+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-21145 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Structure of the bovine BBSome:ARL6:GTP complex | |||||||||
 Map data Map data | BBSome:ARL6:GTP complex. Post-processed map. | |||||||||
 Sample Sample |
| |||||||||
 Keywords Keywords | Cilia / ciliopathy / complex / membrane-protein transport / PROTEIN TRANSPORT | |||||||||
| Function / homology |  Function and homology information Function and homology informationBBSome binding / establishment of anatomical structure orientation / BBSome-mediated cargo-targeting to cilium / multi-ciliated epithelial cell differentiation / receptor localization to non-motile cilium / renal tubule development / BBSome / camera-type eye photoreceptor cell differentiation / smoothened binding / establishment of planar polarity ...BBSome binding / establishment of anatomical structure orientation / BBSome-mediated cargo-targeting to cilium / multi-ciliated epithelial cell differentiation / receptor localization to non-motile cilium / renal tubule development / BBSome / camera-type eye photoreceptor cell differentiation / smoothened binding / establishment of planar polarity / photoreceptor connecting cilium / axonemal microtubule / inner ear receptor cell stereocilium organization / patched binding / olfactory bulb development / membrane coat / non-motile cilium assembly / phosphatidylinositol-3-phosphate binding / establishment of epithelial cell apical/basal polarity / protein localization to cilium / regulation of stress fiber assembly / non-motile cilium / motile cilium / protein targeting to membrane / centrosome cycle / ciliary membrane / pericentriolar material / fat cell differentiation / axoneme / centriolar satellite / protein polymerization / cilium assembly / intracellular transport / vesicle-mediated transport / ciliary basal body / protein localization to plasma membrane / axon guidance / intracellular protein transport / protein localization / multicellular organism growth / phospholipid binding / cilium / regulation of protein localization / sensory perception of smell / protein transport / protein-macromolecule adaptor activity / RNA polymerase II-specific DNA-binding transcription factor binding / neuron projection / GTPase activity / centrosome / GTP binding / membrane / cytoplasm Similarity search - Function | |||||||||
| Biological species |  | |||||||||
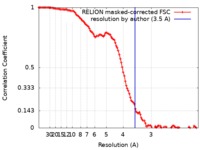
| Method | single particle reconstruction / cryo EM / Resolution: 3.5 Å | |||||||||
 Authors Authors | Singh SK / Gui M | |||||||||
| Funding support |  United States, 1 items United States, 1 items
| |||||||||
 Citation Citation |  Journal: Elife / Year: 2020 Journal: Elife / Year: 2020Title: Structure and activation mechanism of the BBSome membrane protein trafficking complex. Authors: Sandeep K Singh / Miao Gui / Fujiet Koh / Matthew Cj Yip / Alan Brown /  Abstract: Bardet-Biedl syndrome (BBS) is a currently incurable ciliopathy caused by the failure to correctly establish or maintain cilia-dependent signaling pathways. Eight proteins associated with BBS ...Bardet-Biedl syndrome (BBS) is a currently incurable ciliopathy caused by the failure to correctly establish or maintain cilia-dependent signaling pathways. Eight proteins associated with BBS assemble into the BBSome, a key regulator of the ciliary membrane proteome. We report the electron cryomicroscopy (cryo-EM) structures of the native bovine BBSome in inactive and active states at 3.1 and 3.5 Å resolution, respectively. In the active state, the BBSome is bound to an Arf-family GTPase (ARL6/BBS3) that recruits the BBSome to ciliary membranes. ARL6 recognizes a composite binding site formed by BBS1 and BBS7 that is occluded in the inactive state. Activation requires an unexpected swiveling of the β-propeller domain of BBS1, the subunit most frequently implicated in substrate recognition, which widens a central cavity of the BBSome. Structural mapping of disease-causing mutations suggests that pathogenesis results from folding defects and the disruption of autoinhibition and activation. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_21145.map.gz emd_21145.map.gz | 116.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-21145-v30.xml emd-21145-v30.xml emd-21145.xml emd-21145.xml | 43.2 KB 43.2 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_21145_fsc.xml emd_21145_fsc.xml | 11.4 KB | Display |  FSC data file FSC data file |
| Images |  emd_21145.png emd_21145.png | 199 KB | ||
| Masks |  emd_21145_msk_1.map emd_21145_msk_1.map emd_21145_msk_2.map emd_21145_msk_2.map emd_21145_msk_3.map emd_21145_msk_3.map emd_21145_msk_4.map emd_21145_msk_4.map | 125 MB 125 MB 125 MB 125 MB |  Mask map Mask map | |
| Filedesc metadata |  emd-21145.cif.gz emd-21145.cif.gz | 10.3 KB | ||
