+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-20551 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of a MAPK pathway complex | |||||||||
 Map data Map data | Structure of a MAPK pathway complex | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | TRANSFERASE / SIGNALING PROTEIN-Transferase complex | |||||||||
| Function / homology |  Function and homology information Function and homology informationsynaptic target recognition / Golgi reassembly / CD4-positive, alpha-beta T cell differentiation / NOTCH4 Activation and Transmission of Signal to the Nucleus / positive regulation of axon regeneration / CD4-positive or CD8-positive, alpha-beta T cell lineage commitment / negative regulation of synaptic vesicle exocytosis / establishment of Golgi localization / Signalling to p38 via RIT and RIN / respiratory system process ...synaptic target recognition / Golgi reassembly / CD4-positive, alpha-beta T cell differentiation / NOTCH4 Activation and Transmission of Signal to the Nucleus / positive regulation of axon regeneration / CD4-positive or CD8-positive, alpha-beta T cell lineage commitment / negative regulation of synaptic vesicle exocytosis / establishment of Golgi localization / Signalling to p38 via RIT and RIN / respiratory system process / head morphogenesis / ARMS-mediated activation / myeloid progenitor cell differentiation / tube formation / endothelial cell apoptotic process / regulation of synapse maturation / SHOC2 M1731 mutant abolishes MRAS complex function / Gain-of-function MRAS complexes activate RAF signaling / negative regulation of fibroblast migration / positive regulation of D-glucose transmembrane transport / Rap1 signalling / establishment of protein localization to membrane / regulation of T cell differentiation / positive regulation of axonogenesis / negative regulation of protein localization to nucleus / KSRP (KHSRP) binds and destabilizes mRNA / Negative feedback regulation of MAPK pathway / GP1b-IX-V activation signalling / Frs2-mediated activation / stress fiber assembly / face development / MAP kinase kinase activity / thyroid gland development / Regulation of localization of FOXO transcription factors / Interleukin-3, Interleukin-5 and GM-CSF signaling / somatic stem cell population maintenance / positive regulation of peptidyl-serine phosphorylation / synaptic vesicle exocytosis / Activation of BAD and translocation to mitochondria / phosphoserine residue binding / MAP kinase kinase kinase activity / negative regulation of endothelial cell apoptotic process / regulation of ERK1 and ERK2 cascade / SARS-CoV-2 targets host intracellular signalling and regulatory pathways / postsynaptic modulation of chemical synaptic transmission / protein targeting / cellular response to glucose starvation / SARS-CoV-1 targets host intracellular signalling and regulatory pathways / RHO GTPases activate PKNs / Chk1/Chk2(Cds1) mediated inactivation of Cyclin B:Cdk1 complex / ERK1 and ERK2 cascade / positive regulation of stress fiber assembly / negative regulation of TORC1 signaling / positive regulation of substrate adhesion-dependent cell spreading / Transcriptional and post-translational regulation of MITF-M expression and activity / substrate adhesion-dependent cell spreading / lung development / cellular response to calcium ion / negative regulation of innate immune response / thymus development / animal organ morphogenesis / hippocampal mossy fiber to CA3 synapse / TP53 Regulates Metabolic Genes / Translocation of SLC2A4 (GLUT4) to the plasma membrane / Deactivation of the beta-catenin transactivating complex / sperm end piece / protein sequestering activity / regulation of protein stability / RAF activation / Spry regulation of FGF signaling / Signaling by