[English] 日本語
 Yorodumi
Yorodumi- EMDB-45675: Cryo-EM model derived from localized reconstruction of human aden... -
+ Open data
Open data
- Basic information
Basic information
| Entry | 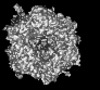 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM model derived from localized reconstruction of human adenovirus (Ad5)-hexon-FX complex at 3.6A resolution | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Adenovirus / Hexon / Coagulation factor X / Coagulation factor II / Prothrombin / Complex / Interactions / VIRUS | |||||||||
| Function / homology |  Function and homology information Function and homology informationT=25 icosahedral viral capsid / coagulation factor Xa / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / microtubule-dependent intracellular transport of viral material towards nucleus / : / positive regulation of TOR signaling / Transport of gamma-carboxylated protein precursors from the endoplasmic reticulum to the Golgi apparatus / : ...T=25 icosahedral viral capsid / coagulation factor Xa / Defective factor IX causes thrombophilia / Defective cofactor function of FVIIIa variant / Defective F9 variant does not activate FX / microtubule-dependent intracellular transport of viral material towards nucleus / : / positive regulation of TOR signaling / Transport of gamma-carboxylated protein precursors from the endoplasmic reticulum to the Golgi apparatus / : / Gamma-carboxylation of protein precursors / Removal of aminoterminal propeptides from gamma-carboxylated proteins / : / phospholipid binding / Golgi lumen / blood coagulation / viral capsid / host cell / positive regulation of cell migration / endoplasmic reticulum lumen / serine-type endopeptidase activity / external side of plasma membrane / calcium ion binding / symbiont entry into host cell / host cell nucleus / structural molecule activity / proteolysis / : / extracellular region / plasma membrane Similarity search - Function | |||||||||
| Biological species |   Human adenovirus 5 / Human adenovirus 5 /  Homo sapiens (human) Homo sapiens (human) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.61 Å | |||||||||
 Authors Authors | Reddy VS / Ma OX | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2024 Journal: Nat Commun / Year: 2024Title: Structure-derived insights from blood factors binding to the surfaces of different adenoviruses. Authors: Haley E Mudrick / Shao-Chia Lu / Janarjan Bhandari / Mary E Barry / Jack R Hemsath / Felix G M Andres / Olivia X Ma / Michael A Barry / Vijay S Reddy /  Abstract: The tropism of adenoviruses (Ads) is significantly influenced by the binding of various blood factors. To investigate differences in their binding, we conducted cryo-EM analysis on complexes of ...The tropism of adenoviruses (Ads) is significantly influenced by the binding of various blood factors. To investigate differences in their binding, we conducted cryo-EM analysis on complexes of several human adenoviruses with human platelet factor-4 (PF4), coagulation factors FII (Prothrombin), and FX. While we observed EM densities for FII and FX bound to all the species-C adenoviruses examined, no densities were seen for PF4, even though PF4 can co-pellet with various Ads. Similar to FX, the γ-carboxyglutamic acid (Gla) domain of FII binds within the surface cavity of hexon trimers. While FII binds equally to species-C Ads: Ad5, Ad6, and Ad657, FX exhibits significantly better binding to Ad5 and Ad657 compared to Ad6. Although only the FX-Gla domain is observed at high-resolution (3.7 Å), the entire FX is visible at low-resolution bound to Ad5 in three equivalent binding modes consistent with the 3-fold symmetric hexon. Only the Gla and kringle-1 domains of FII are visible on all the species-C adenoviruses, where the rigid FII binds in an upright fashion, in contrast to the flexible and bent FX. These data suggest that differential binding of FII and FX may shield certain species-C adenoviruses differently against immune molecules, thereby modulating their tropism. