+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-23938 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Cryo-EM structure of the human SSU processome, state post-A1 | |||||||||
 マップデータ マップデータ | ||||||||||
 試料 試料 |
| |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報preribosome / U6 snRNA 2'-O-ribose methyltransferase activity / nucleolar exosome (RNase complex) / oocyte growth / nucleologenesis / 18S rRNA (adenine1779-N6/adenine1780-N6)-dimethyltransferase / 18S rRNA (adenine(1779)-N(6)/adenine(1780)-N(6))-dimethyltransferase activity / snoRNA localization / nuclear polyadenylation-dependent snoRNA catabolic process / nuclear polyadenylation-dependent snRNA catabolic process ...preribosome / U6 snRNA 2'-O-ribose methyltransferase activity / nucleolar exosome (RNase complex) / oocyte growth / nucleologenesis / 18S rRNA (adenine1779-N6/adenine1780-N6)-dimethyltransferase / 18S rRNA (adenine(1779)-N(6)/adenine(1780)-N(6))-dimethyltransferase activity / snoRNA localization / nuclear polyadenylation-dependent snoRNA catabolic process / nuclear polyadenylation-dependent snRNA catabolic process / granular component / nuclear polyadenylation-dependent antisense transcript catabolic process / regulation of telomerase RNA localization to Cajal body / positive regulation of mRNA cis splicing, via spliceosome / RNA exonuclease activity / t-UTP complex / positive regulation of male gonad development / nuclear polyadenylation-dependent CUT catabolic process / CURI complex / UTP-C complex / TRAMP-dependent tRNA surveillance pathway / regulation of stem cell population maintenance / cytoplasmic exosome (RNase complex) / U4atac snRNP / CUT catabolic process / nuclear polyadenylation-dependent rRNA catabolic process / poly(A)-dependent snoRNA 3'-end processing / exosome (RNase complex) / Mpp10 complex / Pwp2p-containing subcomplex of 90S preribosome / U4atac snRNA binding / histone H2AQ104 methyltransferase activity / rRNA (pseudouridine) methyltransferase activity / nuclear exosome (RNase complex) / exonucleolytic trimming to generate mature 3'-end of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / pre-snoRNP complex / endonucleolytic cleavage in 5'-ETS of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA (adenine-N6,N6-)-dimethyltransferase activity / box C/D sno(s)RNA binding / box C/D sno(s)RNA 3'-end processing / dense fibrillar component / histone mRNA catabolic process / endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / rRNA methyltransferase activity / endonucleolytic cleavage to generate mature 5'-end of SSU-rRNA from (SSU-rRNA, 5.8S rRNA, LSU-rRNA) / nuclear mRNA surveillance / histone methyltransferase binding / positive regulation of rRNA processing / regulation of transcription elongation by RNA polymerase II / tRNA export from nucleus / box C/D methylation guide snoRNP complex / spindle assembly involved in female meiosis / positive regulation of respiratory burst involved in inflammatory response / cilium disassembly / rRNA base methylation / nucleolus organization / epigenetic programming in the zygotic pronuclei / transcription elongation factor activity / Cul4-RING E3 ubiquitin ligase complex / rRNA primary transcript binding / blastocyst formation / sno(s)RNA-containing ribonucleoprotein complex / RNA splicing, via transesterification reactions / U4 snRNA binding / protein localization to nucleolus / negative regulation of RNA splicing / telomerase RNA binding / SUMOylation of RNA binding proteins / RNA catabolic process / rRNA methylation / U2-type precatalytic spliceosome / U3 snoRNA binding / neural crest cell differentiation / box C/D snoRNP assembly / rRNA modification in the nucleus and cytosol / erythrocyte homeostasis / Formation of the ternary complex, and subsequently, the 43S complex / cytoplasmic side of rough endoplasmic reticulum membrane / nuclear-transcribed mRNA catabolic process, nonsense-mediated decay / maturation of 5.8S rRNA / preribosome, small subunit precursor / negative regulation of ubiquitin protein ligase activity / Ribosomal scanning and start codon recognition / snoRNA binding / precatalytic spliceosome / Translation initiation complex formation / positive regulation of transcription by RNA polymerase I / fibroblast growth factor binding / mammalian oogenesis stage / activation-induced cell death of T cells / monocyte chemotaxis / RNA polymerase II complex binding / Protein hydroxylation / Association of TriC/CCT with target proteins during biosynthesis / mTORC1-mediated signalling / SARS-CoV-1 modulates host translation machinery / Peptide chain elongation / positive regulation of intrinsic apoptotic signaling pathway by p53 class mediator / TFIID-class transcription factor complex binding / Selenocysteine synthesis 類似検索 - 分子機能 | |||||||||