| Others |  emd_21145_additional_1.map.gz emd_21145_additional_1.map.gz emd_21145_additional_2.map.gz emd_21145_additional_2.map.gz emd_21145_additional_3.map.gz emd_21145_additional_3.map.gz emd_21145_additional_4.map.gz emd_21145_additional_4.map.gz emd_21145_additional_5.map.gz emd_21145_additional_5.map.gz emd_21145_half_map_1.map.gz emd_21145_half_map_1.map.gz emd_21145_half_map_2.map.gz emd_21145_half_map_2.map.gz | 98.8 MB 115.9 MB 116.4 MB 116.2 MB 116 MB 99.2 MB 99.2 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-21145 http://ftp.pdbj.org/pub/emdb/structures/EMD-21145 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21145 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-21145 | HTTPS FTP |
-Validation report
| Summary document |  emd_21145_validation.pdf.gz emd_21145_validation.pdf.gz | 1.1 MB | Display |  EMDB validaton report EMDB validaton report |
|---|---|---|---|---|
| Full document |  emd_21145_full_validation.pdf.gz emd_21145_full_validation.pdf.gz | 1.1 MB | Display | |
| Data in XML |  emd_21145_validation.xml.gz emd_21145_validation.xml.gz | 18.4 KB | Display | |
| Data in CIF |  emd_21145_validation.cif.gz emd_21145_validation.cif.gz | 24.1 KB | Display | |
| Arichive directory |  https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21145 https://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21145 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21145 ftp://ftp.pdbj.org/pub/emdb/validation_reports/EMD-21145 | HTTPS FTP |
-Related structure data
| Related structure data |  6vbvMC  6vbuC M: atomic model generated by this map C: citing same article ( |
|---|---|
| Similar structure data |
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|---|
| Related items in Molecule of the Month |
- Map
Map
| File |  Download / File: emd_21145.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_21145.map.gz / Format: CCP4 / Size: 125 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Annotation | BBSome:ARL6:GTP complex. Post-processed map. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.06 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
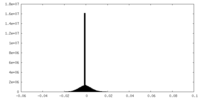
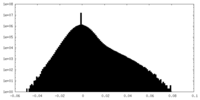
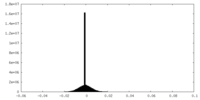
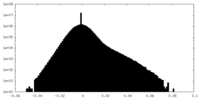
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
+Mask #1
+Mask #2
+Mask #3
+Mask #4
+Additional map: BBSome:ARL6:GTP complex. Unfiltered map
+Additional map: BBSome:ARL6:GTP complex. Chimeric map following multibody refinement.
+Additional map: Post-processed multibody map for the BBSome body
+Additional map: Post-processed multibody map for the BBSome head
+Additional map: Post-processed multibody map for the BBS1-ARL6 interaction.
+Half map: BBSome:ARL6:GTP complex. Half map 1
+Half map: BBSome:ARL6:GTP complex. Half map 2
- Sample components
Sample components
+Entire : Bovine BBSome:ARL6:GTP complex
+Supramolecule #1: Bovine BBSome:ARL6:GTP complex
+Supramolecule #2: BBSome complex
+Supramolecule #3: ADP-ribosylation factor-like protein 6
+Macromolecule #1: Bardet-Biedl syndrome 18 protein
+Macromolecule #2: BBS1 domain-containing protein
+Macromolecule #3: Bardet-Biedl syndrome 2 protein homolog
+Macromolecule #4: Bardet-Biedl syndrome 4 protein homolog
+Macromolecule #5: Bardet-Biedl syndrome 5 protein homolog
+Macromolecule #6: Bardet-Biedl syndrome 7 protein homolog
+Macromolecule #7: Tetratricopeptide repeat domain 8
+Macromolecule #8: Bardet-Biedl syndrome 9
+Macromolecule #9: ADP-ribosylation factor-like protein 6
+Macromolecule #10: CALCIUM ION
+Macromolecule #11: GUANOSINE-5'-TRIPHOSPHATE
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Concentration | 0.7 mg/mL | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Buffer | pH: 7.5 Component:
| ||||||||||||||||||
| Grid | Support film - Material: CARBON / Support film - topology: HOLEY / Details: unspecified | ||||||||||||||||||
| Vitrification | Cryogen name: ETHANE / Chamber humidity: 100 % / Chamber temperature: 293 K / Instrument: FEI VITROBOT MARK II / Details: Grids were blotted for 2 s with a -2 offset.. |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Specialist optics | Energy filter - Name: GIF Bioquantum / Energy filter - Slit width: 25 eV |
| Image recording | Film or detector model: GATAN K3 BIOQUANTUM (6k x 4k) / Number grids imaged: 2 / Number real images: 9408 / Average exposure time: 4.0 sec. / Average electron dose: 56.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm / Nominal defocus max: 2.4 µm / Nominal defocus min: 1.1 µm / Nominal magnification: 81000 |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
+ Image processing
Image processing
-Atomic model buiding 1
| Details | During refinement, the resolution limit was set to 3.5 Angstrom. Secondary structure, Ramachandran and rotamer restraints were applied during refinement. |
|---|---|
| Refinement | Space: REAL / Protocol: OTHER / Overall B value: 43.6 / Target criteria: Correlation coefficient |
| Output model |  PDB-6vbv: |
 Movie
Movie Controller
Controller
















 Z
Z Y
Y X
X