+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-3283 | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
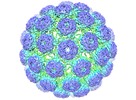
| Title | Cryo-EM structure of BK polyomavirus | ||||||||||||
 Map data Map data | Reconstruction of BK polyomavirus (sharpened/masked) | ||||||||||||
 Sample Sample |
| ||||||||||||
 Keywords Keywords | BKPyV / BK / polyomavirus | ||||||||||||
| Function / homology |  Function and homology information Function and homology informationcaveolin-mediated endocytosis of virus by host cell / T=7 icosahedral viral capsid / virion attachment to host cell / host cell nucleus / structural molecule activity Similarity search - Function | ||||||||||||
| Biological species |  BK polyomavirus BK polyomavirus | ||||||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 7.6 Å | ||||||||||||
 Authors Authors | Hurdiss DL / Morgan EL / Thompson RF / Prescott EL / Panou MM / Macdonald A / Ranson NA | ||||||||||||
 Citation Citation |  Journal: Structure / Year: 2016 Journal: Structure / Year: 2016Title: New Structural Insights into the Genome and Minor Capsid Proteins of BK Polyomavirus using Cryo-Electron Microscopy. Authors: Daniel L Hurdiss / Ethan L Morgan / Rebecca F Thompson / Emma L Prescott / Margarita M Panou / Andrew Macdonald / Neil A Ranson /  Abstract: BK polyomavirus is the causative agent of several diseases in transplant patients and the immunosuppressed. In order to better understand the structure and life cycle of BK, we produced infectious ...BK polyomavirus is the causative agent of several diseases in transplant patients and the immunosuppressed. In order to better understand the structure and life cycle of BK, we produced infectious virions and VP1-only virus-like particles in cell culture, and determined their three-dimensional structures using cryo-electron microscopy (EM) and single-particle image processing. The resulting 7.6-Å resolution structure of BK and 9.1-Å resolution of the virus-like particles are the highest-resolution cryo-EM structures of any polyomavirus. These structures confirm that the architecture of the major structural protein components of these human polyomaviruses are similar to previous structures from other hosts, but give new insight into the location and role of the enigmatic minor structural proteins, VP2 and VP3. We also observe two shells of electron density, which we attribute to a structurally ordered part of the viral genome, and discrete contacts between this density and both VP1 and the minor capsid proteins. | ||||||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_3283.map.gz emd_3283.map.gz | 35.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-3283-v30.xml emd-3283-v30.xml emd-3283.xml emd-3283.xml | 8.8 KB 8.8 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_3283_fsc.xml emd_3283_fsc.xml | 13.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_3283.tif emd_3283.tif | 2.4 MB | ||
| Others |  emd_3283_additional_1.map emd_3283_additional_1.map | 216 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-3283 http://ftp.pdbj.org/pub/emdb/structures/EMD-3283 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3283 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-3283 | HTTPS FTP |
-Related structure data
| Related structure data |  5fuaMC  3284C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_3283.map.gz / Format: CCP4 / Size: 210.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_3283.map.gz / Format: CCP4 / Size: 210.9 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Reconstruction of BK polyomavirus (sharpened/masked) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.92 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
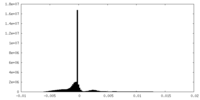
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Supplemental map: emd 3283 additional 1.map
| File | emd_3283_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
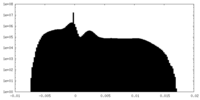
| Density Histograms |
- Sample components
Sample components
-Entire : BK polyomavirus
| Entire | Name:  BK polyomavirus BK polyomavirus |
|---|---|
| Components |
|
-Supramolecule #1000: BK polyomavirus
| Supramolecule | Name: BK polyomavirus / type: sample / ID: 1000 / Oligomeric state: Icosohedral / Number unique components: 1 |
|---|
-Supramolecule #1: BK polyomavirus
| Supramolecule | Name: BK polyomavirus / type: virus / ID: 1 / NCBI-ID: 10629 / Sci species name: BK polyomavirus / Sci species strain: Dunlop / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host (natural) | Organism:  Homo sapiens (human) / synonym: VERTEBRATES Homo sapiens (human) / synonym: VERTEBRATES |
| Host system | Recombinant cell: Vero cells |
| Virus shell | Shell ID: 1 / Diameter: 498 Å |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.9 / Details: 10mM HEPES pH 7.9, 1mM CaCl2, 1mM MgCl2, 5mM KCl |
|---|---|
| Grid | Details: Quantifoil R2/1 EM grids |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Instrument: FEI VITROBOT MARK IV / Method: 6.5 seconds blot before plunging |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI F20 |
|---|---|
| Date | Dec 15, 2014 |
| Image recording | Category: CCD / Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Number real images: 432 / Average electron dose: 40 e/Å2 Details: 4 e-/A2/s, a 4 frames per second frame rate, and a 10 s exposure |
| Tilt angle min | 0 |
| Tilt angle max | 0 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Nominal defocus max: 4.8 µm / Nominal defocus min: 0.6 µm / Nominal magnification: 19000 |
| Sample stage | Specimen holder model: GATAN LIQUID NITROGEN |
| Experimental equipment |  Model: Tecnai F20 / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)