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Supplemental images |
|---|
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_45675.map.gz emd_45675.map.gz | 719.1 KB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-45675-v30.xml emd-45675-v30.xml emd-45675.xml emd-45675.xml | 20.2 KB 20.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_45675_fsc.xml emd_45675_fsc.xml | 7.2 KB | Display |  FSC data file FSC data file |
| Images |  emd_45675.png emd_45675.png | 52 KB | ||
| Filedesc metadata |  emd-45675.cif.gz emd-45675.cif.gz | 7.1 KB | ||
| Others |  emd_45675_half_map_1.map.gz emd_45675_half_map_1.map.gz emd_45675_half_map_2.map.gz emd_45675_half_map_2.map.gz | 23 MB 23 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-45675 http://ftp.pdbj.org/pub/emdb/structures/EMD-45675 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45675 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-45675 | HTTPS FTP |
-Related structure data
| Related structure data |  9cliMC  9clnC  9clsC  9cm2C  9cm9C  9cmoC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data | Similarity search - Function & homology  F&H Search F&H Search |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_45675.map.gz / Format: CCP4 / Size: 4.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_45675.map.gz / Format: CCP4 / Size: 4.3 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. generated in cubic-lattice coordinate | ||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.408 Å | ||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
|
-Supplemental data
-Half map: #2
| File | emd_45675_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_45675_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : Human adenovirus 5 (Ad5) in complex with Coagulation factor FX
| Entire | Name: Human adenovirus 5 (Ad5) in complex with Coagulation factor FX |
|---|---|
| Components |
|
-Supramolecule #1: Human adenovirus 5 (Ad5) in complex with Coagulation factor FX
| Supramolecule | Name: Human adenovirus 5 (Ad5) in complex with Coagulation factor FX type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#2 / Details: Localized reconstruction. |
|---|---|
| Source (natural) | Organism:   Human adenovirus 5 Human adenovirus 5 |
| Molecular weight | Theoretical: 400 KDa |
-Supramolecule #2: Hexon protein
| Supramolecule | Name: Hexon protein / type: organelle_or_cellular_component / ID: 2 / Parent: 1 / Macromolecule list: #1 |
|---|---|
| Source (natural) | Organism:   Human adenovirus 5 Human adenovirus 5 |
-Supramolecule #3: Human Coagulation Factor X
| Supramolecule | Name: Human Coagulation Factor X / type: organelle_or_cellular_component / ID: 3 / Parent: 2 / Macromolecule list: #2 |
|---|---|
| Source (natural) | Organism: |
-Macromolecule #1: Hexon protein
| Macromolecule | Name: Hexon protein / type: protein_or_peptide / ID: 1 / Details: Adenovirus hexon protein / Number of copies: 3 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism:   Human adenovirus 5 Human adenovirus 5 |
| Molecular weight | Theoretical: 108.107617 KDa |
| Sequence | String: MATPSMMPQW SYMHISGQDA SEYLSPGLVQ FARATETYFS LNNKFRNPTV APTHDVTTDR SQRLTLRFIP VDREDTAYSY KARFTLAVG DNRVLDMAST YFDIRGVLDR GPTFKPYSGT AYNALAPKGA PNPCEWDEAA TALEINLEEE DDDNEDEVDE Q AEQQKTHV ...String: MATPSMMPQW SYMHISGQDA SEYLSPGLVQ FARATETYFS LNNKFRNPTV APTHDVTTDR SQRLTLRFIP VDREDTAYSY KARFTLAVG DNRVLDMAST YFDIRGVLDR GPTFKPYSGT AYNALAPKGA PNPCEWDEAA TALEINLEEE DDDNEDEVDE Q AEQQKTHV FGQAPYSGIN ITKEGIQIGV EGQTPKYADK TFQPEPQIGE SQWYETEINH AAGRVLKKTT PMKPCYGSYA KP TNENGGQ GILVKQQNGK LESQVEMQFF STTEATAGNG DNLTPKVVLY SEDVDIETPD THISYMPTIK EGNSRELMGQ QSM PNRPNY IAFRDNFIGL MYYNSTGNMG VLAGQASQLN AVVDLQDRNT ELSYQLLLDS IGDRTRYFSM WNQAVDSYDP DVRI IENHG TEDELPNYCF PLGGVINTET LTKVKPKTGQ ENGWEKDATE FSDKNEIRVG NNFAMEINLN ANLWRNFLYS NIALY LPDK LKYSPSNVKI SDNPNTYDYM NKRVVAPGLV DCYINLGARW SLDYMDNVNP FNHHRNAGLR YRSMLLGNGR YVPFHI QVP QKFFAIKNLL LLPGSYTYEW NFRKDVNMVL QSSLGNDLRV DGASIKFDSI CLYATFFPMA HNTASTLEAM LRNDTND QS FNDYLSAANM LYPIPANATN VPISIPSRNW AAFRGWAFTR LKTKETPSLG SGYDPYYTYS GSIPYLDGTF YLNHTFKK V AITFDSSVSW PGNDRLLTPN EFEIKRSVDG EGYNVAQCNM TKDWFLVQML ANYNIGYQGF YIPESYKDRM YSFFRNFQP MSRQVVDDTK YKDYQQVGIL HQHNNSGFVG YLAPTMREGQ AYPANFPYPL IGKTAVDSIT QKKFLCDRTL WRIPFSSNFM SMGALTDLG QNLLYANSAH ALDMTFEVDP MDEPTLLYVL FEVFDVVRVH RPHRGVIETV YLRTPFSAGN ATT UniProtKB: Hexon protein |
-Macromolecule #2: Coagulation factor X
| Macromolecule | Name: Coagulation factor X / type: protein_or_peptide / ID: 2 / Number of copies: 1 / Enantiomer: LEVO / EC number: coagulation factor Xa |
|---|---|
| Source (natural) | Organism:  Homo sapiens (human) Homo sapiens (human) |
| Molecular weight | Theoretical: 55.286738 KDa |
| Sequence | String: MGRPLHLVLL SASLAGLLLL GESLFIRREQ ANNILARVTR ANSFL(CGU)(CGU)MKK GHL(CGU)R(CGU)CM(CGU) (CGU)TCSY(CGU)(CGU)AR(CGU) VF(CGU)DSDKTN(CGU) FWNKYKDGDQ CETSPCQNQG KCKDGLGEYT CTCLEG FEG KNCELFTRKL ...String: MGRPLHLVLL SASLAGLLLL GESLFIRREQ ANNILARVTR ANSFL(CGU)(CGU)MKK GHL(CGU)R(CGU)CM(CGU) (CGU)TCSY(CGU)(CGU)AR(CGU) VF(CGU)DSDKTN(CGU) FWNKYKDGDQ CETSPCQNQG KCKDGLGEYT CTCLEG FEG KNCELFTRKL CSLDNGDCDQ FCHEEQNSVV CSCARGYTLA DNGKACIPTG PYPCGKQTLE RRKRSVAQAT SSSGEAP DS ITWKPYDAAD LDPTENPFDL LDFNQTQPER GDNNLTRIVG GQECKDGECP WQALLINEEN EGFCGGTILS EFYILTAA H CLYQAKRFKV RVGDRNTEQE EGGEAVHEVE VVIKHNRFTK ETYDFDIAVL RLKTPITFRM NVAPACLPER DWAESTLMT QKTGIVSGFG RTHEKGRQST RLKMLEVPYV DRNSCKLSSS FIITQNMFCA GYDTKQEDAC QGDSGGPHVT RFKDTYFVTG IVSWGEGCA RKGKYGIYTK VTAFLKWIDR SMKTRGLPKA KSHAPEVITS SPLK UniProtKB: Coagulation factor X |
-Macromolecule #3: CALCIUM ION
| Macromolecule | Name: CALCIUM ION / type: ligand / ID: 3 / Number of copies: 7 / Formula: CA |
|---|---|
| Molecular weight | Theoretical: 40.078 Da |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 2 mg/mL |
|---|---|
| Buffer | pH: 7.2 / Component - Concentration: 20.0 mM / Component - Name: Hepes |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 300 / Pretreatment - Type: GLOW DISCHARGE / Pretreatment - Time: 60 sec. |
| Vitrification | Cryogen name: ETHANE |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K3 BIOCONTINUUM (6k x 4k) / Number grids imaged: 3 / Number real images: 10000 / Average electron dose: 81.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 100.0 µm / Calibrated defocus max: 5.0 µm / Calibrated defocus min: 1.0 µm / Calibrated magnification: 81000 / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 4.0 µm / Nominal defocus min: 2.5 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller




















 Y (Sec.)
Y (Sec.) X (Row.)
X (Row.) Z (Col.)
Z (Col.)






































