-Search query
-Search result
Showing 1 - 50 of 793 items for (author: bak & j)

EMDB-45655: 
Cryo-EM structure of alpha5beta1 integrin in complex with NeoNectin

EMDB-29618: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with apoRF3, RF1, P- and E-site tRNAPhe (State I-B)

EMDB-29619: 
ApoRF3 bound to an E. coli non-rotated ribosome termination complex, from focused classification and refinement (State I-B)

EMDB-29620: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with apoRF3, RF1, P- and E-site tRNAPhe (Composite state I-B)

EMDB-29621: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF1, P- and E-site tRNAPhe (State I-A)

EMDB-29625: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF3-GDPCP, RF1, P- and E-site tRNAPhe (State II-A)

EMDB-29626: 
RF3-GDPCP bound to an E. coli non-rotated ribosome termination complex, from focused classification and refinement (State II-A)

EMDB-29627: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF3-GDPCP, RF1, P- and E-site tRNAPhe (Composite state II-A)

EMDB-29628: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF1, P- and E-site tRNAPhe (State II-D)

EMDB-29629: 
Cryo-EM structure of an E. coli rotated ribosome bound with RF3-GDPCP and p/E-tRNAPhe (State II-B)

EMDB-29630: 
RF3-GDPCP bound to an E. coli rotated ribosome, from focused classification and refinement (State II-B)

EMDB-29631: 
Cryo-EM structure of an E. coli rotated ribosome bound with RF3-GDPCP and p/E-tRNAPhe (Composite state II-B)

EMDB-29632: 
Cryo-EM structure of an E. coli rotated ribosome bound with RF3-GDPCP and p/E-tRNAPhe (State II-C)

EMDB-29633: 
RF3-GDPCP bound to an E. coli rotated ribosome, from focused classification and refinement (State II-C)

EMDB-29634: 
Cryo-EM structure of an E. coli rotated ribosome bound with RF3-GDPCP and p/E-tRNAPhe (Composite state II-C)

PDB-8fzd: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with apoRF3, RF1, P- and E-site tRNAPhe (Composite state I-B)

PDB-8fze: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF1, P- and E-site tRNAPhe (State I-A)

PDB-8fzg: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF3-GDPCP, RF1, P- and E-site tRNAPhe (Composite state II-A)

PDB-8fzh: 
Cryo-EM structure of an E. coli non-rotated ribosome termination complex bound with RF1, P- and E-site tRNAPhe (State II-D)

PDB-8fzi: 
Cryo-EM structure of an E. coli rotated ribosome bound with RF3-GDPCP and p/E-tRNAPhe (Composite state II-B)

PDB-8fzj: 
Cryo-EM structure of an E. coli rotated ribosome bound with RF3-GDPCP and p/E-tRNAPhe (Composite state II-C)

EMDB-18438: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 1

EMDB-18439: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2

EMDB-18440: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 3

EMDB-18443: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 4

EMDB-18460: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1

EMDB-18461: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 2

PDB-8qrk: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 1

PDB-8qrl: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 2

PDB-8qrm: 
mt-SSU assembly intermediate in GTPBP8 knock-out cells, state 3

PDB-8qrn: 
mt-SSU in GTPBP8 knock-out cells, state 4

PDB-8qu1: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 1

PDB-8qu5: 
mt-LSU assembly intermediate in GTPBP8 knock-out cells, state 2

EMDB-29622: 
Cryo-EM structure of an E. coli rotated ribosome complex bound with RF3-ppGpp and p/E-tRNAPhe (State I-C)

EMDB-29623: 
RF3-ppGpp bound to an E. coli rotated ribosome, from focused classification and refinement (State I-C)
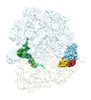
EMDB-29624: 
Cryo-EM structure of an E. coli rotated ribosome complex bound with RF3-ppGpp and p/E-tRNAPhe (Composite state I-C)

PDB-8fzf: 
Cryo-EM structure of an E. coli rotated ribosome complex bound with RF3-ppGpp and p/E-tRNAPhe (Composite state I-C)

EMDB-18592: 
E.coli DNA gyrase in complex with 217 bp substrate DNA and LEI-800

PDB-8qqi: 
E.coli DNA gyrase in complex with 217 bp substrate DNA and LEI-800

EMDB-42981: 
Prefusion-stabilized Respirovirus type 3 Fusion protein

PDB-8v5a: 
Prefusion-stabilized Respirovirus type 3 Fusion protein

EMDB-28966: 
CryoEM map of de novo designed oligomeric protein C4-71_6x

EMDB-28967: 
CryoEM map of de novo designed oligomeric protein C4-71_8x
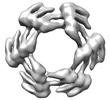
EMDB-28968: 
CryoEM map of de novo designed oligomeric protein C6-71

EMDB-28969: 
CryoEM map of de novo designed oligomeric protein C6-71_6x

EMDB-28970: 
CryoEM map of de novo designed oligomeric protein C6-71_8x

EMDB-28971: 
CryoEM map of de novo designed oligomeric protein C8-71_6x

EMDB-28972: 
CryoEM map of de novo designed oligomeric protein C8-71_8x
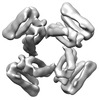
EMDB-28973: 
CryoEM map of de novo designed oligomeric protein C4-81

EMDB-28974: 
CryoEM map of designed oligomeric protein C4-71
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
