-検索条件
-検索結果
検索 (著者・登録者: rheinberger & j)の結果140件中、1から50件目までを表示しています
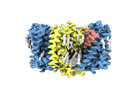
EMDB-18621: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)
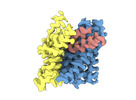
EMDB-18622: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)
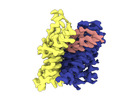
EMDB-18623: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)
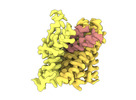
EMDB-18624: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)
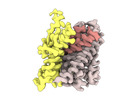
EMDB-18625: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)
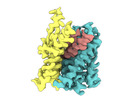
EMDB-18626: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)
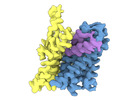
EMDB-18627: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)
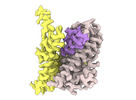
EMDB-18628: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)

PDB-8qro: 
ASCT2 trimer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS)

PDB-8qrp: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.1)

PDB-8qrq: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.2)

PDB-8qrr: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the outward-facing state (OFS.3)

PDB-8qrs: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-up)

PDB-8qru: 
ASCT2 protomer in lipid nanodiscs with bound glutamine and Na+ ions in the intermediate outward-facing state (iOFS-down)

PDB-8qrv: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the outward-facing state (OFS)

PDB-8qrw: 
ASCT2 protomer in lipid nanodiscs under low Na+ concentration in the intermediate outward-facing state (iOFS-up)

EMDB-17596: 
Ligand-free SpSLC9C1 in lipid nanodiscs, dimer

EMDB-17598: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 1

EMDB-17599: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 2

EMDB-17601: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 3

EMDB-17602: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 4

EMDB-17603: 
Ligand-free SpSLC9C1 in detergent, asymmetric class

EMDB-17604: 
Ligand-free SpSLC9C1 in detergent, symmetric class

EMDB-17605: 
cAMP-bound SpSLC9C1 in lipid nanodiscs, dimer

EMDB-17607: 
cAMP-bound SpSLC9C1 in lipid nanodiscs, protomer state 1

EMDB-17621: 
cGMP-bound SpSLC9C1 in lipid nanodiscs, dimer

EMDB-17622: 
cGMP-bound SpSLC9C1 in lipid nanodiscs, protomer

EMDB-17625: 
cAMP-bound SpSLC9C1 in lipid nanodiscs, protomer state 2

PDB-8pcz: 
Ligand-free SpSLC9C1 in lipid nanodiscs, dimer

PDB-8pd2: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 1

PDB-8pd3: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 2

PDB-8pd5: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 3

PDB-8pd7: 
Ligand-free SpSLC9C1 in lipid nanodiscs, protomer state 4

PDB-8pd8: 
cAMP-bound SpSLC9C1 in lipid nanodiscs, dimer

PDB-8pd9: 
cAMP-bound SpSLC9C1 in lipid nanodiscs, protomer state 1

PDB-8pdu: 
cGMP-bound SpSLC9C1 in lipid nanodiscs, dimer

PDB-8pdv: 
cGMP-bound SpSLC9C1 in lipid nanodiscs, protomer

EMDB-16120: 
Cryo-EM structure of the folate-specific ECF transporter complex in MSP2N2 lipid nanodiscs bound to ATP and ADP

EMDB-16121: 
Cryo-EM structure of the folate-specific ECF transporter complex in MSP2N2 lipid nanodiscs bound to AMP-PNP

EMDB-16122: 
Cryo-EM structure of the wild-type solitary ECF module in MSP2N2 lipid nanodiscs in the ATPase open and nucleotide-free conformation

EMDB-16123: 
Cryo-EM structure of the wild-type solitary ECF module in DDM micelles in the ATPase open and nucleotide-free conformation

EMDB-16124: 
Cryo-EM structure of the mutant solitary ECF module 2EQ in MSP2N2 lipid nanodiscs in the ATPase closed and ATP-bound conformation

PDB-8bmp: 
Cryo-EM structure of the folate-specific ECF transporter complex in MSP2N2 lipid nanodiscs bound to ATP and ADP

PDB-8bmq: 
Cryo-EM structure of the folate-specific ECF transporter complex in MSP2N2 lipid nanodiscs bound to AMP-PNP

PDB-8bmr: 
Cryo-EM structure of the wild-type solitary ECF module in MSP2N2 lipid nanodiscs in the ATPase open and nucleotide-free conformation

PDB-8bms: 
Cryo-EM structure of the mutant solitary ECF module 2EQ in MSP2N2 lipid nanodiscs in the ATPase closed and ATP-bound conformation
EMDB-14347: 
Cryo-EM map of the WT KdpFABC complex in the E2-P conformation, stabilised with the inhibitor orthovanadate
EMDB-14911: 
Cryo-EM map of the WT KdpFABC complex in the E1-P tight conformation, stabilised with the inhibitor orthovanadate
EMDB-14912: 
Cryo-EM map of the WT KdpFABC complex in the E1-P tight conformation, under turnover conditions
EMDB-14913: 
Cryo-EM map of the WT KdpFABC complex in the E1_ATPearly conformation, under turnover conditions
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
