[English] 日本語
 Yorodumi
Yorodumi- EMDB-32209: Coxsackievirus B3 at pH7.4 (VP3-234E) incubation with coxsackievi... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-32209 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Coxsackievirus B3 at pH7.4 (VP3-234E) incubation with coxsackievirus and adenovirus receptor for 10min | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationAV node cell-bundle of His cell adhesion involved in cell communication / cell adhesive protein binding involved in AV node cell-bundle of His cell communication / homotypic cell-cell adhesion / AV node cell to bundle of His cell communication / epithelial structure maintenance / regulation of AV node cell action potential / gamma-delta T cell activation / germ cell migration / apicolateral plasma membrane / transepithelial transport ...AV node cell-bundle of His cell adhesion involved in cell communication / cell adhesive protein binding involved in AV node cell-bundle of His cell communication / homotypic cell-cell adhesion / AV node cell to bundle of His cell communication / epithelial structure maintenance / regulation of AV node cell action potential / gamma-delta T cell activation / germ cell migration / apicolateral plasma membrane / transepithelial transport / cell-cell junction organization / connexin binding / cardiac muscle cell development / heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules / intercalated disc / bicellular tight junction / cell adhesion molecule binding / neutrophil chemotaxis / acrosomal vesicle / filopodium / mitochondrion organization / PDZ domain binding / Cell surface interactions at the vascular wall / adherens junction / neuromuscular junction / beta-catenin binding / Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell / cell-cell junction / integrin binding / cell junction / heart development / virus receptor activity / growth cone / cell body / actin cytoskeleton organization / basolateral plasma membrane / defense response to virus / neuron projection / membrane raft / signaling receptor binding / protein-containing complex / extracellular space / extracellular region / nucleoplasm / identical protein binding / plasma membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |   Coxsackievirus B3 / Coxsackievirus B3 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
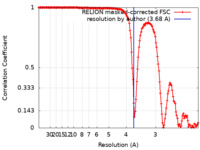
| Method | single particle reconstruction / cryo EM / Resolution: 3.68 Å | |||||||||
 Authors Authors | Wang QL / Liu CC | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2022 Journal: Proc Natl Acad Sci U S A / Year: 2022Title: Molecular basis of differential receptor usage for naturally occurring CD55-binding and -nonbinding coxsackievirus B3 strains. Authors: Qingling Wang / Qian Yang / Congcong Liu / Guoqing Wang / Hao Song / Guijun Shang / Ruchao Peng / Xiao Qu / Sheng Liu / Yingzi Cui / Peiyi Wang / Wenbo Xu / Xin Zhao / Jianxun Qi / Mengsu Yang / George F Gao /  Abstract: Receptor usage defines cell tropism and contributes to cell entry and infection. Coxsackievirus B (CVB) engages coxsackievirus and adenovirus receptor (CAR), and selectively utilizes the decay- ...Receptor usage defines cell tropism and contributes to cell entry and infection. Coxsackievirus B (CVB) engages coxsackievirus and adenovirus receptor (CAR), and selectively utilizes the decay-accelerating factor (DAF; CD55) to infect cells. However, the differential receptor usage mechanism for CVB remains elusive. This study identified VP3-234 residues (234Q/N/V/D/E) as critical population selection determinants during CVB3 