+ データを開く
データを開く
- 基本情報
基本情報
| 登録情報 | データベース: EMDB / ID: EMD-13004 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| タイトル | Pol II-CSB-CSA-DDB1-UVSSA (Structure1) | |||||||||
 マップデータ マップデータ | Main map. | |||||||||
 試料 試料 |
| |||||||||
 キーワード キーワード | transcription / DNA repair | |||||||||
| 機能・相同性 |  機能・相同性情報 機能・相同性情報RNA polymerase inhibitor activity / negative regulation of double-strand break repair via nonhomologous end joining / nucleotide-excision repair complex / regulation of transcription-coupled nucleotide-excision repair / : / response to auditory stimulus / regulation of transcription elongation by RNA polymerase II / DNA protection / single strand break repair / B-WICH complex ...RNA polymerase inhibitor activity / negative regulation of double-strand break repair via nonhomologous end joining / nucleotide-excision repair complex / regulation of transcription-coupled nucleotide-excision repair / : / response to auditory stimulus / regulation of transcription elongation by RNA polymerase II / DNA protection / single strand break repair / B-WICH complex / Formation of RNA Pol II elongation complex / Formation of the Early Elongation Complex / Transcriptional regulation by small RNAs / RNA Polymerase II Pre-transcription Events / TP53 Regulates Transcription of DNA Repair Genes / FGFR2 alternative splicing / RNA polymerase II transcribes snRNA genes / mRNA Capping / mRNA Splicing - Minor Pathway / Processing of Capped Intron-Containing Pre-mRNA / RNA Polymerase II Promoter Escape / RNA Polymerase II Transcription Pre-Initiation And Promoter Opening / RNA Polymerase II Transcription Initiation / RNA Polymerase II Transcription Elongation / RNA Polymerase II Transcription Initiation And Promoter Clearance / RNA Pol II CTD phosphorylation and interaction with CE / Estrogen-dependent gene expression / response to superoxide / Formation of TC-NER Pre-Incision Complex / Dual incision in TC-NER / Gap-filling DNA repair synthesis and ligation in TC-NER / mRNA Splicing - Major Pathway / double-strand break repair via classical nonhomologous end joining / photoreceptor cell maintenance / positive regulation by virus of viral protein levels in host cell / RNA polymerase binding / chromatin-protein adaptor activity / positive regulation of DNA-templated transcription, elongation / spindle assembly involved in female meiosis / response to UV-B / epigenetic programming in the zygotic pronuclei / ATP-dependent chromatin remodeler activity / positive regulation of transcription by RNA polymerase III / UV-damage excision repair / biological process involved in interaction with symbiont / regulation of mitotic cell cycle phase transition / ATP-dependent DNA damage sensor activity / WD40-repeat domain binding / positive regulation of nuclear-transcribed mRNA poly(A) tail shortening / Cul4A-RING E3 ubiquitin ligase complex / Cul4-RING E3 ubiquitin ligase complex / organelle membrane / Cul4B-RING E3 ubiquitin ligase complex / positive regulation of transcription by RNA polymerase I / ubiquitin ligase complex scaffold activity / negative regulation of reproductive process / negative regulation of developmental process / RNA polymerase II complex binding / RNA Polymerase I Transcription Initiation / protein tyrosine kinase activator activity / maintenance of transcriptional fidelity during transcription elongation by RNA polymerase II / pyrimidine dimer repair / viral release from host cell / positive regulation of translational initiation / cullin family protein binding / response to X-ray / positive regulation of transcription initiation by RNA polymerase II / ectopic germ cell programmed cell death / ATP-dependent activity, acting on DNA / RNA polymerase I complex / positive regulation of viral genome replication / RNA polymerase III complex / response to UV / transcription elongation by RNA polymerase I / protein autoubiquitination / RNA polymerase II, core complex / positive regulation of double-strand break repair via homologous recombination / tRNA transcription by RNA polymerase III / ubiquitin-like ligase-substrate adaptor activity / site of DNA damage / transcription by RNA polymerase I / JNK cascade / protein localization to chromatin / transcription-coupled nucleotide-excision repair / translation initiation factor binding / proteasomal protein catabolic process / neurogenesis / positive regulation of gluconeogenesis / DNA damage checkpoint signaling / regulation of DNA-templated transcription elongation / positive regulation of RNA splicing / positive regulation of DNA repair / transcription elongation factor complex / helicase activity / ERCC6 (CSB) and EHMT2 (G9a) positively regulate rRNA expression / response to gamma radiation / nucleotide-excision repair / sperm end piece / transcription initiation at RNA polymerase II promoter / P-body 類似検索 - 分子機能 | |||||||||
| 生物種 |   Homo sapiens (ヒト) Homo sapiens (ヒト) | |||||||||
| 手法 | 単粒子再構成法 / クライオ電子顕微鏡法 / 解像度: 2.8 Å | |||||||||
