-Search query
-Search result
Showing 1 - 50 of 69 items for (author: case & jb)

EMDB-48323: 
Structure of the IFIT2-IFIT3 heterodimer from Mus musculus

EMDB-28198: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with LLNL-199

EMDB-28199: 
Cryo-EM map of SARS-CoV-2 Omicron BA.2 spike in complex with 2130-1-0114-112

EMDB-29530: 
SARS-CoV-2 XBB.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-29531: 
SARS-CoV-2 BQ.1.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-40240: 
SARS-CoV-2 BN.1 spike RBD bound to the human ACE2 ectodomain and the S309 neutralizing antibody Fab fragment

EMDB-26200: 
Cryo-EM structure of SARS-CoV-2 spike in complex with FSR22, an anti-SARS-CoV-2 DARPin

EMDB-26201: 
Cryo-EM structure of SARS-CoV-2 spike in complex with FSR22, an anti-SARS-CoV-2 DARPin (Local refinement of FSR22 and RBD)

EMDB-27749: 
Cryo-EM structure of SARS-CoV-2 RBD in complex with anti-SARS-CoV-2 DARPin,SR22, and two antibody Fabs, S309 and CR3022

EMDB-27750: 
Cryo-EM structure of SARS-CoV-2 RBD in complex with anti-SARS-CoV-2 DARPin,SR16m, and two antibody Fabs, S309 and CR3022

EMDB-24894: 
Ab16 Fab in complex with SARS-CoV-2 Spike (6P)

EMDB-24895: 
Ab20 in complex with SARS-CoV-2 spike (6P)

EMDB-26507: 
SARS-CoV-2 spike in complex with Multivalent miniprotein inhibitor FUS231-P24 (2RBDs open)

EMDB-26508: 
SARS-CoV-2 spike in complex with multivalent miniprotein inhibitor FUS231-P24 (3RBDs open)

EMDB-26509: 
SARS-CoV-2 spike in complex with multivalent miniprotein inhibitor FUS31-G10 (2RBDs open)

EMDB-26510: 
SARS-CoV-2 spike in complex with multivalent miniprotein inhibitor FUS31-G10 (3RBDs open)

EMDB-26511: 
SARS-CoV-2 spike in complex with AHB2-2GS-SB175 (local refinement of the RBD and AHB2)

EMDB-26512: 
SARS-CoV-2 spike in complex with AHB2-2GS-SB175

EMDB-25448: 
Negative-stain EM reconstruction of SpFN_1B-06-PL, a SARS-CoV-2 spike fused to H.pylori ferritin nanoparticle vaccine candidate
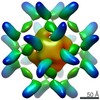
EMDB-25449: 
RFN_131, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Receptor-Binding Domain

EMDB-25450: 
pCoV146, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike Receptor-Binding and N-Terminal Domains

EMDB-25451: 
pCoV111, a Ferritin-based Nanoparticle Vaccine Candidate Displaying the SARS-CoV-2 Spike S1 Subunit

EMDB-13857: 
Beta049 fab in complex with SARS-CoV2 beta-Spike glycoprotein, The Beta mAb response underscores the antigenic distance to other SARS-CoV-2 variants

EMDB-13868: 
Beta-50 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13869: 
COVOX-222 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13870: 
Beta-43 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13871: 
Beta-26 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13872: 
Beta-32 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13873: 
Beta-53 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13874: 
Beta-44 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-13875: 
Beta-06 fab in complex with SARS-CoV-2 beta-Spike glycoprotein

EMDB-22748: 
SARS-CoV-2 Spike in complex with neutralizing Fab 2B04 (one up, two down conformation)

EMDB-22749: 
SARS-CoV-2 Spike RBD in complex with neutralizing Fab 2B04 (local refinement)

EMDB-22750: 
SARS-CoV-2 Spike in complex with neutralizing Fab 2H04 (three down conformation)

EMDB-22751: 
SARS-CoV-2 Spike RBD in complex with neutralizing Fab 2H04 (local refinement)

EMDB-22752: 
SARS-CoV-2 Spike in complex with neutralizing Fab 2B04 (two up, one down conformation)

EMDB-22753: 
SARS-CoV-2 Spike in complex with neutralizing Fab 2H04 (one up, two down conformation)

EMDB-23898: 
SARS-CoV-2 Spike in complex with neutralizing Fab SARS2-38 (three down conformation)

EMDB-23899: 
SARS-CoV-2 Spike RBD in complex with neutralizing Fab SARS2-38 (local refinement)

EMDB-12274: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab

EMDB-12275: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-88 Fab

EMDB-12276: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-150 Fab

EMDB-12277: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-40 Fab

EMDB-12278: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-316 Fab

EMDB-12279: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-384 Fab

EMDB-12280: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-253H55L Fab

EMDB-12281: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-253H55L Fab

EMDB-12282: 
EM structure of SARS-CoV-2 Spike glycoprotein in complex with COVOX-253H165L Fab

EMDB-12283: 
EM structure of SARS-CoV-2 Spike glycoprotein (all RBD down) in complex with COVOX-159

EMDB-12284: 
EM structure of SARS-CoV-2 Spike glycoprotein (one RBD up) in complex with COVOX-159
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
