-Search query
-Search result
Showing all 27 items for (author: sapiro & g)

EMDB-7770: 
Atomic resolution cryo-EM structure of beta-galactosidase
Method: single particle / : Subramaniam S, Bartesaghi A, Banerjee S, Zhu X

PDB-6cvm: 
Atomic resolution cryo-EM structure of beta-galactosidase
Method: single particle / : Subramaniam S, Bartesaghi A, Banerjee S, Zhu X, Milne JLS

EMDB-5682: 
Molecular structure of the native influenza hemagglutinin H1 on virions
Method: subtomogram averaging / : Harris AK, Meyerson JR, Matsuoka Y, Kuybeda O, Moran A, Bliss D, Das SR, Yewdell JW, Sapiro G, Subbarao K, Subramaniam S

EMDB-5683: 
Molecular structure of the native influenza hemagglutinin H1 on virions
Method: subtomogram averaging / : Harris AK, Meyerson JR, Matsuoka Y, Kuybeda O, Moran A, Bliss D, Das SR, Yewdell JW, Sapiro G, Subbarao K, Subramaniam S

EMDB-5684: 
Molecular structure of the native influenza hemagglutinin H1 on virions complexed with broadly neutralizing stem antibody C179
Method: subtomogram averaging / : Harris AK, Meyerson JR, Matsuoka Y, Kuybeda O, Moran A, Bliss D, Das SR, Yewdell JW, Sapiro G, Subbarao K, Subramaniam S

EMDB-5685: 
Molecular structure of the native influenza hemagglutinin H1 on virions complexed with broadly neutralizing stem antibody C179
Method: subtomogram averaging / : Harris AK, Meyerson JR, Matsuoka Y, Kuybeda O, Moran A, Bliss D, Das SR, Yewdell JW, Sapiro G, Subbarao K, Subramaniam S

EMDB-5686: 
Molecular structure of the native influenza hemagglutinin H3 on virions
Method: subtomogram averaging / : Harris AK, Meyerson JR, Matsuoka Y, Kuybeda O, Moran A, Bliss D, Das SR, Yewdell JW, Sapiro G, Subbarao K, Subramaniam S

EMDB-5687: 
Molecular structure of the native influenza hemagglutinin H3 on virions
Method: subtomogram averaging / : Harris AK, Meyerson JR, Matsuoka Y, Kuybeda O, Moran A, Bliss D, Das SR, Yewdell JW, Sapiro G, Subbarao K, Subramaniam S

EMDB-2221: 
GroEL at sub-nanometer resolution by Constrained Single Particle Tomography
Method: subtomogram averaging / : Bartesaghi A, Lecumberry F, Sapiro G, Subramaniam S

PDB-2ynj: 
GroEL at sub-nanometer resolution by Constrained Single Particle Tomography
Method: single particle / : Bartesaghi A, Lecumberry F, Sapiro G, Subramaniam S

EMDB-5455: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with soluble 2-domain CD4
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5456: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with 17b IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5457: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC01 IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5458: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC03 IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S
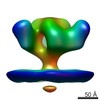
EMDB-5459: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC02 Fab
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5460: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC02 IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S
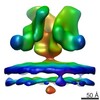
EMDB-5461: 
Cryo-electron tomography of native, trimeric HIV-1 BaL Env in complex with VRC01 IgG and 17b IgG
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5462: 
Cryo-electron tomography of a trimeric soluble HIV Env construct, gp140 SOSIP, in complex with Fab 17b
Method: subtomogram averaging / : Tran EEH, Borgnia MJ, Kuybeda O, Schauder DM, Bartesaghi A, Frank GA, Sapiro G, Milne JLS, Subramaniam S

EMDB-5018: 
Molecular Structure of the Native HIV-1 gp120 trimer bound to b12: Spike region
Method: subtomogram averaging / : Bartesaghi A, Borgnia M, Liu J, Sapiro G, Subramaniam S

EMDB-5019: 
Molecular Structure of Unliganded Native HIV-1 gp120 trimer: Spike region
Method: subtomogram averaging / : Bartesaghi A, Borgnia M, Liu J, Sapiro G, Subramaniam S

EMDB-5020: 
Molecular Structure of the Native HIV-1 gp120 trimer bound to CD4 and 17b: Spike region
Method: subtomogram averaging / : Bartesaghi A, Borgnia M, Liu J, Sapiro G, Subramaniam S

EMDB-5021: 
Molecular Structure of the Native HIV-1 gp120 trimer bound to b12: Membrane region
Method: subtomogram averaging / : Bartesaghi A, Borgnia M, Liu J, Sapiro G, Subramaniam S

EMDB-5022: 
Molecular Structure of Unliganded Native HIV-1 gp120 trimer: Membrane region
Method: subtomogram averaging / : Bartesaghi A, Borgnia M, Liu J, Sapiro G, Subramaniam S

EMDB-5023: 
Molecular Structure of the Native HIV-1 gp120 trimer bound to CD4 and 17b: Membrane region
Method: subtomogram averaging / : Bartesaghi A, Borgnia M, Liu J, Sapiro G, Subramaniam S

PDB-3dnl: 
Molecular structure for the HIV-1 gp120 trimer in the b12-bound state
Method: single particle / : Borgnia MJ, Liu J, Bartesaghi A, Sapiro G, Subramaniam S

PDB-3dnn: 
Molecular structure for the HIV-1 gp120 trimer in the unliganded state
Method: single particle / : Borgnia MJ, Liu J, Bartesaghi A, Sapiro G, Subramaniam S

PDB-3dno: 
Molecular structure for the HIV-1 gp120 trimer in the CD4-bound state
Method: single particle / : Borgnia MJ, Liu J, Bartesaghi A, Sapiro G, Subramaniam S
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
