-Search query
-Search result
Showing 1 - 50 of 175 items for (author: james & nr)

EMDB-41024: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

EMDB-41025: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

EMDB-41026: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

EMDB-41027: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

EMDB-41034: 
MD64 N332-GT5 SOSIP

EMDB-41035: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

PDB-8t49: 
MD65 N332-GT5 SOSIP in complex with RM_N332_03 Fab and RM20A3 Fab

PDB-8t4a: 
MD65 N332-GT5 SOSIP in complex with RM_N332_36 Fab and RM20A3 Fab

PDB-8t4b: 
MD65 N332-GT5 SOSIP in complex with RM_N332_32 Fab and RM20A3

PDB-8t4d: 
MD65 N332-GT5 SOSIP in complex with RM_N332_08 Fab and RM20A3 Fab

PDB-8t4k: 
MD64 N332-GT5 SOSIP

PDB-8t4l: 
MD65 N332-GT5 SOSIP in complex with RM_N332_07 Fab and RM20A3 Fab

EMDB-43290: 
Cryo-EM structure of Myxococcus xanthus EncA encapsulin shell loaded with EncD cargo
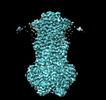
EMDB-40180: 
MsbA bound to cerastecin C

PDB-8gk7: 
MsbA bound to cerastecin C

EMDB-38095: 
MRE-269 bound Prostacyclin Receptor G protein complex
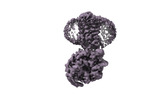
EMDB-38096: 
Treprostinil bound Prostacyclin Receptor G protein complex

PDB-8x79: 
MRE-269 bound Prostacyclin Receptor G protein complex

PDB-8x7a: 
Treprostinil bound Prostacyclin Receptor G protein complex

EMDB-28531: 
Cryo-EM structure of SARS-CoV-2 Spike trimer S2D14 in the 3-RBD Down conformation

EMDB-28532: 
Cryo-EM structure of SARS-CoV-2 Spike trimer S2D14 with two RBDs in the open conformation

EMDB-28533: 
Cryo-EM structure of SARS-CoV-2 Spike trimer S2D14 with two RBDs exposed

PDB-8epn: 
Cryo-EM structure of SARS-CoV-2 Spike trimer S2D14 in the 3-RBD Down conformation

PDB-8epp: 
Cryo-EM structure of SARS-CoV-2 Spike trimer S2D14 with two RBDs in the open conformation

PDB-8epq: 
Cryo-EM structure of SARS-CoV-2 Spike trimer S2D14 with two RBDs exposed

EMDB-29428: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% CHAPS, Fully Closed

EMDB-29430: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% CHAPS, One RBD Open

EMDB-29431: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% CHAPS, Two RBD Open

EMDB-29432: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% CHAPS, Three RBD Open

EMDB-29434: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% CHAPS, Fully Closed

EMDB-29435: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% CHAPS, One RBD Open

EMDB-29436: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% CHAPS, Two RBD Open

EMDB-29438: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% CHAPS, Three RBD Open

EMDB-29444: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% DDM, Fully Closed

EMDB-29445: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% DDM, One RBD Open

EMDB-29446: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.01% DDM, Two RBD Open

EMDB-29460: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% DDM, Fully Closed

EMDB-29461: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% DDM, One RBD Open

EMDB-29462: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% DDM, Two RBD Open

EMDB-29463: 
SARS-CoV-2 Spike D614G variant, pH 7.4, 0.5% DDM, Three RBD Open

EMDB-29464: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Fully Closed

EMDB-29465: 
SARS-CoV-2 Spike D614G variant, pH 5.0, One RBD Open (RBD observed)

EMDB-29466: 
SARS-CoV-2 Spike D614G variant, pH 5.0, One RBD Open (RBD scattered)

EMDB-29467: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Two RBD Open (two RBD observed)

EMDB-29468: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Two RBD Open (one RBD observed, one RBD scattered)

EMDB-29469: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Two RBD Open (two RBD scattered)

EMDB-29470: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Three RBD Open (three RBD observed)

EMDB-29471: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Three RBD Open (two RBD observed, one RBD scattered)

EMDB-29472: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Three RBD Open (one RBD observed, two RBD scattered)

EMDB-29473: 
SARS-CoV-2 Spike D614G variant, pH 5.0, Three RBD Open (three RBD scattered)
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
