[English] 日本語
 Yorodumi
Yorodumi- EMDB-30260: Cryo-EM structure of mature Coxsackievirus A10 in complex with KR... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-30260 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of mature Coxsackievirus A10 in complex with KRM1 at pH 5.5 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Picornavirus / Coxsackievirus A10 / pH 5.5 / mature particle / KRM1 / complex / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationSignaling by LRP5 mutants / Negative regulation of TCF-dependent signaling by WNT ligand antagonists / cell communication / negative regulation of axon regeneration / negative regulation of ossification / regulation of canonical Wnt signaling pathway / limb development / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / TCF dependent signaling in response to WNT / picornain 2A ...Signaling by LRP5 mutants / Negative regulation of TCF-dependent signaling by WNT ligand antagonists / cell communication / negative regulation of axon regeneration / negative regulation of ossification / regulation of canonical Wnt signaling pathway / limb development / symbiont-mediated suppression of host cytoplasmic pattern recognition receptor signaling pathway via inhibition of MDA-5 activity / TCF dependent signaling in response to WNT / picornain 2A / symbiont-mediated suppression of host mRNA export from nucleus / symbiont genome entry into host cell via pore formation in plasma membrane / picornain 3C / T=pseudo3 icosahedral viral capsid / negative regulation of canonical Wnt signaling pathway / host cell cytoplasmic vesicle membrane / Wnt signaling pathway / ribonucleoside triphosphate phosphatase activity / transmembrane signaling receptor activity / nucleoside-triphosphate phosphatase / channel activity / monoatomic ion transmembrane transport / DNA replication / RNA helicase activity / endocytosis involved in viral entry into host cell / symbiont-mediated activation of host autophagy / RNA-directed RNA polymerase / cysteine-type endopeptidase activity / viral RNA genome replication / RNA-directed RNA polymerase activity / neuronal cell body / apoptotic process / DNA-templated transcription / virion attachment to host cell / host cell nucleus / structural molecule activity / signal transduction / proteolysis / RNA binding / zinc ion binding / ATP binding / membrane / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Coxsackievirus A10 / Coxsackievirus A10 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.0 Å | |||||||||
 Authors Authors | Cui Y / Peng R / Gao GF / Qi J | |||||||||
| Funding support |  China, 1 items China, 1 items
| |||||||||
 Citation Citation |  Journal: Proc Natl Acad Sci U S A / Year: 2020 Journal: Proc Natl Acad Sci U S A / Year: 2020Title: Molecular basis of Coxsackievirus A10 entry using the two-in-one attachment and uncoating receptor KRM1. Authors: Yingzi Cui / Ruchao Peng / Hao Song / Zhou Tong / Xiao Qu / Sheng Liu / Xin Zhao / Yan Chai / Peiyi Wang / George F Gao / Jianxun Qi /  Abstract: KREMEN1 (KRM1) has been identified as a functional receptor for Coxsackievirus A10 (CV-A10), a causative agent of hand-foot-and-mouth disease (HFMD), which poses a great threat to infants globally. ...KREMEN1 (KRM1) has been identified as a functional receptor for Coxsackievirus A10 (CV-A10), a causative agent of hand-foot-and-mouth disease (HFMD), which poses a great threat to infants globally. However, the underlying mechanisms for the viral entry process are not well understood. Here we determined the atomic structures of different forms of CV-A10 viral particles and its complex with KRM1 in both neutral and acidic conditions. These structures reveal that KRM1 selectively binds to the mature viral particle above the canyon of the viral protein 1 (VP1) subunit and contacts across two adjacent asymmetry units. The key residues for receptor binding are conserved among most KRM1-dependent enteroviruses, suggesting a uniform mechanism for receptor binding. Moreover, the binding of KRM1 induces the release of pocket factor, a process accelerated under acidic conditions. Further biochemical studies confirmed that receptor binding at acidic pH enabled CV-A10 virion uncoating in vitro. Taken together, these findings provide high-resolution snapshots of CV-A10 entry and identify KRM1 as a two-in-one receptor for enterovirus infection. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_30260.map.gz emd_30260.map.gz | 94.8 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-30260-v30.xml emd-30260-v30.xml emd-30260.xml emd-30260.xml | 19.2 KB 19.2 KB | Display Display |  EMDB header EMDB header |
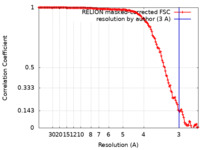
| FSC (resolution estimation) |  emd_30260_fsc.xml emd_30260_fsc.xml | 14.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_30260.png emd_30260.png | 289.4 KB | ||
| Filedesc metadata |  emd-30260.cif.gz emd-30260.cif.gz | 7.1 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-30260 http://ftp.pdbj.org/pub/emdb/structures/EMD-30260 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30260 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-30260 | HTTPS FTP |
-Related structure data
| Related structure data |  7bzuMC  7bznC  7bzoC  7bztC  7c4tC  7c4wC  7c4yC  7c4zC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_30260.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_30260.map.gz / Format: CCP4 / Size: 244.1 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.32 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Coxsackievirus A10
| Entire | Name:   Coxsackievirus A10 Coxsackievirus A10 |
|---|---|
| Components |
|
-Supramolecule #1: Coxsackievirus A10
| Supramolecule | Name: Coxsackievirus A10 / type: virus / ID: 1 / Parent: 0 / Macromolecule list: #1-#5 / NCBI-ID: 42769 / Sci species name: Coxsackievirus A10 / Virus type: VIRION / Virus isolate: STRAIN / Virus enveloped: No / Virus empty: No |
|---|
-Macromolecule #1: Capsid protein VP1
| Macromolecule | Name: Capsid protein VP1 / type: protein_or_peptide / ID: 1 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A10 Coxsackievirus A10 |
| Molecular weight | Theoretical: 33.204332 KDa |
| Sequence | String: GDPVEDIIHD ALGSTARRAI SSVTNAESAA NTTPSSHRLE TGRVPALQAA ETGATSNATD ENMIETRCVV NRNGVLETTI NHFFSRSGL VGVVNLTDGG TDTTGYATWD IDIMGFVQLR RKCEMFTYMR FNAEFTFVTT TKNGEARPYM LQYMYVPPGA P KPTGRDAF ...String: GDPVEDIIHD ALGSTARRAI SSVTNAESAA NTTPSSHRLE TGRVPALQAA ETGATSNATD ENMIETRCVV NRNGVLETTI NHFFSRSGL VGVVNLTDGG TDTTGYATWD IDIMGFVQLR RKCEMFTYMR FNAEFTFVTT TKNGEARPYM LQYMYVPPGA P KPTGRDAF QWQTATNPSV FVKLTDPPAQ VSVPFMSPAS AYQWFYDGYP TFGQHPETSN TTYGLCPNNM MGTFAVRVVS RE ASQLKLQ TRVYMKLKHV RAWVPRPIRS QPYLLKNFPN YDSSKITNSA RDRSSIKQAN M |
