-Search query
-Search result
Showing 1 - 50 of 276 items for (author: bart & sm)

EMDB-41248: 
Structure of AT118-H Nanobody Antagonist in Complex with the Angiotensin II Type I Receptor

EMDB-41249: 
Structure of AT118-L Nanobody Antagonist in Complex with the Angiotensin II Type I Receptor and Losartan

EMDB-44644: 
SARS CoV2 spike in complex with NTD-directed antibody Fab DH1052

EMDB-40853: 
CH505 Disulfide Stapled SOSIP Bound to b12 Fab
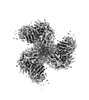
EMDB-40854: 
CH505 Disulfide Stapled SOSIP Bound to CH235.12 Fab

EMDB-41810: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-41820: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41823: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

EMDB-41838: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to partially open HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env

PDB-8u1d: 
Cryo-EM structure of vaccine-elicited CD4 binding site antibody DH1285 bound to HIV-1 CH505TFchim.6R.SOSIP.664v4.1 Env Local Refinement

EMDB-15964: 
Subtomogram averaging of SARS-CoV-2 nsp3-4 delta Ubl1-Mac1 obtained from cryo-ET of cryo-FIB milled VeroE6 cells transfected with nsp3-4 delta Ubl1-Mac1

EMDB-15965: 
Subtomogram average of SARS-CoV-2 nsp3-4 delate Ubl1-Ubl2 obtained from cryo-ET of cryo-FIB milled VeroE6 cells transfected with nsp3-4 delta Ubl1-Ubl2

EMDB-15963: 
Subtomogram average SARS-CoV-2 nsp3-4 from cryo-ET of VeroE6 expressing nsp3-4 (filtered to 20A resolution).

EMDB-15925: 
Cryo-electron tomogram acquired on cryo-lamella of VeroE6 cells transfected with SARS-CoV-2 HA-nsp3-4-V5, plunge frozen at 16 hpt

EMDB-15926: 
Cryo-electron tomogram of VeroE6 cells transfected with HA-nsp3-deltaUbl1-Ubl2-nsp4-V5, plunge frozen at 16 hpt

EMDB-15927: 
Cryo-electron tomogram of VeroE6 cells transfected with HA-nsp3-deltaUbl1-Mac1-nsp4-V5. Cells were plunge-frozen at 16h post-transfection and subjected to cryo-FIB milling.

EMDB-15928: 
Cryo-electron tomogram of VeroE6 cells transfected with HA-GG>AA-nsp4-V5, plunge-frozen at 16 hpt and subjected to cryo-FIB milling.

EMDB-15929: 
Cryo-electron tomogram of VeroE6 cells transfected with nsp3-4, vitrified by plunge-freezing at 16 hours post transfection and processed by cryo-FIB milling.

EMDB-27703: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace

PDB-8dtk: 
Structure of RBD directed antibody DH1047 in complex with SARS-CoV-2 spike: Local refinement of RBD-Fab interace
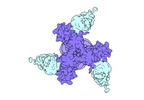
EMDB-27706: 
Vaccine elicited Antibody MU89 bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer
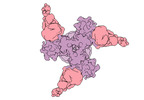
EMDB-27776: 
Vaccine elicited Antibody MU89+S27Y bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer

PDB-8dto: 
Vaccine elicited Antibody MU89 bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer

PDB-8dy6: 
Vaccine elicited Antibody MU89+S27Y bound to CH848.D949.10.17_N133D_N138T.DS.SOSIP.664 HIV-1 Env trimer

EMDB-40273: 
CryoEM structure of VRC01-CH848.0358.80

EMDB-40274: 
CryoEM structure of VRC01-CH848.0836.10

EMDB-40275: 
CryoEM structure of DH270.6-CH848.0526.25

EMDB-40277: 
CryoEM structure of DH270.6-CH848.10.17

EMDB-40278: 
CryoEM structure of DH270.5-CH848.10.17

EMDB-40279: 
CryoEM structure of VRC01-CH848.10.17

EMDB-40280: 
CryoEM structure of DH270.4-CH848.10.17

EMDB-40281: 
CryoEM structure of VRC01-CH848.0526.25

EMDB-40282: 
CryoEM structure of DH270.UCA.G57R-CH848.10.17DT

EMDB-40283: 
CryoEM structure of DH270.UCA-CH848.10.17DT

EMDB-40284: 
CryoEM structure of DH270.3-CH848.10.17

EMDB-40285: 
CryoEM structure of DH270.I5.6-CH848.10.17

EMDB-40286: 
CryoEM structure of DH270.I4.6-CH848.10.17

EMDB-40287: 
CryoEM structure of DH270.I3-CH848.10.17

EMDB-40288: 
CryoEM structure of DH270.I2-CH848.10.17

EMDB-40289: 
CryoEM structure of DH270.2-CH848.10.17

EMDB-40290: 
CryoEM structure of DH270.1-CH848.10.17

EMDB-40291: 
CryoEM structure of DH270.I1.6-CH848.10.17

EMDB-16480: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 10D12 heavy-chain-only antibody (3 RBDs up)

EMDB-16481: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 10D12 heavy-chain-only antibody (2 RBDs up)

EMDB-16490: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 10D12 heavy-chain-only antibody (local refinement)

PDB-8c8p: 
Structure of the SARS-CoV-2 spike glycoprotein in complex with the 10D12 heavy-chain-only antibody (local refinement)
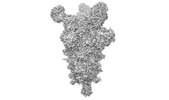
EMDB-26676: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment

EMDB-26677: 
Three RBD-down state of SARS-CoV-2 D614G spike in complex with the SP1-77 neutralizing antibody Fab fragment (local refinement of the RBD and Fab variable domains)

EMDB-26678: 
An antibody from single human VH-rearranging mouse neutralizes all SARS-CoV-2 variants through BA.5 by inhibiting membrane fusion

EMDB-27043: 
Negative stain electron microscopy single particle reconstruction of monoclonal antibody SP1-77 Fab in complex with SARS-CoV2 2P spike
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
