1QSP
 
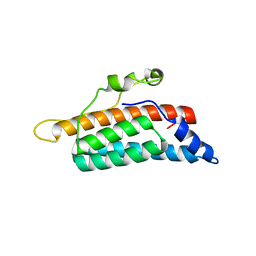 | | CRYSTAL STRUCTURE OF THE YEAST PHOSPHORELAY PROTEIN YPD1 | | Descriptor: | YPD1 | | Authors: | Xu, Q, West, A.H. | | Deposit date: | 1999-06-22 | | Release date: | 1999-10-13 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Conservation of structure and function among histidine-containing phosphotransfer (HPt) domains as revealed by the crystal structure of YPD1.
J.Mol.Biol., 292, 1999
|
|
1OXK
 
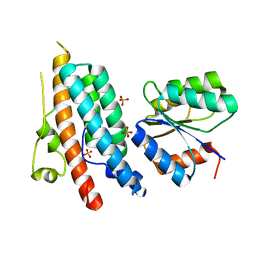 | |
1OXB
 
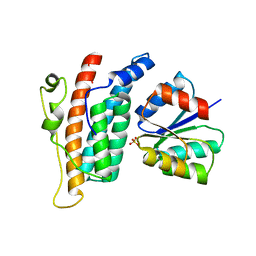 | |
1RY5
 
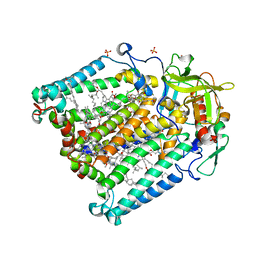 | | PHOTOSYNTHETIC REACTION CENTER MUTANT FROM RHODOBACTER SPHAEROIDES WITH ASP L213 REPLACED WITH ASN | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, CARDIOLIPIN, ... | | Authors: | Xu, Q, Axelrod, H.L, Abresch, E.C, Paddock, M.L, Okamura, M.Y, Feher, G. | | Deposit date: | 2003-12-19 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | X-Ray Structure Determination of Three Mutants of the Bacterial Photosynthetic Reaction Centers from Rb. sphaeroides; Altered Proton Transfer Pathways.
STRUCTURE, 12, 2004
|
|
1S00
 
 | | PHOTOSYNTHETIC REACTION CENTER DOUBLE MUTANT FROM RHODOBACTER SPHAEROIDES WITH ASP L213 REPLACED WITH ASN AND ARG M233 REPLACED WITH CYS IN THE CHARGE-SEPARATED D+QAQB- STATE | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, FE (II) ION, ... | | Authors: | Xu, Q, Axelrod, H.L, Abresch, E.C, Paddock, M.L, Okamura, M.Y, Feher, G. | | Deposit date: | 2003-12-29 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | X-Ray Structure Determination of Three Mutants of the Bacterial Photosynthetic Reaction Centers from Rb. sphaeroides; Altered Proton Transfer Pathways.
STRUCTURE, 12, 2004
|
|
1RVJ
 
 | | PHOTOSYNTHETIC REACTION CENTER DOUBLE MUTANT FROM RHODOBACTER SPHAEROIDES WITH ASP L213 REPLACED WITH ASN AND ARG H177 REPLACED WITH HIS | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, CARDIOLIPIN, ... | | Authors: | Xu, Q, Axelrod, H.L, Abresch, E.C, Paddock, M.L, Okamura, M.Y, Feher, G. | | Deposit date: | 2003-12-14 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.75 Å) | | Cite: | X-Ray Structure Determination of Three Mutants of the Bacterial Photosynthetic Reaction
Centers from Rb. sphaeroides; Altered Proton Transfer Pathways.
Structure, 12, 2004
|
|
1RZZ
 
 | | PHOTOSYNTHETIC REACTION CENTER DOUBLE MUTANT FROM RHODOBACTER SPHAEROIDES WITH ASP L213 REPLACED WITH ASN AND ARG M233 REPLACED WITH CYS IN THE CHARGE-NEUTRAL DQAQB STATE (TETRAGONAL FORM) | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, FE (II) ION, ... | | Authors: | Xu, Q, Axelrod, H.L, Abresch, E.C, Paddock, M.L, Okamura, M.Y, Feher, G. | | Deposit date: | 2003-12-29 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.4 Å) | | Cite: | X-Ray Structure Determination of Three Mutants of the Bacterial Photosynthetic Reaction Centers from Rb. sphaeroides; Altered Proton Transfer Pathways.
STRUCTURE, 12, 2004
|
|
1RZH
 
