+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-8465 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | 3D cryo-EM reconstruction of MsbA-nanodisc with ADP | |||||||||
 Map data Map data | 3D cryo-EM density map of MsbA-nanodisc with ADP | |||||||||
 Sample Sample |
| |||||||||
| Function / homology |  Function and homology information Function and homology informationABC-type lipid A-core oligosaccharide transporter / ATPase-coupled lipid transmembrane transporter activity / ABC-type transporter activity / ATP hydrolysis activity / ATP binding / identical protein binding / plasma membrane Similarity search - Function | |||||||||
| Biological species |  | |||||||||
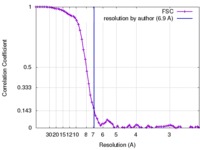
| Method | single particle reconstruction / cryo EM / Resolution: 6.9 Å | |||||||||
 Authors Authors | Mi W / Walz T / Liao M | |||||||||
 Citation Citation |  Journal: Nature / Year: 2017 Journal: Nature / Year: 2017Title: Structural basis of MsbA-mediated lipopolysaccharide transport. Authors: Wei Mi / Yanyan Li / Sung Hwan Yoon / Robert K Ernst / Thomas Walz / Maofu Liao /   Abstract: Lipopolysaccharide (LPS) in the outer membrane of Gram-negative bacteria is critical for the assembly of their cell envelopes. LPS synthesized in the cytoplasmic leaflet of the inner membrane is ...Lipopolysaccharide (LPS) in the outer membrane of Gram-negative bacteria is critical for the assembly of their cell envelopes. LPS synthesized in the cytoplasmic leaflet of the inner membrane is flipped to the periplasmic leaflet by MsbA, an ATP-binding cassette transporter. Despite substantial efforts, the structural mechanisms underlying MsbA-driven LPS flipping remain elusive. Here we use single-particle cryo-electron microscopy to elucidate the structures of lipid-nanodisc-embedded MsbA in three functional states. The 4.2 Å-resolution structure of the transmembrane domains of nucleotide-free MsbA reveals that LPS binds deep inside MsbA at the height of the periplasmic leaflet, establishing extensive hydrophilic and hydrophobic interactions with MsbA. Two sub-nanometre-resolution structures of MsbA with ADP-vanadate and ADP reveal an unprecedented closed and an inward-facing conformation, respectively. Our study uncovers the structural basis for LPS recognition, delineates the conformational transitions of MsbA to flip LPS, and paves the way for structural characterization of other lipid flippases. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_8465.map.gz emd_8465.map.gz | 25.2 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-8465-v30.xml emd-8465-v30.xml emd-8465.xml emd-8465.xml | 11.5 KB 11.5 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_8465_fsc.xml emd_8465_fsc.xml | 8.1 KB | Display |  FSC data file FSC data file |
| Images |  emd_8465.png emd_8465.png | 43.7 KB | ||
| Others |  emd_8465_additional.map.gz emd_8465_additional.map.gz | 20.7 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-8465 http://ftp.pdbj.org/pub/emdb/structures/EMD-8465 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8465 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-8465 | HTTPS FTP |
-Validation report
| Summary document |  emd_8465_validation.pdf.gz emd_8465_validation.pdf.gz | 78.4 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_8465_full_validation.pdf.gz emd_8465_full_validation.pdf.gz | 77.5 KB | Display | |
| Data in XML |  emd_8465_validation.xml.gz emd_8465_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8465 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8465 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8465 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-8465 | HTTPS FTP |
-Related structure data
| Related structure data |  8467C  8469C  8669C  8670C  8671C  5ttpC  5tv4C C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_8465.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_8465.map.gz / Format: CCP4 / Size: 27 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D cryo-EM density map of MsbA-nanodisc with ADP | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.23 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: 3D cryo-EM density map of MsbA-nanodisc with ADP,...
| File | emd_8465_additional.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | 3D cryo-EM density map of MsbA-nanodisc with ADP, without low-pass filter or amplitude modification | ||||||||||||
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : MsbA reconstituted in lipid nanodiscs
| Entire | Name: MsbA reconstituted in lipid nanodiscs |
|---|---|
| Components |
|
-Supramolecule #1: MsbA reconstituted in lipid nanodiscs
| Supramolecule | Name: MsbA reconstituted in lipid nanodiscs / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  Recombinant plasmid: pET-19b |
-Macromolecule #1: MsbA
| Macromolecule | Name: MsbA / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  |
| Sequence | String: MHNDKDLSTW QTFRRLWPTI APFKAGLIVA GVALILNAAS DTFMLSLLKP LLDDGFGKTD RSVLVWMPLV VIGLMILRGI TSYVSSYCIS WVSGKVVMTM RRRLFGHMMG MPVSFFDKQS TGTLLSRITY DSEQVASSSS GALITVVREG ASIIGLFIMM FYYSWQLSII ...String: MHNDKDLSTW QTFRRLWPTI APFKAGLIVA GVALILNAAS DTFMLSLLKP LLDDGFGKTD RSVLVWMPLV VIGLMILRGI TSYVSSYCIS WVSGKVVMTM RRRLFGHMMG MPVSFFDKQS TGTLLSRITY DSEQVASSSS GALITVVREG ASIIGLFIMM FYYSWQLSII LIVLAPIVSI AIRVVSKRFR NISKNMQNTM GQVTTSAEQM LKGHKEVLIF GGQEVETKRF DKVSNRMRLQ GMKMVSASSI SDPIIQLIAS LALAFVLYAA SFPSVMDSLT AGTITVVFSS MIALMRPLKS LTNVNAQFQR GMAACQTLFT ILDSEQEKDE GKRVIERATG DVEFRNVTFT YPGRDVPALR NINLKIPAGK TVALVGRSGS GKSTIASLIT RFYDIDEGEI LMDGHDLREY TLASLRNQVA LVSQNVHLFN DTVANNIAYA RTEQYSREQI EEAARMAYAM DFINKMDNGL DTVIGENGVL LSGGQRQRIA IARALLRDSP ILILDEATSA LDTESERAIQ AALDELQKNR TSLVIAHRLS TIEKADEIVV VEDGVIVERG THNDLLEHRG VYAQLHKMQF GQ |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.4 |
|---|---|
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI POLARA 300 |
|---|---|
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Average electron dose: 47.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Tecnai Polara / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller











 Z
Z Y
Y X
X