virus evolution, contributing to diverse binding affinities to CD55. Cryoelectron microscopy (cryo-EM) structures of CD55-binding/nonbinding isolates and their complexes with CD55 or CAR were obtained under both neutral and acidic conditions, and the molecular mechanism of VP3-234 residues determining CD55 affinity/specificity for naturally occurring CVB3 strains was elucidated. Structural and biochemical studies in vitro revealed the dynamic entry process of CVB3 and the function of the uncoating receptor CAR with different pH preferences. This work provides detailed insight into the molecular mechanism of CVB infection and contributes to an in-depth understanding of enterovirus attachment receptor usage. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_32209.map.gz emd_32209.map.gz | 76.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-32209-v30.xml emd-32209-v30.xml emd-32209.xml emd-32209.xml | 21.5 KB 21.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_32209_fsc.xml emd_32209_fsc.xml | 15.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_32209.png emd_32209.png | 108.5 KB | ||
| Masks |  emd_32209_msk_1.map emd_32209_msk_1.map | 307.5 MB |  Mask map Mask map | |
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-32209 http://ftp.pdbj.org/pub/emdb/structures/EMD-32209 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32209 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-32209 | HTTPS FTP |
-Validation report
| Summary document |  emd_32209_validation.pdf.gz emd_32209_validation.pdf.gz | 532.5 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_32209_full_validation.pdf.gz emd_32209_full_validation.pdf.gz | 532.1 KB | Display | |
| Data in XML |  emd_32209_validation.xml.gz emd_32209_validation.xml.gz | 14.5 KB | Display | |
| Data in CIF |  emd_32209_validation.cif.gz emd_32209_validation.cif.gz | 19.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32209 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32209 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32209 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-32209 | HTTPS FTP |
-Related structure data
| Related structure data |  7vymMC  7vxhC  7vxzC  7vy0C  7vy5C  7vy6C  7vykC  7vylC  7w14C  7w17C M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_32209.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_32209.map.gz / Format: CCP4 / Size: 307.5 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_32209_msk_1.map emd_32209_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Coxsackievirus B3
| Entire | Name:   Coxsackievirus B3 Coxsackievirus B3 |
|---|---|
| Components |
|
-Supramolecule #1: Coxsackievirus B3
| Supramolecule | Name: Coxsackievirus B3 / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:   Coxsackievirus B3 Coxsackievirus B3 |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Supramolecule #3: Coxsackievirus and adenovirus receptor
| Supramolecule | Name: Coxsackievirus and adenovirus receptor / type: complex / ID: 3 / Parent: 1 / Macromolecule list: #5 |
|---|
-Supramolecule #2: Homo sapiens
| Supramolecule | Name: Homo sapiens / type: virus / ID: 2 / Parent: 1 / Macromolecule list: #1-#4 / NCBI-ID: 9606 / Sci species name: Homo sapiens / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|---|
| Host system | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus B3 Coxsackievirus B3 |
