登録情報 データベース : EMDB / ID : EMD-24729タイトル CryoEM structure of RBD domain of COVID-19 in complex with Legobody sharpened map for the complex 複合体 : The complex of RBD domain of COVID-19 with Legobodyタンパク質・ペプチド : Maltodextrin-binding protein,Immunoglobulin G-binding protein A,Immunoglobulin G-binding protein Gタンパク質・ペプチド : Fab_8D3_2 heavy chainタンパク質・ペプチド : Fab_8D3_2 light chainタンパク質・ペプチド : Nb_RBDタンパク質・ペプチド : Spike protein S1 / / / / 機能・相同性 分子機能 ドメイン・相同性 構成要素
/ / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / / 生物種 Streptococcus sp. (バクテリア) / Mus musculus (ハツカネズミ) / Vicugna pacos (アルパカ) / 手法 / / 解像度 : 3.6 Å Wu XD / Rapoport TA 資金援助 Organization Grant number 国 National Institutes of Health/National Institute of General Medical Sciences (NIH/NIGMS) R01GM052586
ジャーナル : Proc Natl Acad Sci U S A / 年 : 2021タイトル : Cryo-EM structure determination of small proteins by nanobody-binding scaffolds (Legobodies).著者 : Xudong Wu / Tom A Rapoport / 要旨 : We describe a general method that allows structure determination of small proteins by single-particle cryo-electron microscopy (cryo-EM). The method is based on the availability of a target-binding ... We describe a general method that allows structure determination of small proteins by single-particle cryo-electron microscopy (cryo-EM). The method is based on the availability of a target-binding nanobody, which is then rigidly attached to two scaffolds: 1) a Fab fragment of an antibody directed against the nanobody and 2) a nanobody-binding protein A fragment fused to maltose binding protein and Fab-binding domains. The overall ensemble of ∼120 kDa, called Legobody, does not perturb the nanobody-target interaction, is easily recognizable in EM images due to its unique shape, and facilitates particle alignment in cryo-EM image processing. The utility of the method is demonstrated for the KDEL receptor, a 23-kDa membrane protein, resulting in a map at 3.2-Å overall resolution with density sufficient for de novo model building, and for the 22-kDa receptor-binding domain (RBD) of SARS-CoV-2 spike protein, resulting in a map at 3.6-Å resolution that allows analysis of the binding interface to the nanobody. The Legobody approach thus overcomes the current size limitations of cryo-EM analysis. 履歴 登録 2021年8月22日 - ヘッダ(付随情報) 公開 2021年10月6日 - マップ公開 2021年10月6日 - 更新 2025年6月4日 - 現状 2025年6月4日 処理サイト : RCSB / 状態 : 公開
すべて表示 表示を減らす
 データを開く
データを開く 基本情報
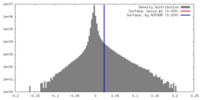
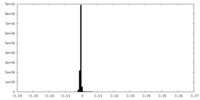
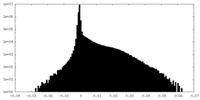
基本情報 マップデータ
マップデータ 試料
試料 キーワード
キーワード 機能・相同性情報
機能・相同性情報 Streptococcus sp. (バクテリア) /
Streptococcus sp. (バクテリア) / 


 データ登録者
データ登録者 米国, 1件
米国, 1件  引用
引用 ジャーナル: Proc Natl Acad Sci U S A / 年: 2021
ジャーナル: Proc Natl Acad Sci U S A / 年: 2021
 構造の表示
構造の表示 ムービービューア
ムービービューア SurfView
SurfView Molmil
Molmil Jmol/JSmol
Jmol/JSmol ダウンロードとリンク
ダウンロードとリンク emd_24729.map.gz
emd_24729.map.gz EMDBマップデータ形式
EMDBマップデータ形式 emd-24729-v30.xml
emd-24729-v30.xml emd-24729.xml
emd-24729.xml EMDBヘッダ
EMDBヘッダ emd_24729.png
emd_24729.png emd-24729.cif.gz
emd-24729.cif.gz emd_24729_additional_1.map.gz
emd_24729_additional_1.map.gz http://ftp.pdbj.org/pub/emdb/structures/EMD-24729
http://ftp.pdbj.org/pub/emdb/structures/EMD-24729 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24729
ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24729 emd_24729_validation.pdf.gz
emd_24729_validation.pdf.gz EMDB検証レポート
EMDB検証レポート emd_24729_full_validation.pdf.gz
emd_24729_full_validation.pdf.gz emd_24729_validation.xml.gz
emd_24729_validation.xml.gz emd_24729_validation.cif.gz
emd_24729_validation.cif.gz https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24729
https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24729 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24729
ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-24729 リンク
リンク EMDB (EBI/PDBe) /
EMDB (EBI/PDBe) /  EMDataResource
EMDataResource マップ
マップ ダウンロード / ファイル: emd_24729.map.gz / 形式: CCP4 / 大きさ: 46.4 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES)
ダウンロード / ファイル: emd_24729.map.gz / 形式: CCP4 / 大きさ: 46.4 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) 試料の構成要素
試料の構成要素 Streptococcus sp. (バクテリア)
Streptococcus sp. (バクテリア)

 Homo sapiens (ヒト)
Homo sapiens (ヒト)
 Homo sapiens (ヒト)
Homo sapiens (ヒト)


 Homo sapiens (ヒト)
Homo sapiens (ヒト) 解析
解析 試料調製
試料調製 電子顕微鏡法
電子顕微鏡法 FIELD EMISSION GUN
FIELD EMISSION GUN
 ムービー
ムービー コントローラー
コントローラー























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)