[English] 日本語
 Yorodumi
Yorodumi- EMDB-12032: Cryo-EM structure of the contractile injection system crown trime... -
+ Open data
Open data
- Basic information
Basic information
| Entry | Database: EMDB / ID: EMD-12032 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Title | Cryo-EM structure of the contractile injection system crown trimer from Anabaena PCC7120 | |||||||||
 Map data Map data | ||||||||||
 Sample Sample |
| |||||||||
| Biological species |  Nostoc sp. PCC 7120 = FACHB-418 (bacteria) Nostoc sp. PCC 7120 = FACHB-418 (bacteria) | |||||||||
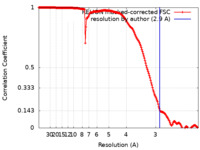
| Method | single particle reconstruction / cryo EM / Resolution: 2.9 Å | |||||||||
 Authors Authors | Eisenstein F / Weiss GL / Pilhofer M | |||||||||
 Citation Citation |  Journal: Nat Microbiol / Year: 2022 Journal: Nat Microbiol / Year: 2022Title: Structure of a thylakoid-anchored contractile injection system in multicellular cyanobacteria. Authors: Gregor L Weiss / Fabian Eisenstein / Ann-Katrin Kieninger / Jingwei Xu / Hannah A Minas / Milena Gerber / Miki Feldmüller / Iris Maldener / Karl Forchhammer / Martin Pilhofer /    Abstract: Contractile injection systems (CISs) mediate cell-cell interactions by phage tail-like structures, using two distinct modes of action: extracellular CISs are released into the medium, while type 6 ...Contractile injection systems (CISs) mediate cell-cell interactions by phage tail-like structures, using two distinct modes of action: extracellular CISs are released into the medium, while type 6 secretion systems (T6SSs) are attached to the cytoplasmic membrane and function upon cell-cell contact. Here, we characterized a CIS in the multicellular cyanobacterium Anabaena, with features distinct from extracellular CISs and T6SSs. Cryo-electron tomography of focused ion beam-milled cells revealed that CISs were anchored in thylakoid membrane stacks, facing the cell periphery. Single particle cryo-electron microscopy showed that this unique in situ localization was mediated by extensions of tail fibre and baseplate components. On stress, cyanobacteria induced the formation of ghost cells, presenting thylakoid-anchored CISs to the environment. Functional assays suggest that these CISs may mediate ghost cell formation and/or interactions of ghost cells with other organisms. Collectively, these data provide a framework for understanding the evolutionary re-engineering of CISs and potential roles of these CISs in cyanobacterial programmed cell death. | |||||||||
| History |
|
- Structure visualization
Structure visualization
| Movie |
 Movie viewer Movie viewer |
|---|---|
| Structure viewer | EM map:  SurfView SurfView Molmil Molmil Jmol/JSmol Jmol/JSmol |
| Supplemental images |
- Downloads & links
Downloads & links
-EMDB archive
| Map data |  emd_12032.map.gz emd_12032.map.gz | 393.4 MB |  EMDB map data format EMDB map data format | |
|---|---|---|---|---|
| Header (meta data) |  emd-12032-v30.xml emd-12032-v30.xml emd-12032.xml emd-12032.xml | 17.7 KB 17.7 KB | Display Display |  EMDB header EMDB header |
| FSC (resolution estimation) |  emd_12032_fsc.xml emd_12032_fsc.xml | 18 KB | Display |  FSC data file FSC data file |
| Images |  emd_12032.png emd_12032.png | 53.8 KB | ||
| Masks |  emd_12032_msk_1.map emd_12032_msk_1.map | 512 MB |  Mask map Mask map | |
| Others |  emd_12032_half_map_1.map.gz emd_12032_half_map_1.map.gz emd_12032_half_map_2.map.gz emd_12032_half_map_2.map.gz | 407.3 MB 407.3 MB | ||
