-検索条件
-検索結果
検索 (著者・登録者: james & li)の結果2,285件中、1から50件目までを表示しています
EMDB-42950: 
Structure of CCP5 class1
EMDB-42951: 
Structure of CCP5 class2
EMDB-42952: 
Structure of CCP5 class3
EMDB-42971: 
CCP5 in complex with microtubules class1
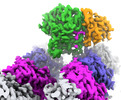
EMDB-42972: 
CCP5 in complex with microtubules class2
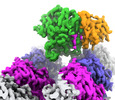
EMDB-42973: 
CCP5 in complex with microtubules class3

PDB-8v3q: 
Structure of CCP5 class1

PDB-8v3r: 
Structure of CCP5 class2

PDB-8v3s: 
Structure of CCP5 class3

PDB-8v4k: 
CCP5 in complex with microtubules class1

PDB-8v4l: 
CCP5 in complex with microtubules class2

PDB-8v4m: 
CCP5 in complex with microtubules class3

EMDB-18136: 
ATP-bound IstB in complex to duplex DNA

EMDB-18144: 
IstA-IstB(E167Q) Strand Transfer Complex

PDB-8q3w: 
ATP-bound IstB in complex to duplex DNA

PDB-8q4d: 
IstA-IstB(E167Q) Strand Transfer Complex

EMDB-41433: 
Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter

EMDB-41437: 
Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter

EMDB-41439: 
Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter

EMDB-41448: 
Escherichia coli RNA polymerase unwinding intermediate (I1c) at the lambda PR promoter

EMDB-41456: 
Escherichia coli RNA polymerase closed complex intermediate at the lambda PR promoter

PDB-8to1: 
Escherichia coli RNA polymerase unwinding intermediate (I1a) at the lambda PR promoter

PDB-8to6: 
Escherichia coli RNA polymerase unwinding intermediate (I1d) at the lambda PR promoter

PDB-8to8: 
Escherichia coli RNA polymerase unwinding intermediate (I1b) at the lambda PR promoter

PDB-8toe: 
Escherichia coli RNA polymerase unwinding intermediate (I1c) at the lambda PR promoter

PDB-8tom: 
Escherichia coli RNA polymerase closed complex intermediate at the lambda PR promoter

EMDB-17295: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

PDB-8oyt: 
Stabilised BA.1 SARS-CoV-2 spike with H6 nanobodies in '3 up' RBD conformation

EMDB-18438: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 1

EMDB-18439: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2

EMDB-18440: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 3

EMDB-18443: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 4

EMDB-18460: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1

EMDB-18461: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 2

PDB-8qrk: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 1

PDB-8qrl: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2

PDB-8qrm: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 3

PDB-8qrn: 
mt-SSU in GTPBP8 knock-out cells, state 4

PDB-8qu1: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1

PDB-8qu5: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 2
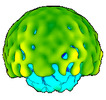
EMDB-50358: 
In vitro-induced genome-releasing intermediate of Rhodobacter microvirus Ebor computed with C5 symmetry

EMDB-41569: 
Cryo-EM structure of HmAb64 scFv in complex with CNE40 SOSIP trimer

PDB-8tr3: 
Cryo-EM structure of HmAb64 scFv in complex with CNE40 SOSIP trimer

EMDB-50356: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

EMDB-50357: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
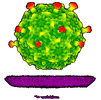
EMDB-50359: 
Rhodobacter microvirus Ebor attached to B10 host cell reconstructed by single particle analysis with applied C5 symmetry
EMDB-50360: 
Rhodobacter microvirus Ebor attached to the outer membrane vesicle

EMDB-50361: 
Rhodobacter microvirus Ebor attached to the host cell reconstructed by subtomogram averaging

PDB-9ffg: 
Empty capsid of Rhodobacter microvirus Ebor computed with I4 symmetry

PDB-9ffh: 
Native capsid of Rhodobacter microvirus Ebor computed with I4 symmetry
ページ:
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