 | | PHOTOSYNTHETIC REACTION CENTER DOUBLE MUTANT FROM RHODOBACTER SPHAEROIDES WITH ASP L213 REPLACED WITH ASN AND ARG M233 REPLACED WITH CYS IN THE CHARGE-NEUTRAL DQAQB STATE (TRIGONAL FORM) | | Descriptor: | BACTERIOCHLOROPHYLL A, BACTERIOPHEOPHYTIN A, CARDIOLIPIN, ... | | Authors: | Xu, Q, Axelrod, H.L, Abresch, E.C, Paddock, M.L, Okamura, M.Y, Feher, G. | | Deposit date: | 2003-12-24 | | Release date: | 2004-04-13 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | X-Ray Structure Determination of Three Mutants of the Bacterial Photosynthetic Reaction Centers from Rb. sphaeroides; Altered Proton Transfer Pathways.
STRUCTURE, 12, 2004
|
|
3HFE
 
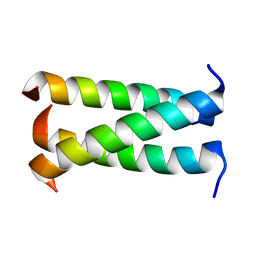 | | A trimeric form of the Kv7.1 A domain Tail | | Descriptor: | Potassium voltage-gated channel subfamily KQT member 1 | | Authors: | Xu, Q, Minor, D.L. | | Deposit date: | 2009-05-11 | | Release date: | 2009-09-01 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of a trimeric form of the K(V)7.1 (KCNQ1) A-domain tail coiled-coil reveals structural plasticity and context dependent changes in a putative coiled-coil trimerization motif.
Protein Sci., 18, 2009
|
|
3HFC
 | | A trimeric form of the Kv7.1 A domain Tail, L602M/L606M mutant Semet | | Descriptor: | Potassium voltage-gated channel subfamily KQT member 1 | | Authors: | Xu, Q, Minor, D.L. | | Deposit date: | 2009-05-11 | | Release date: | 2009-09-01 | | Last modified: | 2024-11-27 | | Method: | X-RAY DIFFRACTION (2.45 Å) | | Cite: | Crystal structure of a trimeric form of the K(V)7.1 (KCNQ1) A-domain tail coiled-coil reveals structural plasticity and context dependent changes in a putative coiled-coil trimerization motif.
Protein Sci., 18, 2009
|
|
4GOW
 
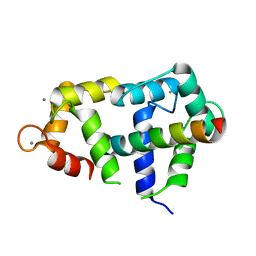 | | Crystal Structure of Ca2+/CaM:Kv7.4 (KCNQ4) B helix complex | | Descriptor: | CALCIUM ION, Calmodulin, Potassium voltage-gated channel subfamily KQT member 4 | | Authors: | Xu, Q, Chang, A, Tolia, A, Minor Jr, D.L. | | Deposit date: | 2012-08-20 | | Release date: | 2012-12-12 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structure of a Ca(2+)/CaM:Kv7.4 (KCNQ4) B-helix complex provides insight into M current modulation.
J.Mol.Biol., 425, 2013
|
|
6JYX
 
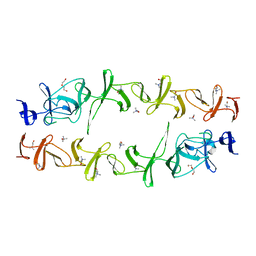 | | Structure of CbpJ from Streptococcus Pneumoniae TIGR4 | | Descriptor: | CHOLINE ION, Choline binding protein J, DI(HYDROXYETHYL)ETHER | | Authors: | Xu, Q, Zhang, J.W, Li, Q, Jiang, Y.L. | | Deposit date: | 2019-04-29 | | Release date: | 2019-06-05 | | Last modified: | 2023-11-22 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structure of the choline-binding protein CbpJ from Streptococcus pneumoniae.
Biochem.Biophys.Res.Commun., 514, 2019
|
|
7EFT
 | |
9GAF
 