| 生物種 |  Homo sapiens (ヒト) / Homo sapiens (ヒト) /  Human (ヒト) Human (ヒト) | |||||||||
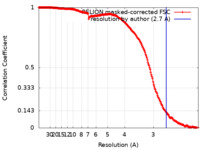
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.7 Å | |||||||||
 データ登録者 データ登録者 | Vanden Broeck A / Singh S / Klinge S | |||||||||
| 資金援助 |  米国, 2件 米国, 2件
| |||||||||
 引用 引用 |  ジャーナル: Science / 年: 2021 ジャーナル: Science / 年: 2021タイトル: Nucleolar maturation of the human small subunit processome. 著者: Sameer Singh / Arnaud Vanden Broeck / Linamarie Miller / Malik Chaker-Margot / Sebastian Klinge /  要旨: The human small subunit processome mediates early maturation of the small ribosomal subunit by coupling RNA folding to subsequent RNA cleavage and processing steps. We report the high-resolution ...The human small subunit processome mediates early maturation of the small ribosomal subunit by coupling RNA folding to subsequent RNA cleavage and processing steps. We report the high-resolution cryo–electron microscopy structures of maturing human small subunit (SSU) processomes at resolutions of 2.7 to 3.9 angstroms. These structures reveal the molecular mechanisms that enable crucial progressions during SSU processome maturation. RNA folding states within these particles are communicated to and coordinated with key enzymes that drive irreversible steps such as targeted exosome-mediated RNA degradation, protein-guided site-specific endonucleolytic RNA cleavage, and tightly controlled RNA unwinding. These conserved mechanisms highlight the SSU processome’s impressive structural plasticity, which endows this 4.5-megadalton nucleolar assembly with the distinctive ability to mature the small ribosomal subunit from within. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_23938.map.gz emd_23938.map.gz | 619.1 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-23938-v30.xml emd-23938-v30.xml emd-23938.xml emd-23938.xml | 118.9 KB 118.9 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
| FSC (解像度算出) |  emd_23938_fsc.xml emd_23938_fsc.xml | 19.7 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_23938.png emd_23938.png | 163.1 KB | ||
| マスクデータ |  emd_23938_msk_1.map emd_23938_msk_1.map | 669.9 MB |  マスクマップ マスクマップ | |
| その他 |  emd_23938_additional_1.map.gz emd_23938_additional_1.map.gz emd_23938_half_map_1.map.gz emd_23938_half_map_1.map.gz emd_23938_half_map_2.map.gz emd_23938_half_map_2.map.gz | 616.4 MB 536.2 MB 536.3 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23938 http://ftp.pdbj.org/pub/emdb/structures/EMD-23938 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23938 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23938 | HTTPS FTP |
-検証レポート
| 文書・要旨 |  emd_23938_validation.pdf.gz emd_23938_validation.pdf.gz | 842.5 KB | 表示 |  EMDB検証レポート EMDB検証レポート |
|---|---|---|---|---|
| 文書・詳細版 |  emd_23938_full_validation.pdf.gz emd_23938_full_validation.pdf.gz | 842 KB | 表示 | |
| XML形式データ |  emd_23938_validation.xml.gz emd_23938_validation.xml.gz | 27.8 KB | 表示 | |
| CIF形式データ |  emd_23938_validation.cif.gz emd_23938_validation.cif.gz | 37.6 KB | 表示 | |
| アーカイブディレクトリ |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23938 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23938 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23938 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-23938 | HTTPS FTP |
-関連構造データ
| 関連構造データ |  7mqaMC  7mq8C  7mq9C  7mqjC C: 同じ文献を引用 ( M: このマップから作成された原子モデル |
|---|---|
| 類似構造データ | |
| 電子顕微鏡画像生データ |  EMPIAR-10781 (タイトル: Nucleolar maturation of the human small subunit processome EMPIAR-10781 (タイトル: Nucleolar maturation of the human small subunit processomeData size: 74.6 TB Data #1: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 1 [micrographs - multiframe] Data #2: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 2 [micrographs - multiframe] Data #3: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 3 [micrographs - multiframe] Data #4: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 4 [micrographs - multiframe] Data #5: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 5 [micrographs - multiframe] Data #6: Unaligned multi-frame micrograph movies of human SSU processomes - Dataset 6 [micrographs - multiframe] Data #7: Aligned and averaged micrographs of human SSU processomes - Dataset 1 [micrographs - single frame] Data #8: Aligned and averaged micrographs of human SSU processomes - Dataset 2 [micrographs - single frame] Data #9: Aligned and averaged micrographs of human SSU processomes - Dataset 3 [micrographs - single frame] Data #10: Aligned and averaged micrographs of human SSU processomes - Dataset 4 [micrographs - single frame] Data #11: Aligned and averaged micrographs of human SSU processomes - Dataset 5 [micrographs - single frame] Data #12: Aligned and averaged micrographs of human SSU processomes - Dataset 6 [micrographs - single frame]) |
