-Search query
-Search result
Showing 1 - 50 of 197 items for (author: saha & s)

EMDB-36488: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Composite map)

EMDB-37212: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Receptor original map)

EMDB-37214: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Ligand/CCL7 focused map)

PDB-8jps: 
Structure of Duffy Antigen Receptor for Chemokines (DARC)/ACKR1 in complex with the chemokine, CCL7 (Composite map)

EMDB-17295: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

PDB-8oyt: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

EMDB-40825: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

PDB-8sx3: 
10E8-GT10.2 immunogen in complex with human Fab 10E8 and mouse Fab W6-10

EMDB-17296: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

PDB-8oyu: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '2 up 1 down' RBD conformation

EMDB-41024: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

EMDB-41025: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

EMDB-41026: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

EMDB-41027: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

EMDB-41034: 
MD64 N332-GT5 SOSIP

EMDB-41035: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

PDB-8t49: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

PDB-8t4a: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

PDB-8t4b: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

PDB-8t4d: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

PDB-8t4k: 
MD64 N332-GT5 SOSIP

PDB-8t4l: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

EMDB-18110: 
The fibrillar and amorphous states of polyQ Q97

EMDB-18114: 
phagophore in fibrillar polyQ
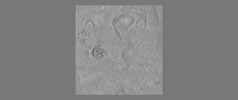
EMDB-18115: 
phagophore and lysosomes with amorphous polyQ

EMDB-18116: 
autolysosome, lysosome next to polyQ fibrils are empty

EMDB-18117: 
autophagosome and autolysosomes are empty next to fibrillar polyQ
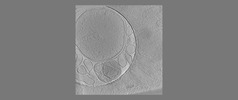
EMDB-18118: 
Isolated autophagosome

EMDB-37850: 
Cryo-EM structure of native H. thermoluteolus TH-1 GroEL

EMDB-37853: 
Cryo-EM structure of H. thermoluteolus GroEL-GroES2 football complex

EMDB-37862: 
Cryo-EM structure of H. thermophilus GroEL-GroES2 asymmetric football complex

EMDB-37863: 
Cryo-EM structure of H. thermophilus GroEL-GroES bullet complex

PDB-8wu4: 
Cryo-EM structure of native H. thermoluteolus TH-1 GroEL

PDB-8wuc: 
Cryo-EM structure of H. thermoluteolus GroEL-GroES2 football complex

PDB-8wuw: 
Cryo-EM structure of H. thermophilus GroEL-GroES2 asymmetric football complex

PDB-8wux: 
Cryo-EM structure of H. thermophilus GroEL-GroES bullet complex

EMDB-35817: 
Structure of Niacin-GPR109A-G protein complex

EMDB-35822: 
Structure of MK6892-GPR109A-G-protein complex

EMDB-35831: 
Structure of GSK256073-GPR109A-G-protein complex

EMDB-36193: 
Structure of Acipimox-GPR109A-G protein complex

EMDB-36280: 
Structure of MMF-GPR109A-G protein complex

PDB-8iy9: 
Structure of Niacin-GPR109A-G protein complex

PDB-8iyh: 
Structure of MK6892-GPR109A-G-protein complex

PDB-8iyw: 
Structure of GSK256073-GPR109A-G-protein complex

PDB-8jer: 
Structure of Acipimox-GPR109A-G protein complex

PDB-8jhn: 
Structure of MMF-GPR109A-G protein complex

EMDB-16626: 
Human heparan sulfate N-deacetylase-N-sulfotransferase 1 in complex with calcium, 3'-phosphoadenosine-5'-phosphosulfate, and nanobody nAb7 (original map)

EMDB-16627: 
Human heparan sulfate N-deacetylase-N-sulfotransferase 1 in complex with calcium, 3'-phosphoadenosine-5'-phosphosulfate, and nanobody nAb7 (locally refined map of N-terminal and deacetylase domains)

EMDB-16629: 
Human heparan sulfate N-deacetylase-N-sulfotransferase 1 in complex with calcium, 3'-phosphoadenosine-5'-phosphosulfate, and nanobody nAb7 (locally refined map of deacetylase and sulfotransferase domains)

EMDB-16661: 
Human heparan sulfate N-deacetylase-N-sulfotransferase 1 in complex with calcium, 3'-phosphoadenosine-5'-phosphosulfate, and nanobody nAb13 (original map)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
