-Search query
-Search result
Showing 1 - 50 of 52 items for (author: gourdon & pe)

EMDB-18202: 
Copper-transporting ATPase HMA4 in E1 state apo

EMDB-18203: 
Copper-transporting ATPase HMA4 in E1 state with Cu
EMDB-18204: 
Copper-transporting ATPase HMA4 in E2P state with AlF

EMDB-18205: 
Copper-transporting ATPase HMA4 in E2P state with BeF

PDB-8q73: 
Copper-transporting ATPase HMA4 in E1 state apo

PDB-8q74: 
Copper-transporting ATPase HMA4 in E1 state with Cu

PDB-8q75: 
Copper-transporting ATPase HMA4 in E2P state with AlF

PDB-8q76: 
Copper-transporting ATPase HMA4 in E2P state with BeF

EMDB-16510: 
AQP7_inhibitor

PDB-8c9h: 
AQP7_inhibitor

EMDB-18464: 
Cryo-EM structure of MmpL3 from Mycobacterium smegmatis reconstituted into peptidiscs
EMDB-15527: 
AQP7 dimer of tetramers_C1

EMDB-15528: 
AQP7 dimer of tetramers_D4

PDB-8amw: 
AQP7 dimer of tetramers_C1

PDB-8amx: 
AQP7 dimer of tetramers_D4
EMDB-14745: 
Transient receptor potential cation channel subfamily V member 2,Enhanced green fluorescent protein

EMDB-14746: 
C16-2
EMDB-14747: 
Probenecid
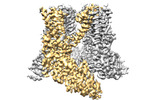
EMDB-14748: 
TRPV2-C16+Pro-2
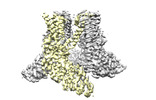
EMDB-14749: 
Transient receptor potential cation channel subfamily V member 2,Enhanced green fluorescent protein

PDB-7zjd: 
Transient receptor potential cation channel subfamily V member 2,Enhanced green fluorescent protein

PDB-7zje: 
C16-2

PDB-7zjg: 
Probenecid

PDB-7zjh: 
TRPV2-C16+Pro-2

PDB-7zji: 
Transient receptor potential cation channel subfamily V member 2,Enhanced green fluorescent protein
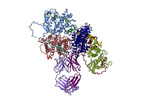
EMDB-14438: 
VAR2 complex with PAM1.4

EMDB-14446: 
VAR2CSA APO

PDB-7z12: 
VAR2 complex with PAM1.4

PDB-7z1h: 
VAR2CSA APO

EMDB-13606: 
Structure of CtAtm1 in the inward-facing open conformation

EMDB-13607: 
Structure of CtAtm1 in the occluded conformation with ATP bound

EMDB-13609: 
Structure of CtAtm1 in the inward-open with Glutathione-complexed [2Fe-2S] cluster bound
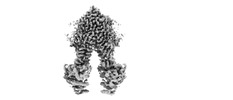
EMDB-13610: 
Structure of CtAtm1 in the inward-facing partially occluded with cargo bound

EMDB-13612: 
Structure of CtAtm1(E603Q) in the inward-facing open conformation

PDB-7pqx: 
Structure of CtAtm1 in the inward-facing open conformation

PDB-7pr1: 
Structure of CtAtm1 in the occluded conformation with ATP bound

PDB-7pro: 
Structure of CtAtm1 in the inward-open with Glutathione-complexed [2Fe-2S] cluster bound

PDB-7pru: 
Structure of CtAtm1 in the inward-facing partially occluded with cargo bound

PDB-7psd: 
Structure of CtAtm1(E603Q) in the inward-facing open conformation

EMDB-12018: 
VAR2CSA full ectodomain in present of plCS, DBL1-DBL4

EMDB-12477: 
Cryo-EM structure of VAR2CSA FCR3 domain DBL5/6

EMDB-12017: 
VAR2CSA full ectodomain

EMDB-4645: 
CryoEM structure of the human ClC-1 chloride channel, membrane domain

EMDB-4646: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 1

EMDB-4647: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 1

EMDB-4649: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 2

EMDB-4657: 
CryoEM structure of the human ClC-1 chloride channel, low pH

PDB-6qv6: 
CryoEM structure of the human ClC-1 chloride channel, membrane domain

PDB-6qvb: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 3

PDB-6qvc: 
CryoEM structure of the human ClC-1 chloride channel, CBS state 1
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