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_23938.map.gz / 形式: CCP4 / 大きさ: 669.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_23938.map.gz / 形式: CCP4 / 大きさ: 669.9 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.08 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
-マスク #1
| ファイル |  emd_23938_msk_1.map emd_23938_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-追加マップ: #1
| ファイル | emd_23938_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #2
| ファイル | emd_23938_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
-ハーフマップ: #1
| ファイル | emd_23938_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 投影像・断面図 |
| ||||||||||||
| 密度ヒストグラム |
- 試料の構成要素
試料の構成要素
+全体 : Human SSU processome
+超分子 #1: Human SSU processome
+分子 #1: 5'ETS rRNA
+分子 #2: 18S rRNA
+分子 #3: U3 snoRNA
+分子 #4: 40S ribosomal protein S18
+分子 #5: 40S ribosomal protein S4, X isoform
+分子 #6: 40S ribosomal protein S5
+分子 #7: 40S ribosomal protein S6
+分子 #8: 40S ribosomal protein S7
+分子 #9: 40S ribosomal protein S8
+分子 #10: 40S ribosomal protein S9
+分子 #11: 40S ribosomal protein S12
+分子 #12: 40S ribosomal protein S16
+分子 #13: 40S ribosomal protein S11
+分子 #14: 40S ribosomal protein S24
+分子 #15: 40S ribosomal protein S28
+分子 #16: WD repeat-containing protein 75
+分子 #17: Nucleolar protein 11
+分子 #18: U3 small nucleolar RNA-associated protein 15 homolog
+分子 #19: WD repeat-containing protein 43
+分子 #20: HEAT repeat-containing protein 1
+分子 #21: U3 small nucleolar RNA-associated protein 4 homolog
+分子 #22: Periodic tryptophan protein 2 homolog
+分子 #23: U3 small nucleolar RNA-associated protein 6 homolog
+分子 #24: WD repeat-containing protein 3
+分子 #25: Transducin beta-like protein 3
+分子 #26: U3 small nucleolar RNA-associated protein 18 homolog
+分子 #27: WD repeat-containing protein 36
+分子 #28: DDB1- and CUL4-associated factor 13
+分子 #29: WD repeat-containing protein 46
+分子 #30: Active regulator of SIRT1
+分子 #31: U3 small nucleolar ribonucleoprotein protein IMP3
+分子 #32: U3 small nucleolar ribonucleoprotein protein MPP10
+分子 #33: Something about silencing protein 10
+分子 #34: Nucleolar protein 7
+分子 #35: 40S ribosomal protein S13
+分子 #36: 40S ribosomal protein S14
+分子 #37: Nucleolar protein 6
+分子 #38: Ribosomal RNA-processing protein 7 homolog A
+分子 #39: Probable dimethyladenosine transferase
+分子 #40: 40S ribosomal protein S3a
+分子 #41: 40S ribosomal protein S15a
+分子 #42: 40S ribosomal protein S19
+分子 #43: 40S ribosomal protein S27
+分子 #44: RRP12-like protein
+分子 #45: Probable ATP-dependent RNA helicase DHX37
+分子 #46: Ubiquitin-40S ribosomal protein S27a
+分子 #47: 40S ribosomal protein S17
+分子 #48: Exosome component 10
+分子 #49: Nucleolar protein 56
+分子 #50: Nucleolar protein 58
+分子 #51: rRNA 2'-O-methyltransferase fibrillarin
+分子 #52: NHP2-like protein 1
+分子 #53: U3 small nucleolar RNA-interacting protein 2
+分子 #54: RNA 3'-terminal phosphate cyclase-like protein
+分子 #55: Ribosome biogenesis protein BMS1 homolog
+分子 #56: Ribosomal RNA small subunit methyltransferase NEP1
+分子 #57: rRNA-processing protein FCF1 homolog
+分子 #58: U3 small nucleolar ribonucleoprotein protein IMP4
+分子 #59: Small subunit processome component 20 homolog
+分子 #60: Deoxynucleotidyltransferase terminal-interacting protein 2
+分子 #61: 40S ribosomal protein S23
+分子 #62: U3 small nucleolar RNA-associated protein 14 homolog A
+分子 #63: Nucleolar protein 14
+分子 #64: Nucleolar complex protein 4 homolog
+分子 #65: RNA-binding protein PNO1
+分子 #66: Unassigned peptides
+分子 #67: Probable U3 small nucleolar RNA-associated protein 11
+分子 #68: Bystin
+分子 #69: MAGNESIUM ION
+分子 #70: ADENOSINE-5'-TRIPHOSPHATE
+分子 #71: ZINC ION
+分子 #72: GUANOSINE-5'-TRIPHOSPHATE
+分子 #73: S-ADENOSYL-L-HOMOCYSTEINE
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 緩衝液 | pH: 7.6 |
|---|---|
| グリッド | モデル: Quantifoil R2/2 / 材質: GOLD / メッシュ: 400 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 支持フィルム - Film thickness: 3.0 nm / 前処理 - タイプ: GLOW DISCHARGE |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 90 % / チャンバー内温度: 283 K / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 特殊光学系 | エネルギーフィルター - スリット幅: 20 eV |
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 実像数: 84904 / 平均電子線量: 58.0 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD / Cs: 0.01 mm / 最大 デフォーカス(公称値): 2.7 µm 最小 デフォーカス(公称値): 0.7000000000000001 µm |
| 試料ステージ | 試料ホルダーモデル: FEI TITAN KRIOS AUTOGRID HOLDER |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー















































































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)