[English] 日本語
 Yorodumi
Yorodumi- EMDB-22305: CryoEM structure of designed helical fusion protein C4_nat_HFuse-7900 -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-22305 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | CryoEM structure of designed helical fusion protein C4_nat_HFuse-7900 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | helical bundle / helical repeat / DE NOVO PROTEIN | |||||||||
| Biological species | synthetic construct (others) | |||||||||
| Method | single particle reconstruction / cryo EM / Resolution: 3.7 Å | |||||||||
 Authors Authors | Redler RL / Edman NI | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Design of multi-scale protein complexes by hierarchical building block fusion. Authors: Yang Hsia / Rubul Mout / William Sheffler / Natasha I Edman / Ivan Vulovic / Young-Jun Park / Rachel L Redler / Matthew J Bick / Asim K Bera / Alexis Courbet / Alex Kang / T J Brunette / Una ...Authors: Yang Hsia / Rubul Mout / William Sheffler / Natasha I Edman / Ivan Vulovic / Young-Jun Park / Rachel L Redler / Matthew J Bick / Asim K Bera / Alexis Courbet / Alex Kang / T J Brunette / Una Nattermann / Evelyn Tsai / Ayesha Saleem / Cameron M Chow / Damian Ekiert / Gira Bhabha / David Veesler / David Baker /  Abstract: A systematic and robust approach to generating complex protein nanomaterials would have broad utility. We develop a hierarchical approach to designing multi-component protein assemblies from two ...A systematic and robust approach to generating complex protein nanomaterials would have broad utility. We develop a hierarchical approach to designing multi-component protein assemblies from two classes of modular building blocks: designed helical repeat proteins (DHRs) and helical bundle oligomers (HBs). We first rigidly fuse DHRs to HBs to generate a large library of oligomeric building blocks. We then generate assemblies with cyclic, dihedral, and point group symmetries from these building blocks using architecture guided rigid helical fusion with new software named WORMS. X-ray crystallography and cryo-electron microscopy characterization show that the hierarchical design approach can accurately generate a wide range of assemblies, including a 43 nm diameter icosahedral nanocage. The computational methods and building block sets described here provide a very general route to de novo designed protein nanomaterials. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_22305.map.gz emd_22305.map.gz | 49.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-22305-v30.xml emd-22305-v30.xml emd-22305.xml emd-22305.xml | 11 KB 11 KB | Display Display |  EMDB header EMDB header |
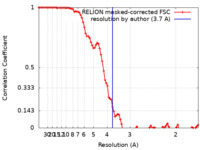
| FSC (resolution estimation) |  emd_22305_fsc.xml emd_22305_fsc.xml | 9 KB | Display |  FSC data file FSC data file |
| Images |  emd_22305.png emd_22305.png | 11 KB | ||
| Filedesc metadata |  emd-22305.cif.gz emd-22305.cif.gz | 5.2 KB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-22305 http://ftp.pdbj.org/pub/emdb/structures/EMD-22305 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22305 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-22305 | HTTPS FTP |
-Validation report
| Summary document |  emd_22305_validation.pdf.gz emd_22305_validation.pdf.gz | 595.2 KB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_22305_full_validation.pdf.gz emd_22305_full_validation.pdf.gz | 594.8 KB | Display | |
| Data in XML |  emd_22305_validation.xml.gz emd_22305_validation.xml.gz | 10.5 KB | Display | |
| Data in CIF |  emd_22305_validation.cif.gz emd_22305_validation.cif.gz | 13.7 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22305 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22305 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22305 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-22305 | HTTPS FTP |
-Related structure data
| Related structure data |  6xssMC  6xh5C  6xi6C  6xnsC  6xt4C C: citing same article ( M: atomic model generated by this map |
|---|---|
| Similar structure data | |
| EM raw data |  EMPIAR-10599 (Title: CryoEM map of designed helical fusion protein C4_nat_HF-7900 EMPIAR-10599 (Title: CryoEM map of designed helical fusion protein C4_nat_HF-7900Data size: 2.0 TB Data #1: Unaligned multiframe movies of C4_nat_HF-7900 [micrographs - multiframe]) |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_22305.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_22305.map.gz / Format: CCP4 / Size: 59.6 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Voxel size | X=Y=Z: 0.859 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
- Sample components
Sample components
-Entire : Designed fusion of helical bundle and helical repeat proteins
| Entire | Name: Designed fusion of helical bundle and helical repeat proteins |
|---|---|
| Components |
|
-Supramolecule #1: Designed fusion of helical bundle and helical repeat proteins
| Supramolecule | Name: Designed fusion of helical bundle and helical repeat proteins type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
-Macromolecule #1: C4_nat_HFuse-7900
| Macromolecule | Name: C4_nat_HFuse-7900 / type: protein_or_peptide / ID: 1 / Number of copies: 4 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Molecular weight | Theoretical: 28.706484 KDa |
| Recombinant expression | Organism:  |
| Sequence | String: ASSWVMLGLL LSLLNRLSLA AEAYKKAIEL DPNDALAWLL LGSVLLLLGR EEEAEEAARK AIELKPEMDS ARRLEGIIEL IRRAREAAE RAQEAAERTG DPRVRELARE LKRLAQEAAE EVRRDPDSKD VNEALKLIVE AIEAAVRALE AAERTGDPEV R ELARELVR ...String: ASSWVMLGLL LSLLNRLSLA AEAYKKAIEL DPNDALAWLL LGSVLLLLGR EEEAEEAARK AIELKPEMDS ARRLEGIIEL IRRAREAAE RAQEAAERTG DPRVRELARE LKRLAQEAAE EVRRDPDSKD VNEALKLIVE AIEAAVRALE AAERTGDPEV R ELARELVR LAVEAAEEVQ RNPSSSDVNE ALKLIVEAID AAVRALEAAE KTGDPEVREL ARELVRLAVE AAEEVQRNPS SE EVNEALK DIVKAIQEAV ESL |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 1 mg/mL |
|---|---|
| Buffer | pH: 8 |
| Grid | Model: Quantifoil R1.2/1.3 / Material: COPPER / Mesh: 400 / Support film - Material: GRAPHENE OXIDE / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 295 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TALOS ARCTICA |
|---|---|
| Image recording | Film or detector model: GATAN K3 (6k x 4k) / Average electron dose: 57.05 e/Å2 |
| Electron beam | Acceleration voltage: 200 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
| Experimental equipment |  Model: Talos Arctica / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller