+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-9193 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of Plasmodium falciparum CyRPA/Ripr invasion complex | |||||||||
 Map data Map data | Cryo-electron microscopy structure of Plasmodium falciparum CyRPA/PfRipr complex | |||||||||
 Sample Sample |
| |||||||||
| Biological species |  | |||||||||
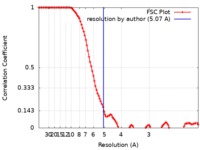
| Method | single particle reconstruction / cryo EM / Resolution: 5.07 Å | |||||||||
 Authors Authors | Wilson W / Alan C / Zhiheng Y | |||||||||
 Citation Citation |  Journal: Nature / Year: 2019 Journal: Nature / Year: 2019Title: Structure of Plasmodium falciparum Rh5-CyRPA-Ripr invasion complex. Authors: Wilson Wong / Rick Huang / Sebastien Menant / Chuan Hong / Jarrod J Sandow / Richard W Birkinshaw / Julie Healer / Anthony N Hodder / Usheer Kanjee / Christopher J Tonkin / Denise Heckmann / ...Authors: Wilson Wong / Rick Huang / Sebastien Menant / Chuan Hong / Jarrod J Sandow / Richard W Birkinshaw / Julie Healer / Anthony N Hodder / Usheer Kanjee / Christopher J Tonkin / Denise Heckmann / Vladislav Soroka / Teit Max Moscote Søgaard / Thomas Jørgensen / Manoj T Duraisingh / Peter E Czabotar / Willem A de Jongh / Wai-Hong Tham / Andrew I Webb / Zhiheng Yu / Alan F Cowman /    Abstract: Plasmodium falciparum causes the severe form of malaria that has high levels of mortality in humans. Blood-stage merozoites of P. falciparum invade erythrocytes, and this requires interactions ...Plasmodium falciparum causes the severe form of malaria that has high levels of mortality in humans. Blood-stage merozoites of P. falciparum invade erythrocytes, and this requires interactions between multiple ligands from the parasite and receptors in hosts. These interactions include the binding of the Rh5-CyRPA-Ripr complex with the erythrocyte receptor basigin, which is an essential step for entry into human erythrocytes. Here we show that the Rh5-CyRPA-Ripr complex binds the erythrocyte cell line JK-1 significantly better than does Rh5 alone, and that this binding occurs through the insertion of Rh5 and Ripr into host membranes as a complex with high molecular weight. We report a cryo-electron microscopy structure of the Rh5-CyRPA-Ripr complex at subnanometre resolution, which reveals the organization of this essential invasion complex and the mode of interactions between members of the complex, and shows that CyRPA is a critical mediator of complex assembly. Our structure identifies blades 4-6 of the β-propeller of CyRPA as contact sites for Rh5 and Ripr. The limited contacts between Rh5-CyRPA and CyRPA-Ripr are consistent with the dissociation of Rh5 and Ripr from CyRPA for membrane insertion. A comparision of the crystal structure of Rh5-basigin with the cryo-electron microscopy structure of Rh5-CyRPA-Ripr suggests that Rh5 and Ripr are positioned parallel to the erythrocyte membrane before membrane insertion. This provides information on the function of this complex, and thereby provides insights into invasion by P. falciparum. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_9193.map.gz emd_9193.map.gz | 95.7 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-9193-v30.xml emd-9193-v30.xml emd-9193.xml emd-9193.xml | 9.7 KB 9.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_9193_fsc.xml emd_9193_fsc.xml | 12.8 KB | Display |  FSC data file FSC data file |
| Images |  emd_9193.png emd_9193.png | 93.9 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-9193 http://ftp.pdbj.org/pub/emdb/structures/EMD-9193 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9193 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-9193 | HTTPS FTP |
-Validation report
| Summary document |  emd_9193_validation.pdf.gz emd_9193_validation.pdf.gz | 78.7 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_9193_full_validation.pdf.gz emd_9193_full_validation.pdf.gz | 77.8 KB | Display | |
| Data in XML |  emd_9193_validation.xml.gz emd_9193_validation.xml.gz | 494 B | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9193 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9193 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9193 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-9193 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_9193.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_9193.map.gz / Format: CCP4 / Size: 103 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | Cryo-electron microscopy structure of Plasmodium falciparum CyRPA/PfRipr complex | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.04 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
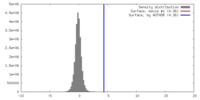
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : CyRPR/PfRipr complex
| Entire | Name: CyRPR/PfRipr complex |
|---|---|
| Components |
|
-Supramolecule #1: CyRPR/PfRipr complex
| Supramolecule | Name: CyRPR/PfRipr complex / type: complex / ID: 1 / Parent: 0 |
|---|---|
| Source (natural) | Organism:  |
| Recombinant expression | Organism:  unidentified baculovirus unidentified baculovirus |
| Molecular weight | Theoretical: 163 kDa/nm |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8.5 / Details: 20 mM Tris, pH8.5 150 mM NaCl |
|---|---|
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 277 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Spherical aberration corrector: Microscope was equipped with a Cs corrector with two hexapole elements Energy filter - Name: GIF Quantum LS / Energy filter - Slit width: 20 eV |
| Image recording | Film or detector model: GATAN K2 SUMMIT (4k x 4k) / Detector mode: SUPER-RESOLUTION / Digitization - Dimensions - Width: 3838 pixel / Digitization - Dimensions - Height: 3710 pixel / Digitization - Sampling interval: 5.0 µm / Number grids imaged: 4 / Number real images: 12974 / Average exposure time: 10.0 sec. / Average electron dose: 92.5 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 70.0 µm / Calibrated magnification: 48077 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 0.01 mm / Nominal magnification: 105000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller














 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)