high-kinase activity BRAF mutants / MAP2K and MAPK activation / Negative regulation of NOTCH4 signaling / visual learning / cellular response to xenobiotic stimulus / epidermal growth factor receptor signaling pathway / centriolar satellite / Signaling by RAF1 mutants / Signaling by moderate kinase activity BRAF mutants / Paradoxical activation of RAF signaling by kinase inactive BRAF / Signaling downstream of RAS mutants / Negative regulation of MAPK pathway / Signaling by BRAF and RAF1 fusions / melanosome / intracellular protein localization / T cell differentiation in thymus / long-term synaptic potentiation / MAPK cascade / regulation of cell population proliferation / T cell receptor signaling pathway / sperm principal piece / presynapse / cell body / angiogenesis / scaffold protein binding / sperm midpiece / protein phosphatase binding / blood microparticle / DNA-binding transcription factor binding / vesicle Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 6.8 Å | |||||||||
 Authors Authors | Park E / Rawson S | |||||||||
| Funding support |  United States, 2 items United States, 2 items
| |||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Architecture of autoinhibited and active BRAF-MEK1-14-3-3 complexes. Authors: Eunyoung Park / Shaun Rawson / Kunhua Li / Byeong-Won Kim / Scott B Ficarro / Gonzalo Gonzalez-Del Pino / Humayun Sharif / Jarrod A Marto / Hyesung Jeon / Michael J Eck /  Abstract: RAF family kinases are RAS-activated switches that initiate signalling through the MAP kinase cascade to control cellular proliferation, differentiation and survival. RAF activity is tightly ...RAF family kinases are RAS-activated switches that initiate signalling through the MAP kinase cascade to control cellular proliferation, differentiation and survival. RAF activity is tightly regulated and inappropriate activation is a frequent cause of cancer; however, the structural basis for RAF regulation is poorly understood at present. Here we use cryo-electron microscopy to determine autoinhibited and active-state structures of full-length BRAF in complexes with MEK1 and a 14-3-3 dimer. The reconstruction reveals an inactive BRAF-MEK1 complex restrained in a cradle formed by the 14-3-3 dimer, which binds the phosphorylated S365 and S729 sites that flank the BRAF kinase domain. The BRAF cysteine-rich domain occupies a central position that stabilizes this assembly, but the adjacent RAS-binding domain is poorly ordered and peripheral. The 14-3-3 cradle maintains autoinhibition by sequestering the membrane-binding cysteine-rich domain and blocking dimerization of the BRAF kinase domain. In the active state, these inhibitory interactions are released and a single 14-3-3 dimer rearranges to bridge the C-terminal pS729 binding sites of two BRAFs, which drives the formation of an active, back-to-back BRAF dimer. Our structural snapshots provide a foundation for understanding normal RAF regulation and its mutational disruption in cancer and developmental syndromes. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_20551.map.gz emd_20551.map.gz | 42.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-20551-v30.xml emd-20551-v30.xml emd-20551.xml emd-20551.xml | 12.9 KB 12.9 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_20551.png emd_20551.png | 55.4 KB | ||
| Filedesc metadata |  emd-20551.cif.gz emd-20551.cif.gz | 6 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-20551 http://ftp.pdbj.org/pub/emdb/structures/EMD-20551 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20551 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-20551 | HTTPS FTP |
-Related structure data
