[English] 日本語
 Yorodumi
Yorodumi- EMDB-24683: Cryo-EM Structure of the Sodium-driven Chloride/Bicarbonate Excha... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-24683 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM Structure of the Sodium-driven Chloride/Bicarbonate Exchanger NDCBE (SLC4A8) | |||||||||
 Map data Map data | postprocess 3d map. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | NDCBE / sodium-driven chloride/bicarbonate exchanger / cryo-EM / MEMBRANE PROTEIN | |||||||||
| Biological species |  | |||||||||
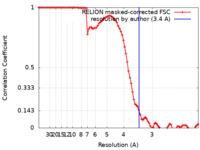
| Method | single particle reconstruction / cryo EM / Resolution: 3.4 Å | |||||||||
 Authors Authors | Wang WG / Tsirulnikov K | |||||||||
| Funding support |  United States, United States,  Canada, 2 items Canada, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Cryo-EM structure of the sodium-driven chloride/bicarbonate exchanger NDCBE. Authors: Weiguang Wang / Kirill Tsirulnikov / Hristina R Zhekova / Gülru Kayık / Hanif Muhammad Khan / Rustam Azimov / Natalia Abuladze / Liyo Kao / Debbie Newman / Sergei Yu Noskov / Z Hong Zhou / ...Authors: Weiguang Wang / Kirill Tsirulnikov / Hristina R Zhekova / Gülru Kayık / Hanif Muhammad Khan / Rustam Azimov / Natalia Abuladze / Liyo Kao / Debbie Newman / Sergei Yu Noskov / Z Hong Zhou / Alexander Pushkin / Ira Kurtz /   Abstract: SLC4 transporters play significant roles in pH regulation and cellular sodium transport. The previously solved structures of the outward facing (OF) conformation for AE1 (SLC4A1) and NBCe1 (SLC4A4) ...SLC4 transporters play significant roles in pH regulation and cellular sodium transport. The previously solved structures of the outward facing (OF) conformation for AE1 (SLC4A1) and NBCe1 (SLC4A4) transporters revealed an identical overall fold despite their different transport modes (chloride/bicarbonate exchange versus sodium-carbonate cotransport). However, the exact mechanism determining the different transport modes in the SLC4 family remains unknown. In this work, we report the cryo-EM 3.4 Å structure of the OF conformation of NDCBE (SLC4A8), which shares transport properties with both AE1 and NBCe1 by mediating the electroneutral exchange of sodium-carbonate with chloride. This structure features a fully resolved extracellular loop 3 and well-defined densities corresponding to sodium and carbonate ions in the tentative substrate binding pocket. Further, we combine computational modeling with functional studies to unravel the molecular determinants involved in NDCBE and SLC4 transport. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_24683.map.gz emd_24683.map.gz | 54.5 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-24683-v30.xml emd-24683-v30.xml emd-24683.xml emd-24683.xml | 21.4 KB 21.4 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_24683_fsc.xml emd_24683_fsc.xml | 8.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_24683.png emd_24683.png | 104.4 KB | ||
| Filedesc metadata |  emd-24683.cif.gz emd-24683.cif.gz | 6.6 KB | ||
| Others |  emd_24683_additional_1.map.gz emd_24683_additional_1.map.gz emd_24683_half_map_1.map.gz emd_24683_half_map_1.map.gz emd_24683_half_map_2.map.gz emd_24683_half_map_2.map.gz | 54.5 MB 43.9 MB 43.9 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-24683 http://ftp.pdbj.org/pub/emdb/structures/EMD-24683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24683 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-24683 | HTTPS FTP |
-Related structure data
| Related structure data |  7rtmMC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_24683.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_24683.map.gz / Format: CCP4 / Size: 58.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | postprocess 3d map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.07 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: postprocess 3d map.
| File | emd_24683_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | postprocess 3d map. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map1.
| File | emd_24683_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map1. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: half map2.