 データ登録者 データ登録者 | Kokic G / Cramer P | |||||||||
| 資金援助 |  ドイツ, 2件 ドイツ, 2件
| |||||||||
 引用 引用 |  ジャーナル: Nature / 年: 2021 ジャーナル: Nature / 年: 2021タイトル: Structural basis of human transcription-DNA repair coupling. 著者: Goran Kokic / Felix R Wagner / Aleksandar Chernev / Henning Urlaub / Patrick Cramer /  要旨: Transcription-coupled DNA repair removes bulky DNA lesions from the genome and protects cells against ultraviolet (UV) irradiation. Transcription-coupled DNA repair begins when RNA polymerase II ...Transcription-coupled DNA repair removes bulky DNA lesions from the genome and protects cells against ultraviolet (UV) irradiation. Transcription-coupled DNA repair begins when RNA polymerase II (Pol II) stalls at a DNA lesion and recruits the Cockayne syndrome protein CSB, the E3 ubiquitin ligase, CRL4 and UV-stimulated scaffold protein A (UVSSA). Here we provide five high-resolution structures of Pol II transcription complexes containing human transcription-coupled DNA repair factors and the elongation factors PAF1 complex (PAF) and SPT6. Together with biochemical and published data, the structures provide a model for transcription-repair coupling. Stalling of Pol II at a DNA lesion triggers replacement of the elongation factor DSIF by CSB, which binds to PAF and moves upstream DNA to SPT6. The resulting elongation complex, EC, uses the CSA-stimulated translocase activity of CSB to pull on upstream DNA and push Pol II forward. If the lesion cannot be bypassed, CRL4 spans over the Pol II clamp and ubiquitylates the RPB1 residue K1268, enabling recruitment of TFIIH to UVSSA and DNA repair. Conformational changes in CRL4 lead to ubiquitylation of CSB and to release of transcription-coupled DNA repair factors before transcription may continue over repaired DNA. | |||||||||
| 履歴 |
|
- 構造の表示
構造の表示
| ムービー |
 ムービービューア ムービービューア |
|---|---|
| 構造ビューア | EMマップ:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| 添付画像 |
- ダウンロードとリンク
ダウンロードとリンク
-EMDBアーカイブ
| マップデータ |  emd_13004.map.gz emd_13004.map.gz | 114.3 MB |  EMDBマップデータ形式 EMDBマップデータ形式 | |
|---|---|---|---|---|
| ヘッダ (付随情報) |  emd-13004-v30.xml emd-13004-v30.xml emd-13004.xml emd-13004.xml | 76.6 KB 76.6 KB | 表示 表示 |  EMDBヘッダ EMDBヘッダ |
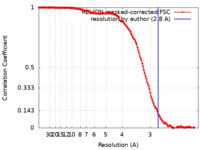
| FSC (解像度算出) |  emd_13004_fsc.xml emd_13004_fsc.xml | 14.1 KB | 表示 |  FSCデータファイル FSCデータファイル |
| 画像 |  emd_13004.png emd_13004.png | 159.2 KB | ||
| マスクデータ |  emd_13004_msk_1.map emd_13004_msk_1.map emd_13004_msk_2.map emd_13004_msk_2.map emd_13004_msk_3.map emd_13004_msk_3.map emd_13004_msk_4.map emd_13004_msk_4.map emd_13004_msk_5.map emd_13004_msk_5.map | 244.1 MB 244.1 MB 244.1 MB 244.1 MB 244.1 MB |  マスクマップ マスクマップ | |
| Filedesc metadata |  emd-13004.cif.gz emd-13004.cif.gz | 15.1 KB | ||
| その他 |  emd_13004_additional_1.map.gz emd_13004_additional_1.map.gz emd_13004_additional_10.map.gz emd_13004_additional_10.map.gz emd_13004_additional_2.map.gz emd_13004_additional_2.map.gz emd_13004_additional_3.map.gz emd_13004_additional_3.map.gz emd_13004_additional_4.map.gz emd_13004_additional_4.map.gz emd_13004_additional_5.map.gz emd_13004_additional_5.map.gz emd_13004_additional_6.map.gz emd_13004_additional_6.map.gz emd_13004_additional_7.map.gz emd_13004_additional_7.map.gz emd_13004_additional_8.map.gz emd_13004_additional_8.map.gz emd_13004_additional_9.map.gz emd_13004_additional_9.map.gz emd_13004_half_map_1.map.gz emd_13004_half_map_1.map.gz emd_13004_half_map_2.map.gz emd_13004_half_map_2.map.gz | 216.5 MB 217 MB 194.2 MB 194.6 MB 194.2 MB 194.6 MB 194.7 MB 194.6 MB 194.8 MB 194.8 MB 225.9 MB 226 MB | ||
| アーカイブディレクトリ |  http://ftp.pdbj.org/pub/emdb/structures/EMD-13004 http://ftp.pdbj.org/pub/emdb/structures/EMD-13004 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13004 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-13004 | HTTPS FTP |
-関連構造データ
- リンク
リンク
| EMDBのページ |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| 「今月の分子」の関連する項目 |
- マップ
マップ
| ファイル |  ダウンロード / ファイル: emd_13004.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) ダウンロード / ファイル: emd_13004.map.gz / 形式: CCP4 / 大きさ: 244.1 MB / タイプ: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 注釈 | Main map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
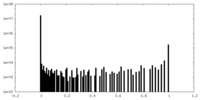
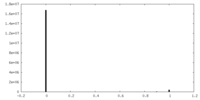
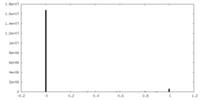
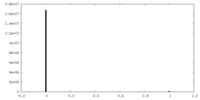
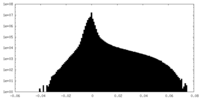
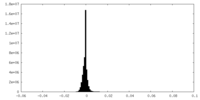
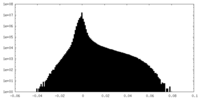
| 投影像・断面図 | 画像のコントロール
画像は Spider により作成 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| ボクセルのサイズ | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 密度 |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 対称性 | 空間群: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| 詳細 | EMDB XML:
CCP4マップ ヘッダ情報:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-添付データ
+マスク #1
+マスク #2
+マスク #3
+マスク #4
+マスク #5
+追加マップ: Half map corresponding to the map focused refined on CSA-UVSSA.