| Archive directory |  http://ftp.pdbj.org/pub/emdb/structures/EMD-12032 http://ftp.pdbj.org/pub/emdb/structures/EMD-12032 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12032 ftp://ftp.pdbj.org/pub/emdb/structures/EMD-12032 | HTTPS FTP |
-Related structure data
- Links
Links
| EMDB pages |  EMDB (EBI/PDBe) / EMDB (EBI/PDBe) /  EMDataResource EMDataResource |
|---|
- Map
Map
| File |  Download / File: emd_12032.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) Download / File: emd_12032.map.gz / Format: CCP4 / Size: 512 MB / Type: IMAGE STORED AS FLOATING POINT NUMBER (4 BYTES) | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & slices | Image control
Images are generated by Spider. | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Voxel size | X=Y=Z: 1.1 Å | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
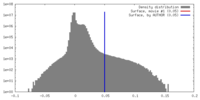
| Density |
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Symmetry | Space group: 1 | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
| Details | EMDB XML:
CCP4 map header:
| ||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
-Supplemental data
-Mask #1
| File |  emd_12032_msk_1.map emd_12032_msk_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
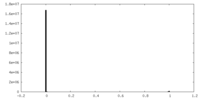
| Density Histograms |
-Half map: #2
| File | emd_12032_half_map_1.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
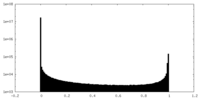
| Density Histograms |
-Half map: #1
| File | emd_12032_half_map_2.map | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Projections & Slices |
| ||||||||||||
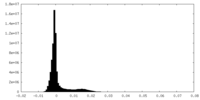
| Density Histograms |
- Sample components
Sample components
-Entire : Base plate complex (C1) of the Anabaena PCC7120 contractile injec...
| Entire | Name: Base plate complex (C1) of the Anabaena PCC7120 contractile injection system |
|---|---|
| Components |
|
-Supramolecule #1: Base plate complex (C1) of the Anabaena PCC7120 contractile injec...
| Supramolecule | Name: Base plate complex (C1) of the Anabaena PCC7120 contractile injection system type: complex / ID: 1 / Parent: 0 / Macromolecule list: #1-#11 / Details: focussed on crown trimer |
|---|---|
| Source (natural) | Organism:  Nostoc sp. PCC 7120 = FACHB-418 (bacteria) Nostoc sp. PCC 7120 = FACHB-418 (bacteria) |
-Experimental details
-Structure determination
| Method | cryo EM |
|---|---|
 Processing Processing | single particle reconstruction |
| Aggregation state | particle |
- Sample preparation
Sample preparation
| Buffer | pH: 7.5 |
|---|---|
| Grid | Model: Quantifoil / Support film - Material: CARBON / Support film - topology: CONTINUOUS |
| Vitrification | Cryogen name: ETHANE-PROPANE / Chamber humidity: 100 % / Chamber temperature: 281 K / Instrument: FEI VITROBOT MARK IV |
- Electron microscopy
Electron microscopy
| Microscope | FEI TITAN KRIOS |
|---|---|
| Image recording | Film or detector model: GATAN K2 QUANTUM (4k x 4k) / Detector mode: COUNTING / Number grids imaged: 4 / Number real images: 19000 / Average electron dose: 52.0 e/Å2 |
| Electron beam | Acceleration voltage: 300 kV / Electron source:  FIELD EMISSION GUN FIELD EMISSION GUN |
| Electron optics | C2 aperture diameter: 50.0 µm / Calibrated defocus max: 1.5 µm / Calibrated defocus min: 0.9 µm / Illumination mode: FLOOD BEAM / Imaging mode: BRIGHT FIELD / Cs: 2.7 mm |
| Sample stage | Specimen holder model: FEI TITAN KRIOS AUTOGRID HOLDER / Cooling holder cryogen: NITROGEN |
| Experimental equipment |  Model: Titan Krios / Image courtesy: FEI Company |
 Movie
Movie Controller
Controller
























 Z (Sec.)
Z (Sec.) Y (Row.)
Y (Row.) X (Col.)
X (Col.)