-検索条件
-検索結果
検索 (著者・登録者: draczkowski & p)の結果全42件を表示しています

EMDB-33942: 
Cryo-EM structure of MERS-CoV spike protein, Two RBD-up conformation 2

EMDB-33943: 
Cryo-EM structure of MERS-CoV spike protein, Two RBD-up conformation 1

EMDB-33944: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 4

EMDB-33945: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 3

EMDB-33946: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 2

EMDB-33947: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 1

EMDB-33948: 
Cryo-EM structure of MERS-CoV spike protein, intermediate conformation

EMDB-33949: 
Cryo-EM structure of MERS-CoV spike protein, all RBD-down conformation

PDB-7ymt: 
Cryo-EM structure of MERS-CoV spike protein, Two RBD-up conformation 2

PDB-7ymv: 
Cryo-EM structure of MERS-CoV spike protein, Two RBD-up conformation 1

PDB-7ymw: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 4

PDB-7ymx: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 2

PDB-7ymy: 
Cryo-EM structure of MERS-CoV spike protein, One RBD-up conformation 1

PDB-7ymz: 
Cryo-EM structure of MERS-CoV spike protein, intermediate conformation

PDB-7yn0: 
Cryo-EM structure of MERS-CoV spike protein, all RBD-down conformation
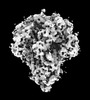
EMDB-32329: 
Cryo-EM map of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

EMDB-32332: 
Subtomogram averaging of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

EMDB-32333: 
Subtomogram averaging of PEDV (Pintung 52) S protein with one protomer in the D0-up conformation and two protomers in the D0-down conformation, determined in situ on intact viral particles

EMDB-32337: 
Subtomogram averaging of PEDV (Pintung 52) S protein with two protomers in the D0-up conformation and one protomer in the D0-down conformation, determined in situ on intact viral particles.

EMDB-32338: 
Cryo-EM map of PEDV S protein with one protomer in the D0-up conformation while the other two in the D0-down conformation
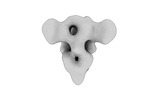
EMDB-32339: 
Subtomogram averaging of PEDV (Pintung 52) S protein with all three protomers in the D0-up conformation determined in situ on intact viral particles.

EMDB-32340: 
Subtomogram averaging of PEDV (Pintung 52) S protein in the postfusion form determined in situ on intact viral particles.

EMDB-33646: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with three D0-up

EMDB-33647: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein one D0-down and two D0-up

EMDB-33648: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-close conformation
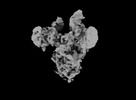
EMDB-33649: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-open conformation

EMDB-33700: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein with three D0-down
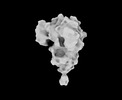
EMDB-33701: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein one D0-up and two D0-down

EMDB-33702: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S protein with three D0-up
EMDB-33703: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I with three D0-down

EMDB-33704: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I one D0-up and two D0-down
EMDB-33705: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I one D0-down and two D0-up
EMDB-33706: 
Cryo-EM map of HEK293F cell-derived PEDV PT52 S T326I with three D0-up

PDB-7w6m: 
Cryo-EM map of PEDV (Pintung 52) S protein with all three protomers in the D0-down conformation determined in situ on intact viral particles.

PDB-7w73: 
Cryo-EM map of PEDV S protein with one protomer in the D0-up conformation while the other two in the D0-down conformation

PDB-7y6s: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein with three D0-up

PDB-7y6t: 
Cryo-EM map of IPEC-J2 cell-derived PEDV PT52 S protein one D0-down and two D0-up

PDB-7y6u: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-close conformation

PDB-7y6v: 
Symmetry-expanded and locally refined protomer structure of IPEC-J2 cell-derived PEDV PT52 S with a CTD-open conformation

EMDB-30634: 
Cryo-EM structure of Ornithine transcarbamylase fused with Ubiquitin in complex with Ubiquitin-carboxy-hydrolase-L1 crosslinked with BS3

EMDB-9891: 
Cryo-EM structure of spike protein of feline infectious peritonitis virus strain UU4

PDB-6jx7: 
Cryo-EM structure of spike protein of feline infectious peritonitis virus strain UU4
 ムービー
ムービー コントローラー
コントローラー 構造ビューア
構造ビューア EMN検索について
EMN検索について



 wwPDBはEMDBデータモデルのバージョン3へ移行します
wwPDBはEMDBデータモデルのバージョン3へ移行します
