[English] 日本語
 Yorodumi
Yorodumi- EMDB-11624: Full-length structure of the substrate-free tyrosine hydroxylase ... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-11624 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Full-length structure of the substrate-free tyrosine hydroxylase (apo-TH). | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Tetramer / catecholamine / brain / Parkinson / OXIDOREDUCTASE | |||||||||
| Function / homology |  Function and homology information Function and homology informationtyrosine 3-monooxygenase / tyrosine 3-monooxygenase activity / dopamine biosynthetic process from tyrosine / embryonic camera-type eye morphogenesis / epinephrine biosynthetic process / serotonin biosynthetic process / Catecholamine biosynthesis / norepinephrine biosynthetic process / hyaloid vascular plexus regression / eye photoreceptor cell development ...tyrosine 3-monooxygenase / tyrosine 3-monooxygenase activity / dopamine biosynthetic process from tyrosine / embryonic camera-type eye morphogenesis / epinephrine biosynthetic process / serotonin biosynthetic process / Catecholamine biosynthesis / norepinephrine biosynthetic process / hyaloid vascular plexus regression / eye photoreceptor cell development / melanosome membrane / synaptic transmission, dopaminergic / mating behavior / dopamine biosynthetic process / eating behavior / regulation of heart contraction / pigmentation / smooth endoplasmic reticulum / anatomical structure morphogenesis / heart morphogenesis / visual perception / animal organ morphogenesis / learning / locomotory behavior / cognition / cytoplasmic side of plasma membrane / memory / synaptic vesicle / heart development / cytoplasmic vesicle / response to ethanol / perikaryon / response to hypoxia / neuron projection / iron ion binding / axon / perinuclear region of cytoplasm / enzyme binding / identical protein binding / nucleus / cytoplasm / cytosol Similarity search - Function | |||||||||
| Biological species |  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 4.1 Å | |||||||||
 Authors Authors | Bueno-Carrasco MT / Cuellar J | |||||||||
| Funding support |  Norway, Norway,  Spain, 2 items Spain, 2 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2022 Journal: Nat Commun / Year: 2022Title: Structural mechanism for tyrosine hydroxylase inhibition by dopamine and reactivation by Ser40 phosphorylation. Authors: María Teresa Bueno-Carrasco / Jorge Cuéllar / Marte I Flydal / César Santiago / Trond-André Kråkenes / Rune Kleppe / José R López-Blanco / Miguel Marcilla / Knut Teigen / Sara Alvira ...Authors: María Teresa Bueno-Carrasco / Jorge Cuéllar / Marte I Flydal / César Santiago / Trond-André Kråkenes / Rune Kleppe / José R López-Blanco / Miguel Marcilla / Knut Teigen / Sara Alvira / Pablo Chacón / Aurora Martinez / José M Valpuesta /    Abstract: Tyrosine hydroxylase (TH) catalyzes the rate-limiting step in the biosynthesis of dopamine (DA) and other catecholamines, and its dysfunction leads to DA deficiency and parkinsonisms. Inhibition by ...Tyrosine hydroxylase (TH) catalyzes the rate-limiting step in the biosynthesis of dopamine (DA) and other catecholamines, and its dysfunction leads to DA deficiency and parkinsonisms. Inhibition by catecholamines and reactivation by S40 phosphorylation are key regulatory mechanisms of TH activity and conformational stability. We used Cryo-EM to determine the structures of full-length human TH without and with DA, and the structure of S40 phosphorylated TH, complemented with biophysical and biochemical characterizations and molecular dynamics simulations. TH presents a tetrameric structure with dimerized regulatory domains that are separated 15 Å from the catalytic domains. Upon DA binding, a 20-residue α-helix in the flexible N-terminal tail of the regulatory domain is fixed in the active site, blocking it, while S40-phosphorylation forces its egress. The structures reveal the molecular basis of the inhibitory and stabilizing effects of DA and its counteraction by S40-phosphorylation, key regulatory mechanisms for homeostasis of DA and TH. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_11624.map.gz emd_11624.map.gz | 11.1 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-11624-v30.xml emd-11624-v30.xml emd-11624.xml emd-11624.xml | 18.8 KB 18.8 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_11624.png emd_11624.png | 41.4 KB | ||
