+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-23170 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Dihedral ring D2_Wm-01_trunc | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species | synthetic construct (others) | |||||||||
| Method | single particle reconstruction / negative staining / Resolution: 19.0 Å | |||||||||
 Authors Authors | Park YJ / Veesler D | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Nat Commun / Year: 2021 Journal: Nat Commun / Year: 2021Title: Design of multi-scale protein complexes by hierarchical building block fusion. Authors: Yang Hsia / Rubul Mout / William Sheffler / Natasha I Edman / Ivan Vulovic / Young-Jun Park / Rachel L Redler / Matthew J Bick / Asim K Bera / Alexis Courbet / Alex Kang / T J Brunette / Una ...Authors: Yang Hsia / Rubul Mout / William Sheffler / Natasha I Edman / Ivan Vulovic / Young-Jun Park / Rachel L Redler / Matthew J Bick / Asim K Bera / Alexis Courbet / Alex Kang / T J Brunette / Una Nattermann / Evelyn Tsai / Ayesha Saleem / Cameron M Chow / Damian Ekiert / Gira Bhabha / David Veesler / David Baker /  Abstract: A systematic and robust approach to generating complex protein nanomaterials would have broad utility. We develop a hierarchical approach to designing multi-component protein assemblies from two ...A systematic and robust approach to generating complex protein nanomaterials would have broad utility. We develop a hierarchical approach to designing multi-component protein assemblies from two classes of modular building blocks: designed helical repeat proteins (DHRs) and helical bundle oligomers (HBs). We first rigidly fuse DHRs to HBs to generate a large library of oligomeric building blocks. We then generate assemblies with cyclic, dihedral, and point group symmetries from these building blocks using architecture guided rigid helical fusion with new software named WORMS. X-ray crystallography and cryo-electron microscopy characterization show that the hierarchical design approach can accurately generate a wide range of assemblies, including a 43 nm diameter icosahedral nanocage. The computational methods and building block sets described here provide a very general route to de novo designed protein nanomaterials. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_23170.map.gz emd_23170.map.gz | 1.9 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-23170-v30.xml emd-23170-v30.xml emd-23170.xml emd-23170.xml | 17.5 KB 17.5 KB | Display Display |  EMDB header EMDB header |
| Images |  emd_23170.png emd_23170.png | 69.5 KB | ||
| Others |  emd_23170_additional_1.map.gz emd_23170_additional_1.map.gz emd_23170_half_map_1.map.gz emd_23170_half_map_1.map.gz emd_23170_half_map_2.map.gz emd_23170_half_map_2.map.gz | 1018.9 KB 1.8 MB 1.8 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-23170 http://ftp.pdbj.org/pub/emdb/structures/EMD-23170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23170 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-23170 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_23170.map.gz / Format: CCP4 / Size: 2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_23170.map.gz / Format: CCP4 / Size: 2 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 6.4 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Additional map: #1
| File | emd_23170_additional_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
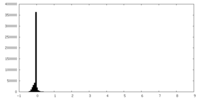
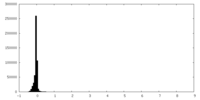
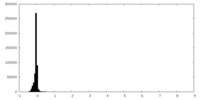
| Density Histograms |
-Half map: #2
| File | emd_23170_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
-Half map: #1
| File | emd_23170_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
| Density Histograms |
- Sample components
Sample components
-Entire : D2_Wm-01_trunc
| Entire | Name: D2_Wm-01_trunc |
|---|---|
| Components |
|
-Supramolecule #1: D2_Wm-01_trunc
| Supramolecule | Name: D2_Wm-01_trunc / type: complex / ID: 1 / Parent: 0 / Macromolecule list: all |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Recombinant expression | Organism:  |
-Macromolecule #1: D2_Wm-01_truncA
| Macromolecule | Name: D2_Wm-01_truncA / type: protein_or_peptide / ID: 1 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Sequence | String: MGTREESLKE QLRSLREQAE LAARLLRLQR EGSSDEDVKE LVAEIIKLIM EQLLLIAEQL LGRS EAAEL ALRAIRLALE LCRQSTDLEE CLRLLKTAIK ALENALRHPD STTAKARLMA ITARLLAQQ LRTQHPDSQA ARDAEKLADQ AERAVRLATR LYEEHPNAEI ...String: MGTREESLKE QLRSLREQAE LAARLLRLQR EGSSDEDVKE LVAEIIKLIM EQLLLIAEQL LGRS EAAEL ALRAIRLALE LCRQSTDLEE CLRLLKTAIK ALENALRHPD STTAKARLMA ITARLLAQQ LRTQHPDSQA ARDAEKLADQ AERAVRLATR LYEEHPNAEI SEMCSQAAYA AALMASIAAI LAQR HPDSQ IARDLIRLAS ELAEMVKRMC ERGGSWGLEH HHHHH |
-Macromolecule #2: D2_Wm-01_truncB
| Macromolecule | Name: D2_Wm-01_truncB / type: protein_or_peptide / ID: 2 / Enantiomer: LEVO |
|---|---|
| Source (natural) | Organism: synthetic construct (others) |
| Sequence | String: MGTREELAKE LLRSLREQAE SLARQLRLQR EGSSDEDVKE LAAEQIKLIM EQLLLIAELT LGRS EAAEL ALDAIRQALE ACRTMDNQEA CTRLLKLAIQ MLELATRAPD AEAAKLALEA AKKAIELAN RHPGSQAAED ATKLAQQAME AVRLALKLYE EHPNADIADL ...String: MGTREELAKE LLRSLREQAE SLARQLRLQR EGSSDEDVKE LAAEQIKLIM EQLLLIAELT LGRS EAAEL ALDAIRQALE ACRTMDNQEA CTRLLKLAIQ MLELATRAPD AEAAKLALEA AKKAIELAN RHPGSQAAED ATKLAQQAME AVRLALKLYE EHPNADIADL CRRAAAEAAE AASKAAELAQ RHPD SQAAR DAIKLASQAA EAVKLACELA QEHPNADKAK LCILLASAAA LLASIAAMLA QRHPDSQEA RDMIRIASEL AELVKEICER |
-Experimental details
-Structure determination
| Method | negative staining |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 8 |
|---|---|
| Staining | Type: NEGATIVE / Material: UF |
- Electron microscopy
Electron microscopy
| Microscope | FEI TECNAI 12 |
|---|---|
| Image recording | Film or detector model: GATAN ULTRASCAN 4000 (4k x 4k) / Average electron dose: 50.0 e/Å2 |
| Electron beam | Acceleration voltage: 120 kV / Electron source: LAB6 |
| Electron optics | Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD |
- Image processing
Image processing
| Startup model | Type of model: OTHER / Details: cryoSPARC ab initio |
|---|---|
| Final reconstruction | Resolution.type: BY AUTHOR / Resolution: 19.0 Å / Resolution method: FSC 0.143 CUT-OFF / Software - Name: cryoSPARC / Number images used: 65970 |
| Initial angle assignment | Type: PROJECTION MATCHING |
| Final angle assignment | Type: PROJECTION MATCHING |
 Movie
Movie Controller
Controller



 UCSF Chimera
UCSF Chimera




















 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)