| File | emd_24683_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | half map2. | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : NDCBE(SLC4A8) dimer
| Entire | Name: NDCBE(SLC4A8) dimer |
|---|---|
| Components |
|
-Supramolecule #1: NDCBE(SLC4A8) dimer
| Supramolecule | Name: NDCBE(SLC4A8) dimer / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  |
-Macromolecule #1: Electroneutral sodium bicarbonate exchanger 1
| Macromolecule | Name: Electroneutral sodium bicarbonate exchanger 1 / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Molecular weight | Theoretical: 64.304094 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: LFGGLVLDVK RKAPWYWSDY RDALSLQCLA SFLFLYCACM SPVITFGGLL GEATEGRISA IESLFGASMT GIAYSLFAGQ PLTILGSTG PVLVFEKILF KFCKDYALSY LSLRACIGLW TAFLCIVLVA TDASSLVCYI TRFTEEAFAS LICIIFIYEA I EKLIHLAE ...String: LFGGLVLDVK RKAPWYWSDY RDALSLQCLA SFLFLYCACM SPVITFGGLL GEATEGRISA IESLFGASMT GIAYSLFAGQ PLTILGSTG PVLVFEKILF KFCKDYALSY LSLRACIGLW TAFLCIVLVA TDASSLVCYI TRFTEEAFAS LICIIFIYEA I EKLIHLAE TYPIHMHSQL DHLSLYYCRC ALPENPNNHT LQYWKEHSIP TADVNWANLT VSECQEMHGE FIGSACGHHG PY TPDVLFW SCILFFATFI VSSTLKTFKT SRYFPTRVRS TVSDFAVFLT IFTMVILDFL IGVPSPKLQV PSVFKPTRDD RGW FISPIG PNPWWTVIAA IIPALLCTIL IFMDQQITAV IINRKEHKLK KGCGYHLDLL VVAIMLGVCS LMGLPWFVAA TVLS ITHVN SLKLESECSA PGEQPKFLGI REQRVTGLMI FVLMGCSVFM TAVLKFIPMP VLYGVFLYMG VSSLQGIQFF DRLKL FGMP AKHQPDFIYL RHVPLRKVHL FTLVQLTCLV LLWVIKASPA AIVFPMMVLA LVFVRKVMDL CFSKRELSWL DDLMPE SKK KKLDDAKK |
-Macromolecule #3: 2-acetamido-2-deoxy-beta-D-glucopyranose
| Macromolecule | Name: 2-acetamido-2-deoxy-beta-D-glucopyranose / type: ligand / ID: 3 / Number of copies: 2 / Formula: NAG |
|---|---|
| Molecular weight | Theoretical: 221.208 Da |
| Chemical component information |  ChemComp-NAG: |
-Macromolecule #4: SODIUM ION
| Macromolecule | Name: SODIUM ION / type: ligand / ID: 4 / Number of copies: 2 |
|---|---|
| Molecular weight | Theoretical: 22.99 Da |
-Macromolecule #5: CARBONATE ION
| Macromolecule | Name: CARBONATE ION / type: ligand / ID: 5 / Number of copies: 2 / Formula: CO3 |
|---|---|
| Molecular weight | Theoretical: 60.009 Da |
| Chemical component information |  ChemComp-CO3: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.8 mg/mL | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 200 / Support film - Material: CARBON / Support film - topology: HOLEY / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 30 sec. / Pretreatment - Atmosphere: OTHER / Details: Glow discharged with Easyglow. | ||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV / Details: blot time 2 seconds. blot force 0. | ||||||||||||
| Details | sample in 0.01% LMNG. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Frames/image: 1-40 / Number grids imaged: 2 / Number real images: 7545 / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated magnification: 130000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Refinement | Space: REAL / Protocol: AB INITIO MODEL / Target criteria: Correlation coefficient |
|---|---|
| Output model |  PDB-7rtm: |
 Movie
Movie Controller
Controller













 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)