-Search query
-Search result
Showing 1 - 50 of 118 items for (author: smith & dy)

EMDB-51365: 
Cryo-EM structure of human SLC45A4 in lipid nanodiscs
Method: single particle / : Markusson S, Newstead S

EMDB-51377: 
Cryo-EM structure of human SLC45A4 in detergent
Method: single particle / : Markusson S, Deme JC, Lea SM, Newstead S

PDB-9ghz: 
Cryo-EM structure of human SLC45A4 in lipid nanodiscs
Method: single particle / : Markusson S, Newstead S

PDB-9giu: 
Cryo-EM structure of human SLC45A4 in detergent
Method: single particle / : Markusson S, Deme JC, Lea SM, Newstead S

EMDB-45808: 
G115 gamma delta TCR/CD3 complex bound by OKT3 Fab
Method: single particle / : Hoque M, Saotome K, Franklin MC

EMDB-45810: 
G115 gamma delta TCR/CD3 complex bound by OKT3 Fab
Method: single particle / : Hoque M, Saotome K, Franklin MC

EMDB-45811: 
G115 gamma delta TCR/CD3 complex bound by OKT3 Fab
Method: single particle / : Hoque M, Saotome K, Franklin MC

EMDB-45814: 
Dimeric 9C2 gamma delta TCR bound by Fab 3
Method: single particle / : Hoque M, Saotome K, Franklin MC

PDB-9cq4: 
G115 gamma delta TCR/CD3 complex bound by OKT3 Fab
Method: single particle / : Hoque M, Saotome K, Franklin MC

PDB-9cq7: 
G115 TCR extracellular domain bound to Fab 1
Method: single particle / : Hoque M, Saotome K, Franklin MC

PDB-9cq8: 
Dimeric 9C2 gamma delta TCR extracellular domain bound by Fab 2
Method: single particle / : Hoque M, Saotome K, Franklin MC

PDB-9cql: 
Dimeric 9C2 gamma delta TCR bound by Fab 3
Method: single particle / : Hoque M, Saotome K, Franklin MC

EMDB-45764: 
CryoEM structure of the APO-BAM complex in DDM detergent
Method: single particle / : Sun D, Tegunov D, Payandeh J

EMDB-45765: 
CryoEM structure of BAM in complex with the PTB1 closed-state inhibitor (in DDM detergent)
Method: single particle / : Sun D, Tegunov D, Payandeh J

EMDB-45766: 
CryoEM structure of the APO-BAM complex in SMA nanodisc
Method: single particle / : Sun D, Tegunov D, Payandeh J

EMDB-45767: 
Structure of BAM complexed with PTB2 ligand in detergent
Method: single particle / : Sun D, Tegunov D, Payandeh J

EMDB-45768: 
CryoEM structure of BAM in complex with the PTB2 open-state inhibitor (in SMA nanodisc)
Method: single particle / : Sun D, Tegunov D, Payandeh J

PDB-9cnw: 
CryoEM structure of the APO-BAM complex in DDM detergent
Method: single particle / : Sun D, Tegunov D, Payandeh J

PDB-9cnx: 
CryoEM structure of BAM in complex with the PTB1 closed-state inhibitor (in DDM detergent)
Method: single particle / : Sun D, Tegunov D, Payandeh J

PDB-9cny: 
CryoEM structure of the APO-BAM complex in SMA nanodisc
Method: single particle / : Sun D, Tegunov D, Payandeh J

PDB-9cnz: 
Structure of BAM complexed with PTB2 ligand in detergent
Method: single particle / : Sun D, Tegunov D, Payandeh J

PDB-9co0: 
CryoEM structure of BAM in complex with the PTB2 open-state inhibitor (in SMA nanodisc)
Method: single particle / : Sun D, Tegunov D, Payandeh J

EMDB-19024: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

EMDB-19025: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

EMDB-19026: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

EMDB-19027: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb3: 
Structure of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb4: 
Structure of the five-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb5: 
Structure of the three-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

PDB-8rb7: 
Structure of the two-fold capsomer of the PNMA2 capsid
Method: single particle / : Erlendsson S, Xu J, Shepherd JD, Briggs JAG

EMDB-41153: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

EMDB-41154: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

PDB-8tcf: 
Integrin alpha-v beta-8 in complex with minibinder B8_BP_dsulf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

PDB-8tcg: 
Integrin alpha-v beta-6 in complex with minibinder B6_BP_dslf
Method: single particle / : Campbell MG, Fernandez A, Roy A, Kraft J, Baker D

EMDB-27847: 
Mouse norovirus strain CR6, attenuated
Method: single particle / : Smith TJ, Sherman M

EMDB-27849: 
Mouse norovirus strain CR6 at pH 5.0
Method: single particle / : Smith TJ

EMDB-27319: 
Mouse Norovirus strain WU23
Method: single particle / : Smith TJ

EMDB-27321: 
Mouse norovirus strain WU23 + 10mM GCDCA
Method: single particle / : Smith TJ

EMDB-27322: 
Murine norovirus strain WU23 at pH 5
Method: single particle / : Smith TJ

EMDB-31528: 
Ewald sphere-corrected Melbournevirus particle
Method: single particle / : Burton-Smith RN, Murata K

EMDB-31529: 
Fivefold block reconstruction of the Melbournevirus capsid
Method: single particle / : Burton-Smith RN, Murata K

EMDB-31530: 
Threefold block reconstruction of Melbournevirus capsid
Method: single particle / : Burton-Smith RN, Murata K

EMDB-31531: 
Twofold block reconstruction of the Melbournevirus capsid
Method: single particle / : Burton-Smith RN, Murata K

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K
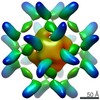
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit
Method: single particle / : Thomas PV, Smith C, Chen WH, Sankhala RS, Hajduczki A, Choe M, Martinez E, Chang W, Peterson CE, Karch C, Gohain N, Kannadka CB, de Val N, Joyce MG, Modjarrad K

EMDB-23970: 
Full length SARS-CoV-2 Nsp2
Method: single particle / : QCRG Structural Biology Consortium

EMDB-23971: 
SARS-CoV-2 Nsp2
Method: single particle / : QCRG Structural Biology Consortium

PDB-7msw: 
Full length SARS-CoV-2 Nsp2
Method: single particle / : QCRG Structural Biology Consortium
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