| Related structure data |  6q0kMC  0541C  6nybC  6pp9C  6q0jC  6q0tC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_20551.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_20551.map.gz / Format: CCP4 / Size: 64 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
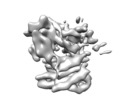
| Annotation | Structure of a MAPK pathway complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 0.85 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : ERK pathway complex
| Entire | Name: ERK pathway complex |
|---|---|
| Components |
|
-Supramolecule #1: ERK pathway complex
| Supramolecule | Name: ERK pathway complex / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 233 kDa/nm |
-Macromolecule #1: Serine/threonine-protein kinase B-raf
| Macromolecule | Name: Serine/threonine-protein kinase B-raf / type: protein_or_peptide / ID: 1 / Number of copies: 2 / Enantiomer: LEVO / EC number: non-specific serine/threonine protein kinase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 89.322812 KDa |
| Recombinant expression | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Sequence | String: MSYYHHHHHH HHDIPTTENL YFQGAMDMAA LSGGGGGGAE PGQALFNGDM EPEAGAGAGA AASSAADPAI PEEVWNIKQM IKLTQEHIE ALLDKFGGEH NPPSIYLEAY EEYTSKLDAL QQREQQLLES LGNGTDFSVS SSASMDTVTS SSSSSLSVLP S SLSVFQNP ...String: MSYYHHHHHH HHDIPTTENL YFQGAMDMAA LSGGGGGGAE PGQALFNGDM EPEAGAGAGA AASSAADPAI PEEVWNIKQM IKLTQEHIE ALLDKFGGEH NPPSIYLEAY EEYTSKLDAL QQREQQLLES LGNGTDFSVS SSASMDTVTS SSSSSLSVLP S SLSVFQNP TDVARSNPKS PQKPIVRVFL PNKQRTVVPA RCGVTVRDSL KKALMMRGLI PECCAVYRIQ DGEKKPIGWD TD ISWLTGE ELHVEVLENV PLTTHNFVRK TFFTLAFCDF CRKLLFQGFR CQTCGYKFHQ RCSTEVPLMC VNYDQLDLLF VSK FFEHHP IPQEEASLAE TALTSGSSPS APASDSIGPQ ILTSPSPSKS IPIPQPFRPA DEDHRNQFGQ RDRSSSAPNV HINT IEPVN IDDLIRDQGF RGDGGSTTGL SATPPASLPG SLTNVKALQK SPGPQRERKS SSSSEDRNRM KTLGRRDSSD DWEIP DGQI TVGQRIGSGS FGTVYKGKWH GDVAVKMLNV TAPTPQQLQA FKNEVGVLRK TRHVNILLFM GYSTKPQLAI VTQWCE GSS LYHHLHIIET KFEMIKLIDI ARQTAQGMDY LHAKSIIHRD LKSNNIFLHE DLTVKIGDFG LATVKSRWSG SHQFEQL SG SILWMAPEVI RMQDKNPYSF QSDVYAFGIV LYELMTGQLP YSNINNRDQI IFMVGRGYLS PDLSKVRSNC PKAMKRLM A ECLKKKRDER PLFPQILASI ELLARSLPKI HRSA(SEP)EPSLN RAGFQTEDFS LYACASPKTP IQAGGYGAFP VHGTS AWSH PQFEK UniProtKB: Serine/threonine-protein kinase B-raf |
-Macromolecule #2: 14-3-3 protein zeta/delta
| Macromolecule | Name: 14-3-3 protein zeta/delta / type: protein_or_peptide / ID: 2 / Number of copies: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 27.777092 KDa |
| Sequence | String: MDKNELVQKA KLAEQAERYD DMAACMKSVT EQGAELSNEE RNLLSVAYKN VVGARRSSWR VVSSIEQKTE GAEKKQQMAR EYREKIETE LRDICNDVLS LLEKFLIPNA SQAESKVFYL KMKGDYYRYL AEVAAGDDKK GIVDQSQQAY QEAFEISKKE M QPTHPIRL ...String: MDKNELVQKA KLAEQAERYD DMAACMKSVT EQGAELSNEE RNLLSVAYKN VVGARRSSWR VVSSIEQKTE GAEKKQQMAR EYREKIETE LRDICNDVLS LLEKFLIPNA SQAESKVFYL KMKGDYYRYL AEVAAGDDKK GIVDQSQQAY QEAFEISKKE M QPTHPIRL GLALNFSVFY YEILNSPEKA CSLAKTAFDE AIAELDTLSE ESYKDSTLIM QLLRDNLTLW TSDTQGDEAE AG EGGEN UniProtKB: 14-3-3 protein zeta/delta |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Details: unspecified |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 60.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: OTHER |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
- Image processing
Image processing
| Startup model | Type of model: OTHER |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 6.8 Å / Resolution method: FSC 0.143 CUT-OFF / Number images used: 66215 |
| Initial angle assignment | Type: MAXIMUM LIKELIHOOD |
| Final angle assignment | Type: MAXIMUM LIKELIHOOD |
-Atomic model buiding 1
| Initial model |
| ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| Refinement | Space: REAL / Protocol: RIGID BODY FIT | ||||||||||
| Output model |  PDB-6q0k: |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)























