-Search query
-Search result
Showing 1 - 50 of 214 items for (author: schmid & mf)

EMDB-49886: 
Cryo-ET map of the VZV capsid 3-fold axis.
Method: subtomogram averaging / : Oliver SL, Chen M

EMDB-49465: 
Reconstruction of the intranuclear varicella-zoster virus capsid.
Method: subtomogram averaging / : Oliver SL

EMDB-49466: 
Reconstruction of the varicella-zoster virus capsid vertex.
Method: subtomogram averaging / : Olver SL

EMDB-49467: 
Reconstruction of the intracellular varicella-zoster virus capsid with portal.
Method: subtomogram averaging / : Oliver SL

EMDB-49468: 
VZV portal vertex cryo-ET reconstruction.
Method: subtomogram averaging / : Oliver SL

EMDB-49469: 
VZV portal cryo-ET reconstruction.
Method: subtomogram averaging / : Oliver SL

EMDB-49470: 
Reconstruction of intracellular varicella zoster virus CAI-capsid with portal.
Method: subtomogram averaging / : Oliver SL

EMDB-49471: 
Reconstruction of the portal vertex from intracellular varicella-zoster virus CAI-capsids.
Method: subtomogram averaging / : Oliver SL

EMDB-49472: 
Reconstruction of the varicella-zoster virus C-capsid with the portal vertex.
Method: subtomogram averaging / : Oliver SL

EMDB-49473: 
VZV C-capsid portal vertex.
Method: subtomogram averaging / : Oliver SL

EMDB-47463: 
Apo TRiC in closed conformation
Method: single particle / : Zhao Y, Chiu W

EMDB-47464: 
TRiC encapsulating tubulin folding nucleus state.
Method: single particle / : Zhao Y, Chiu W

EMDB-47465: 
Tubulin intermediate state I in human chaperonin TRiC
Method: single particle / : Zhao Y, Chiu W

EMDB-47466: 
Tubulin intermediate state II in chaperonin TRiC
Method: single particle / : Zhao Y, Chiu W

EMDB-47467: 
Tubulin intermediate III in chaperonin TRiC
Method: single particle / : Zhao Y, Chiu W

EMDB-47468: 
Native Tubulin in human chaperonin TRIC
Method: single particle / : Zhao Y, Chiu W

EMDB-49591: 
Cryo-ET map of the VZV capsid vertex (5-fold axis).
Method: subtomogram averaging / : Oliver SL, Chen M

EMDB-45793: 
Closed state MmCpn under ATP/AlFx condition by subtomogram averaging
Method: subtomogram averaging / : Yanyan Z

EMDB-45794: 
MmCpn in closed state wit D9 symmetry
Method: subtomogram averaging / : Yanyan Z

EMDB-44925: 
Structure of the LM189-Bound beta2AR-Gi Complex
Method: single particle / : Casiraghi M, Wang H, Kobilka BK

EMDB-41346: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO DJ85-b.01 FAB
Method: single particle / : Pletnev S, Hoyt F, Fischer E, Kwong P

EMDB-41359: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO DJ85-c.01 FAB
Method: single particle / : Pletnev S, Hoyt F, Fischer E, Kwong P

EMDB-41360: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO DJ85-d.01 FAB
Method: single particle / : Pletnev S, Hoyt F, Fischer E, Kwong P

EMDB-41361: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO DJ85-e.01 FAB
Method: single particle / : Pletnev S, Hoyt F, Fischer E, Kwong P

EMDB-41362: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO HERH-c.01 FAB
Method: single particle / : Pletnev S, Hoyt F, Fischer E, Kwong P

EMDB-41426: 
Cryo-EM structure of TRNM-b*01 Fab in complex with HIV-1 Env trimer BG505.DS SOSIP
Method: single particle / : Roark RS, Morano NC, Shapiro LS, Kwong PD

EMDB-41438: 
Cryo-EM structure of HERH-b*01 Fab in complex with HIV-1 Env trimer BG505.DS SOSIP
Method: single particle / : Roark RS, Hoyt F, Hansen B, Fischer E, Shapiro LS, Kwong PD

