_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
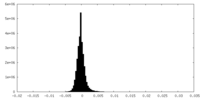
| Density Histograms |
- Sample components
Sample components
-Entire : Homomeric GluA1 AMPA receptor in complex with TARP gamma 3
| Entire | Name: Homomeric GluA1 AMPA receptor in complex with TARP gamma 3 |
|---|---|
| Components |
|
-Supramolecule #1: Homomeric GluA1 AMPA receptor in complex with TARP gamma 3
| Supramolecule | Name: Homomeric GluA1 AMPA receptor in complex with TARP gamma 3 type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Glutamate receptor 1 flip isoform
| Macromolecule | Name: Glutamate receptor 1 flip isoform / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 102.66193 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MPYIFAFFCT GFLGAVVGAD YKDDDDKNFP NNIQIGGLFP NQQSQEHAAF RFALSQLTEP PKLLPQIDIV NISDSFEMTY RFCSQFSKG VYAIFGFYER RTVNMLTSFC GALHVCFITP SFPVDTSNQF VLQLRPELQE ALISIIDHYK WQTFVYIYDA D RGLSVLQR ...String: MPYIFAFFCT GFLGAVVGAD YKDDDDKNFP NNIQIGGLFP NQQSQEHAAF RFALSQLTEP PKLLPQIDIV NISDSFEMTY RFCSQFSKG VYAIFGFYER RTVNMLTSFC GALHVCFITP SFPVDTSNQF VLQLRPELQE ALISIIDHYK WQTFVYIYDA D RGLSVLQR VLDTAAEKNW QVTAVNILTT TEEGYRMLFQ DLEKKKERLV VVDCESERLN AILGQIVKLE KNGIGYHYIL AN LGFMDID LNKFKESGAN VTGFQLVNYT DTIPARIMQQ WRTSDSRDHT RVDWKRPKYT SALTYDGVKV MAEAFQSLRR QRI DISRRG NAGDCLANPA VPWGQGIDIQ RALQQVRFEG LTGNVQFNEK GRRTNYTLHV IEMKHDGIRK IGYWNEDDKF VPAA TDAQA GGDNSSVQNR TYIVTTILED PYVMLKKNAN QFEGNDRYEG YCVELAAEIA KHVGYSYRLE IVSDGKYGAR DPDTK AWNG MVGELVYGRA DVAVAPLTIT LVREEVIDFS KPFMSLGISI MIKKPQKSKP GVFSFLDPLA YEIWMCIVFA YIGVSV VLF LVSRFSPYEW HSEEFEEGRD QTTSDQSNEF GIFNSLWFSL GAFMQQGCDI SPRSLSGRIV GGVWWFFTLI IISSYTA NL AAFLTVERMV SPIESAEDLA KQTEIAYGTL EAGSTKEFFR RSKIAVFEKM WTYMKSAEPS VFVRTTEEGM IRVRKSKG K YAYLLESTMN EYIEQRKPCD TMKVGGNLDS KGYGIATPKG SALRGPVNLA VLKLSEQGVL DKLKSKWWYD KGECGSKDS GSKDKTSALS LSNVAGVFYI LIGGLGLAML VALIEFCYKS RSESKRMKGF CLIPQQSINE AIRTSTLPRN SGAGASGGGG SGENGRVVS QDFPKSMQSI PCMSHSSGMP LGATGL UniProtKB: Glutamate receptor 1 |
-Macromolecule #2: Voltage-dependent calcium channel gamma-3 subunit
| Macromolecule | Name: Voltage-dependent calcium channel gamma-3 subunit / type: protein_or_peptide / ID: 2 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 35.435332 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: RMCDRGIQML ITTVGAFAAF SLMTIAVGTD YWLYSRGVCR TKSTSDNETS RKNEEVMTHS GLWRTCCLEG AFRGVCKKID HFPEDADYE QDTAEYLLRA VRASSVFPIL SVTLLFFGGL CVAASEFHRS RHSVILSAGI FFVSAGLSNI IGIIVYISAN A GDPGQRDS ...String: RMCDRGIQML ITTVGAFAAF SLMTIAVGTD YWLYSRGVCR TKSTSDNETS RKNEEVMTHS GLWRTCCLEG AFRGVCKKID HFPEDADYE QDTAEYLLRA VRASSVFPIL SVTLLFFGGL CVAASEFHRS RHSVILSAGI FFVSAGLSNI IGIIVYISAN A GDPGQRDS KKSYSYGWSF YFGAFSFIIA EIVGVVAVHI YIEKHQQLRA RSHSELLKKS TFARLPPYRY RFRRRSSSRS TE PRSRDLS PISKGFHTIP STDISMFTLS RDPSKLTMGT LLNSDRDHAF LQFHNSTPKE FKESLHNNPA NRRTTPV UniProtKB: Voltage-dependent calcium channel gamma-3 subunit |
-Macromolecule #3: PALMITIC ACID
| Macromolecule | Name: PALMITIC ACID / type: ligand / ID: 3 / Number of copies: 6 / Formula: PLM |
|---|---|
| Molecular weight | Theoretical: 256.424 Da |
| Chemical component information |  ChemComp-PLM: |
-Macromolecule #4: (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate
| Macromolecule | Name: (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate / type: ligand / ID: 4 / Number of copies: 4 / Formula: OLC |
|---|---|
| Molecular weight | Theoretical: 356.54 Da |
| Chemical component information |  ChemComp-OLC: |
-Macromolecule #5: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(tri...
| Macromolecule | Name: (2S)-3-(hexadecanoyloxy)-2-[(9Z)-octadec-9-enoyloxy]propyl 2-(trimethylammonio)ethyl phosphate type: ligand / ID: 5 / Number of copies: 2 / Formula: POV |
|---|---|
| Molecular weight | Theoretical: 760.076 Da |
| Chemical component information |  ChemComp-POV: |
-Macromolecule #6: water
| Macromolecule | Name: water / type: ligand / ID: 6 / Number of copies: 86 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | TFS KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 1.4000000000000001 µm / Nominal defocus min: 1.4000000000000001 µm |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | Servalcat |
|---|---|
| Refinement | Space: RECIPROCAL |
| Output model |  PDB-8c2h: |
 Movie
Movie Controller
Controller





















 Z
Z Y
Y X
X