+追加マップ: Half map corresponding to the map focused refined on CSA-UVSSA.
+追加マップ: Half map corresponding to the map focused refined on CSB.
+追加マップ: Half map corresponding to the map focused refined on CSA-DDB1.
+追加マップ: Half map corresponding to the map focused refined on CSB.
+追加マップ: Half map corresponding to the map focused refined on CSA-DDB1.
+追加マップ: Half map corresponding to the map focused refined on DDB1.
+追加マップ: Half map corresponding to the map focused refined on DDB1.
+追加マップ: Half map corresponding to the map focused refined on Pol II.
+追加マップ: Half map corresponding to the map focused refined on Pol II.
+ハーフマップ: Half map corresponding to the main map.
+ハーフマップ: Half map corresponding to the main map.
- 試料の構成要素
試料の構成要素
+全体 : RNA polymerase II elongation complex with CSB, CSA-DDB1 and UVSSA.
+超分子 #1: RNA polymerase II elongation complex with CSB, CSA-DDB1 and UVSSA.
+超分子 #2: DNA directed RNA polymerase
+超分子 #3: NTS, TS & RNA
+超分子 #4: DNA Repair proteins, CSB element and scaffold protein
+分子 #1: DNA-directed RNA polymerase II subunit RPB1
+分子 #2: DNA-directed RNA polymerase subunit beta
+分子 #3: DNA-directed RNA polymerase II subunit RPB3
+分子 #4: RPOL4c domain-containing protein
+分子 #5: DNA-directed RNA polymerase II subunit E
+分子 #6: DNA-directed RNA polymerase II subunit F
+分子 #7: DNA-directed RNA polymerase II subunit RPB7
+分子 #8: DNA-directed RNA polymerases I, II, and III subunit RPABC3
+分子 #9: DNA-directed RNA polymerase II subunit RPB9
+分子 #10: DNA-directed RNA polymerases I, II, and III subunit RPABC5
+分子 #11: RNA_pol_L_2 domain-containing protein
+分子 #12: RNA polymerase II subunit K
+分子 #16: DNA excision repair protein ERCC-8
+分子 #17: DNA excision repair protein ERCC-6
+分子 #18: CSB element
+分子 #19: UV-stimulated scaffold protein A
+分子 #20: DNA damage-binding protein 1
+分子 #13: NTS
+分子 #14: TS
+分子 #15: RNA
+分子 #21: ZINC ION
+分子 #22: MAGNESIUM ION
-実験情報
-構造解析
| 手法 | クライオ電子顕微鏡法 |
|---|---|
 解析 解析 | 単粒子再構成法 |
| 試料の集合状態 | particle |
- 試料調製
試料調製
| 濃度 | 0.3 mg/mL |
|---|---|
| 緩衝液 | pH: 7.5 |
| グリッド | モデル: Quantifoil R2/1 / 支持フィルム - 材質: CARBON / 支持フィルム - トポロジー: HOLEY / 前処理 - タイプ: GLOW DISCHARGE |
| 凍結 | 凍結剤: ETHANE / チャンバー内湿度: 100 % / 装置: FEI VITROBOT MARK IV |
- 電子顕微鏡法
電子顕微鏡法
| 顕微鏡 | FEI TITAN KRIOS |
|---|---|
| 撮影 | フィルム・検出器のモデル: GATAN K3 (6k x 4k) / 撮影したグリッド数: 1 / 実像数: 10300 / 平均電子線量: 40.4 e/Å2 |
| 電子線 | 加速電圧: 300 kV / 電子線源:  FIELD EMISSION GUN FIELD EMISSION GUN |
| 電子光学系 | 照射モード: FLOOD BEAM / 撮影モード: BRIGHT FIELD |
| 実験機器 |  モデル: Titan Krios / 画像提供: FEI Company |
 ムービー
ムービー コントローラー
コントローラー
































 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)



































































































































 Trichoplusia ni (イラクサキンウワバ)
Trichoplusia ni (イラクサキンウワバ)