EMDB-41440: 
Cryo-EM structure of TRNM-f*01 Fab in complex with HIV-1 Env trimer ConC SOSIP
Method: single particle / : Roark RS, Morano NC, Shapiro LS, Kwong PD

EMDB-41459: 
Cryo-EM structure of HIV-1 Env BG505 DS-SOSIP in complex with antibody GPZ6-b.01 targeting the fusion peptide
Method: single particle / : Zhou T, Morano NC, Roark RS, Kwong PD, Xu J

EMDB-41309: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO HERH-c.01 FAB
Method: single particle / : Morano NC, Hoyt F, Hansen B, Fischer E, Shapiro L

EMDB-41310: 
CRYO-EM STRUCTURE OF HIV-1 BG505DS-SOSIP.664 ENV TRIMER BOUND TO GPZ6-a.01 FAB
Method: single particle / : Morano NC, Becker JE, Shapiro L, Ho DD
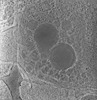
EMDB-28138: 
3D reconstruction of the apical complex of Plasmodium falciparum (3D7) free merozoite
Method: electron tomography / : Segev-Zarko L, Sun SY, Kim CY
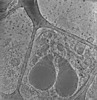
EMDB-28141: 
3D reconstruction of the apical complex of Plasmodium falciparum (3D7) free merozoite
Method: electron tomography / : Segev-Zarko L, Sun SY, Kim CY

EMDB-28142: 
3D Reconstruction of Plasmodium falciparum (3D7) free merozoite
Method: electron tomography / : Segev-Zarko L, Sun SY, Kim CY

EMDB-41840: 
Gaussian Mixture Models based single particle refinement - GPCR (Substance P bound to active human neurokinin 1 receptor in complex with miniGs399 from EMPIAR-10786)
Method: single particle / : Chen M

EMDB-41841: 
Gaussian mixture model based single particle refinement - SARS (SARS-CoV-2 Spike Proteins on intact virions from EMPIAR-10492)
Method: single particle / : Chen M

EMDB-41845: 
Gaussian mixture model based single particle refinement - ABC transporter (inhibitor-bound ABCG2 from EMPIAR-10374)
Method: single particle / : Chen M

EMDB-28125: 
Plasmodium falciparum merozoites apical 2-ring units
Method: subtomogram averaging / : Sun SY, Pintilie GD

EMDB-28126: 
Toxoplasma apical rings
Method: subtomogram averaging / : Sun SY, Pintilie GD

EMDB-29207: 
CryoET tomogram of mitochondria in BACHD mouse model neuron
Method: electron tomography / : Wu GH, Galaz-Montoya JG, Gu Y, Mitchell PG, Wu C, Chiu W

EMDB-29208: 
CryoET tomogram of BACHD mouse model neuron showing sheet aggregates
Method: electron tomography / : Wu GH, Galaz-Montoya JG, Gu Y, Mitchell PG, Wu C, Chiu W

EMDB-29210: 
CryoET tomogram of purified mitochondria from HD patient iPSC-derived neuron (Q109)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29211: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) with PIAS1 hetKO treatment
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-28668: 
CryoET tomogram of iPSC-derived control non-HD neuron (Q18)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
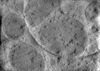
EMDB-28944: 
CryoET tomogram of iPSC-derived control non-HD neuron (Q20)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-28946: 
CryoET tomogram of HD patient iPSC-derived neuron (Q53)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29074: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
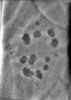
EMDB-29075: 
CryoET tomogram of HD patient iPSC-derived neuron (Q77)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
EMDB-29076: 
CryoET tomogram of HD patient iPSC-derived neuron (Q109)
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W

EMDB-29079: 
CryoET tomogram of HD patient iPSC-derived neuron (Q66) showing sheet aggregate
Method: electron tomography / : Wu GH, Smith-Geater C, Galaz-Montoya JG, Mitchell PG, Thompson LM, Chiu W
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