| Molecular weight | Theoretical: 31.561285 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GPVEDAVTAA IGRVADTVGT GPTNSEAIPA LTAAETGHTS QVVPGDTMQT RHVKNYHSRS ESTVENFLCR SACVYFTEYK NSGSKRYAE WVVTTRQAAQ LRRKLEFFTY IRFDLELTFV ITSTQQPSTT QNQDAQILTH QIMYVPPGGP VPDKVDSYVW Q TSTNPSVF ...String: GPVEDAVTAA IGRVADTVGT GPTNSEAIPA LTAAETGHTS QVVPGDTMQT RHVKNYHSRS ESTVENFLCR SACVYFTEYK NSGSKRYAE WVVTTRQAAQ LRRKLEFFTY IRFDLELTFV ITSTQQPSTT QNQDAQILTH QIMYVPPGGP VPDKVDSYVW Q TSTNPSVF WTEGNAPPRM SIPFLSIGNA YSNFYDGWSD FSRDGVYGIN TLNSMGTLYA RHVNTGGTGP IKSTIRIYFK PK HVKAWIP RPPRLCQYEK AKNVNFQPSG VTTTRQSITA MTNTGAF |
-Macromolecule #2: Capsid protein VP2
| Macromolecule | Name: Capsid protein VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus B3 Coxsackievirus B3 |
| Molecular weight | Theoretical: 28.85649 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: SPTVEECGYS DRVRSITLGN STITTQECAN VVVGYGVWPD YLKDNEATAE DQPTQPDVAT CRFYTLDSVQ WQKTSPGWWW KLPDALSNL GLFGQNMQYH YLGRTGYTIH VQCNASKFHQ GCLLVVCVPE AEMGCATLDN TPSSAELLGG DAAKEFAGEP I ASGSNKLV ...String: SPTVEECGYS DRVRSITLGN STITTQECAN VVVGYGVWPD YLKDNEATAE DQPTQPDVAT CRFYTLDSVQ WQKTSPGWWW KLPDALSNL GLFGQNMQYH YLGRTGYTIH VQCNASKFHQ GCLLVVCVPE AEMGCATLDN TPSSAELLGG DAAKEFAGEP I ASGSNKLV QRVVYNAGMG IGVGNLTIFP HQWINLRTNN SATIVMPYTN SVPMDNMFRH NNVTLMVIPF VPLDYCPGST TY VPITVTI APMNAEYNGL RLAGHQ |
-Macromolecule #3: Capsid protein VP3
| Macromolecule | Name: Capsid protein VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus B3 Coxsackievirus B3 |
| Molecular weight | Theoretical: 26.154695 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GLPTMNTPGS CQFLTSDDFQ SPSAMPQYDV TPEMRIPGEV KNLMEIAEVD SVVPVQNVGE KVNSMEAYQI PVRSNEGSGT QVFGFPLQP GYSSVFSRTL LGEILNYYTH WSGSIKLTFM FCGSAMATGK FLLAYSPLGA GAPTKRVDAM LGTHVVWDVG L QSSCVLCI ...String: GLPTMNTPGS CQFLTSDDFQ SPSAMPQYDV TPEMRIPGEV KNLMEIAEVD SVVPVQNVGE KVNSMEAYQI PVRSNEGSGT QVFGFPLQP GYSSVFSRTL LGEILNYYTH WSGSIKLTFM FCGSAMATGK FLLAYSPLGA GAPTKRVDAM LGTHVVWDVG L QSSCVLCI PWISQTHYRY VASDECTAGG FITCWYQTNI VVPADAQSSC YIMCFVSACN DFSVRLLKDT PFISQENFFQ |
-Macromolecule #4: Capsid protein VP4
| Macromolecule | Name: Capsid protein VP4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus B3 Coxsackievirus B3 |
| Molecular weight | Theoretical: 7.306014 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: GAQVSTQKTG AHETGLNASG NSIIHYTNIN YYKDAASNSA TRQDFAQDPG KFTEPVKDIM IKSLPALN |
-Macromolecule #5: Coxsackievirus and adenovirus receptor
| Macromolecule | Name: Coxsackievirus and adenovirus receptor / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 25.106408 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSITTPEEMI EKAKGETAYL PCKFTLSPED QGPLDIEWLI SPADNQKVDQ VIILYSGDKI YDDYYPDLKG RVHFTSNDLK SGDASINVT NLQLSDIGTY QCKVKKAPGV ANKKIHLVVL VKPSGARCYV DGSEEIGSDF KIKCEPKEGS LPLQYEWQKL S DSQKMPTS ...String: MSITTPEEMI EKAKGETAYL PCKFTLSPED QGPLDIEWLI SPADNQKVDQ VIILYSGDKI YDDYYPDLKG RVHFTSNDLK SGDASINVT NLQLSDIGTY QCKVKKAPGV ANKKIHLVVL VKPSGARCYV DGSEEIGSDF KIKCEPKEGS LPLQYEWQKL S DSQKMPTS WLAEMTSSVI SVKNASSEYS GTYSCTVRNR VGSDQCLLRL NVVPPSNKAL EHHHHHH |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Grid | Model: PELCO Ultrathin Carbon with Lacey Carbon / Support film - Material: CARBON / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Temperature | Min: 70.0 K / Max: 70.0 K |
| Image recording | Film or detector model: FEI FALCON III (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 4096 pixel / Digitization - Dimensions - Height: 4096 pixel / Average exposure time: 1.0 sec. / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated defocus max: 5.0 µm / Calibrated defocus min: 1.8 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.5 µm / Nominal defocus min: 1.5 µm / Nominal magnification: 75000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)