+Search query
-Structure paper
| Title | Cooperation between bHLH transcription factors and histones for DNA access. |
|---|---|
| Journal, issue, pages | Nature, Vol. 619, Issue 7969, Page 385-393, Year 2023 |
| Publish date | Jul 5, 2023 |
 Authors Authors | Alicia K Michael / Lisa Stoos / Priya Crosby / Nikolas Eggers / Xinyu Y Nie / Kristina Makasheva / Martina Minnich / Kelly L Healy / Joscha Weiss / Georg Kempf / Simone Cavadini / Lukas Kater / Jan Seebacher / Luca Vecchia / Deyasini Chakraborty / Luke Isbel / Ralph S Grand / Florian Andersch / Jennifer L Fribourgh / Dirk Schübeler / Johannes Zuber / Andrew C Liu / Peter B Becker / Beat Fierz / Carrie L Partch / Jerome S Menet / Nicolas H Thomä /     |
| PubMed Abstract | The basic helix-loop-helix (bHLH) family of transcription factors recognizes DNA motifs known as E-boxes (CANNTG) and includes 108 members. Here we investigate how chromatinized E-boxes are engaged ...The basic helix-loop-helix (bHLH) family of transcription factors recognizes DNA motifs known as E-boxes (CANNTG) and includes 108 members. Here we investigate how chromatinized E-boxes are engaged by two structurally diverse bHLH proteins: the proto-oncogene MYC-MAX and the circadian transcription factor CLOCK-BMAL1 (refs. ). Both transcription factors bind to E-boxes preferentially near the nucleosomal entry-exit sites. Structural studies with engineered or native nucleosome sequences show that MYC-MAX or CLOCK-BMAL1 triggers the release of DNA from histones to gain access. Atop the H2A-H2B acidic patch, the CLOCK-BMAL1 Per-Arnt-Sim (PAS) dimerization domains engage the histone octamer disc. Binding of tandem E-boxes at endogenous DNA sequences occurs through direct interactions between two CLOCK-BMAL1 protomers and histones and is important for circadian cycling. At internal E-boxes, the MYC-MAX leucine zipper can also interact with histones H2B and H3, and its binding is indirectly enhanced by OCT4 elsewhere on the nucleosome. The nucleosomal E-box position and the type of bHLH dimerization domain jointly determine the histone contact, the affinity and the degree of competition and cooperativity with other nucleosome-bound factors. |
 External links External links |  Nature / Nature /  PubMed:37407816 / PubMed:37407816 /  PubMed Central PubMed Central |
| Methods | EM (single particle) |
| Resolution | 3.3 - 6.2 Å |
| Structure data |  EMDB-17154: Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (consensus and constituent map 1) EMDB-17155, PDB-8osj:  EMDB-17156: Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL-6.2 (DNA conformation 2) EMDB-17157, PDB-8osk:  EMDB-17158: Cryo-EM structure of CLOCK-BMAL1 bound to a nucleosomal E-box at position SHL+5.8 (constituent map 2 from additional focus classification on PAS domains)  EMDB-17159: Cryo-EM map of MYC-MAX-OCT4-LIN28 complex EMDB-17160, PDB-8osl:  EMDB-17161: Cryo-EM structure of CLOCK-BMAL1 bound to the native Por enhancer nucleosome (map 1)  EMDB-17162: MAX-MAX bound to a nucleosome at SHL+5.1 and SHL-6.9. EMDB-17183, PDB-8ots: EMDB-17184, PDB-8ott: |
| Chemicals | 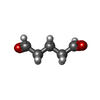 ChemComp-PTD: |
| Source |
|
 Keywords Keywords | GENE REGULATION / E-box / transcription factor / circadian clock / TRANSCRIPTION |
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About Yorodumi Papers
About Yorodumi Papers













 homo sapiens (human)
homo sapiens (human)