| Filedesc metadata |  emd-11624.cif.gz emd-11624.cif.gz | 6 KB | ||
| Others |  emd_11624_half_map_1.map.gz emd_11624_half_map_1.map.gz emd_11624_half_map_2.map.gz emd_11624_half_map_2.map.gz | 19.4 MB 19.4 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-11624 http://ftp.pdbj.org/pub/emdb/structures/EMD-11624 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11624 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-11624 | HTTPS FTP |
-Related structure data
| Related structure data |  7a2gMC  6zn2C  6zvpC  6zzuC  7pimC C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_11624.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_11624.map.gz / Format: CCP4 / Size: 22.2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.05 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Half map: #1
| File | emd_11624_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
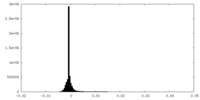
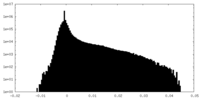
| Density Histograms |
-Half map: #2
| File | emd_11624_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
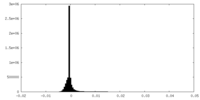
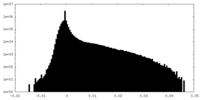
| Density Histograms |
- Sample components
Sample components
-Entire : Tyrosine hydroxylase
| Entire | Name: Tyrosine hydroxylase |
|---|---|
| Components |
|
-Supramolecule #1: Tyrosine hydroxylase
| Supramolecule | Name: Tyrosine hydroxylase / type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
-Macromolecule #1: Tyrosine 3-monooxygenase
| Macromolecule | Name: Tyrosine 3-monooxygenase / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO / EC number: tyrosine 3-monooxygenase |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 47.475598 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: GKAMLNLLFS PRATKPSALS RAVKVFETFE AKIHHLETRP AQRPRAGGPH LEYFVRLEVR RGDLAALLSG VRQVSEDVRS PAGPKVPWF PRKVSELDKC HHLVTKFDPD LDLDHPGFSD QVYRQRRKLI AEIAFQYRHG DPIPRVEYTA EEIATWKEVY T TLKGLYAT ...String: GKAMLNLLFS PRATKPSALS RAVKVFETFE AKIHHLETRP AQRPRAGGPH LEYFVRLEVR RGDLAALLSG VRQVSEDVRS PAGPKVPWF PRKVSELDKC HHLVTKFDPD LDLDHPGFSD QVYRQRRKLI AEIAFQYRHG DPIPRVEYTA EEIATWKEVY T TLKGLYAT HACGEHLEAF ALLERFSGYR EDNIPQLEDV SRFLKERTGF QLRPVAGLLS ARDFLASLAF RVFQCTQYIR HA SSPMHSP EPDCCHELLG HVPMLADRTF AQFSQDIGLA SLGASDEEIE KLSTLYWFTV EFGLCKQNGE VKAYGAGLLS SYG ELLHCL SEEPEIRAFD PEAAAVQPYQ DQTYQSVYFV SESFSDAKDK LRSYASRIQR PFSVKFDPYT LAIDVLDSPQ AVRR SLEGV QDELDTLAHA LSAIG UniProtKB: Tyrosine 3-monooxygenase |
-Macromolecule #2: FE (III) ION
| Macromolecule | Name: FE (III) ION / type: ligand / ID: 2 / Number of copies: 4 / Formula: FE |
|---|---|
| Molecular weight | Theoretical: 55.845 Da |
-Macromolecule #3: water
| Macromolecule | Name: water / type: ligand / ID: 3 / Number of copies: 4 / Formula: HOH |
|---|---|
| Molecular weight | Theoretical: 18.015 Da |
| Chemical component information |  ChemComp-HOH: |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7 Component:
| |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 95 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Quantum LS |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: COUNTING / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Number grids imaged: 1 / Number real images: 3867 / Average electron dose: 39.6 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: OTHER / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 3.0 µm / Nominal defocus min: 1.2 µm / Nominal magnification: 130000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller

















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)