-Macromolecule #2: Capsid protein VP2
| Macromolecule | Name: Capsid protein VP2 / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A10 Coxsackievirus A10 |
| Molecular weight | Theoretical: 27.783105 KDa |
| Sequence | String: SPSVEACGYS DRVAQLTVGN SSITTQEAAN IVLAYGEWPE YCPDTDATAV DKPTRPDVSV NRFYTLDSKM WQENSTGWYW KFPDVLNKT GVFGQNAQFH YLYRSGFCLH VQCNASKFHQ GALLVAVIPE FVIAGRGSNT KPNEAPHPGF TTTFPGTTGA T FHDPYVLD ...String: SPSVEACGYS DRVAQLTVGN SSITTQEAAN IVLAYGEWPE YCPDTDATAV DKPTRPDVSV NRFYTLDSKM WQENSTGWYW KFPDVLNKT GVFGQNAQFH YLYRSGFCLH VQCNASKFHQ GALLVAVIPE FVIAGRGSNT KPNEAPHPGF TTTFPGTTGA T FHDPYVLD SGVPLSQALI YPHQWINLRT NNCATVIVPY INAVPFDSAI NHSNFGLIVI PVSPLKYSSG ATTAIPITIT IA PLNSEFG GLRQAVSQ UniProtKB: Genome polyprotein |
-Macromolecule #3: Capsid protein VP3
| Macromolecule | Name: Capsid protein VP3 / type: protein_or_peptide / ID: 3 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A10 Coxsackievirus A10 |
| Molecular weight | Theoretical: 26.187623 KDa |
| Sequence | String: GIPAELRPGT NQFLTTDDDT AAPILPGFTP TPTIHIPGEV HSLLELCRVE TILEVNNTTE ATGLTRLLIP VSSQNKADEL CAAFMVDPG RIGPWQSTLV GQICRYYTQW SGSLKVTFMF TGSFMATGKM LVAYSPPGSA QPANRETAML GTHVIWDFGL Q SSVSLVIP ...String: GIPAELRPGT NQFLTTDDDT AAPILPGFTP TPTIHIPGEV HSLLELCRVE TILEVNNTTE ATGLTRLLIP VSSQNKADEL CAAFMVDPG RIGPWQSTLV GQICRYYTQW SGSLKVTFMF TGSFMATGKM LVAYSPPGSA QPANRETAML GTHVIWDFGL Q SSVSLVIP WISNTHFRTA KTGGNYDYYT AGVVTLWYQT NYVVPPETPG EAYIIAMGAA QDNFTLKICK DTDEVTQQAV LQ UniProtKB: Genome polyprotein |
-Macromolecule #4: Capsid protein VP4
| Macromolecule | Name: Capsid protein VP4 / type: protein_or_peptide / ID: 4 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Coxsackievirus A10 Coxsackievirus A10 |
| Molecular weight | Theoretical: 7.464104 KDa |
| Sequence | String: MGAQVSTQKS GSHETGNVAT GGSTINFTNI NYYKDSYAAS ATRQDFTQDP KKFTQPVLDS IRELSAPLN UniProtKB: Genome polyprotein |
-Macromolecule #5: KRM1
| Macromolecule | Name: KRM1 / type: protein_or_peptide / ID: 5 / Number of copies: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 41.499027 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: APSPGLGPGP ECFTANGADY RGTQNWTALQ GGKPCLFWNE TFQHPYNTLK YPNGEGGLGE HNYCRNPDGD VSPWCYVAEH EDGVYWKYC EIPACQMPGN LGCYKDHGNP PPLTGTSKTS NKLTIQTCIS FCRSQRFKFA GMESGYACFC GNNPDYWKYG E AASTECNS ...String: APSPGLGPGP ECFTANGADY RGTQNWTALQ GGKPCLFWNE TFQHPYNTLK YPNGEGGLGE HNYCRNPDGD VSPWCYVAEH EDGVYWKYC EIPACQMPGN LGCYKDHGNP PPLTGTSKTS NKLTIQTCIS FCRSQRFKFA GMESGYACFC GNNPDYWKYG E AASTECNS VCFGDHTQPC GGDGRIILFD TLVGACGGNY SAMSSVVYSP DFPDTYATGR VCYWTIRVPG ASHIHFSFPL FD IRDSADM VELLDGYTHR VLARFHGRSR PPLSFNVSLD FVILYFFSDR INQAQGFAVL YQAVKEEGSE NLYFQGGSLP QER PAVNQT VAEVITEQAN LSVSAARSSK VLYVITTSPS HPPQTVPGTH HHHHHHHHH |
-Macromolecule #6: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 6 / Number of copies: 3 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 5.5 |
|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 40.0 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)