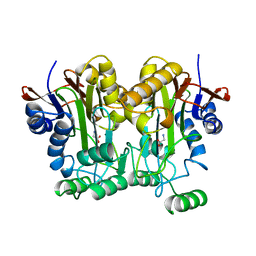 | |
9GAA
 
 | |
9GAC
 
 | |
4L6R
 
 | | Structure of the class B human glucagon G protein coupled receptor | | Descriptor: | DI(HYDROXYETHYL)ETHER, Soluble cytochrome b562 and Glucagon receptor chimera | | Authors: | Siu, F.Y, He, M, de Graaf, C, Han, G.W, Yang, D, Zhang, Z, Zhou, C, Xu, Q, Wacker, D, Joseph, J.S, Liu, W, Lau, J, Cherezov, V, Katritch, V, Wang, M.W, Stevens, R.C, GPCR Network (GPCR) | | Deposit date: | 2013-06-12 | | Release date: | 2013-07-24 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Structure of the human glucagon class B G-protein-coupled receptor.
Nature, 499, 2013
|
|
8ZVQ
 
 | |
9K7D
 
 | | Crystal structure of GH57 family amylopullulanase from Aquifex aeolicus mutant E256Q in complex with Maltohexaose | | Descriptor: | Glycoside hydrolase family 57 N-terminal domain-containing protein, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Zhu, Z.M, Wang, W.W, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S, Yu, F. | | Deposit date: | 2024-10-23 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (1.72 Å) | | Cite: | The crystal structure of GH57 family amylopullulanase reveals its dual binding pockets sharing the same catalytic dyad.
Commun Biol, 8, 2025
|
|
9KYQ
 
 | | GH57 family amylopullulanase from Aquifex aeolicus wild type co-crystallize with gama-cyclodextrin | | Descriptor: | 1,2-ETHANEDIOL, DI(HYDROXYETHYL)ETHER, GLYCEROL, ... | | Authors: | Zhu, Z.M, Wang, W.W, Yu, F, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S. | | Deposit date: | 2024-12-09 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | Mechanistic insights into cyclodextrins as substrates and inhibitors of GH57 family amylopullulanase from Aquifex aeolicus.
J.Struct.Biol., 217, 2025
|
|
9JLW
 
 | | Crystal structure of GH57 family amylopullulanase from Aquifex aeolicus mutant D352N in complex with maltoheptaose | | Descriptor: | GLYCEROL, Glycoside hydrolase family 57 N-terminal domain-containing protein, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, ... | | Authors: | Zhu, Z.M, Wang, W.W, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S, Yu, F. | | Deposit date: | 2024-09-19 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | High-Temperature Crystallization Method for the GH57 Family Hyperthermophilic Amylopullulanase from Aquifex aeolicus
Cryst.Growth Des., 24, 2024
|
|
9JLU
 
 | | GH57 family amylopullulanase from Aquifex aeolicus D352N mutant complex with beta-cyclodextrin | | Descriptor: | Cycloheptakis-(1-4)-(alpha-D-glucopyranose), GLYCEROL, Glycoside hydrolase family 57 N-terminal domain-containing protein | | Authors: | Zhu, Z.M, Wang, W.W, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S, Yu, F. | | Deposit date: | 2024-09-19 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Mechanistic insights into cyclodextrins as substrates and inhibitors of GH57 family amylopullulanase from Aquifex aeolicus.
J.Struct.Biol., 217, 2025
|
|
9JLR
 
 | | Crystal structure of GH57 family amylopullulanase from Aquifex aeolicus mutant E256Q in complex with maltopentaose | | Descriptor: | Glycoside hydrolase family 57 N-terminal domain-containing protein, alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose | | Authors: | Zhu, Z.M, Wang, W.W, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S, Yu, F. | | Deposit date: | 2024-09-19 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (2.79 Å) | | Cite: | High-Temperature Crystallization Method for the GH57 Family Hyperthermophilic Amylopullulanase from Aquifex aeolicus
Cryst.Growth Des., 24, 2024
|
|
9JLS
 
 | | Crystal structure of GH57 family amylopullulanase from Aquifex aeolicus wild type in complex with acarbose | | Descriptor: | 4,6-dideoxy-4-{[(1S,4R,5S,6S)-4,5,6-trihydroxy-3-(hydroxymethyl)cyclohex-2-en-1-yl]amino}-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose-(1-4)-alpha-D-glucopyranose, Glycoside hydrolase family 57 N-terminal domain-containing protein | | Authors: | Zhu, Z.M, Wang, W.W, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S, Yu, F. | | Deposit date: | 2024-09-19 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (3.06 Å) | | Cite: | High-Temperature Crystallization Method for the GH57 Family Hyperthermophilic Amylopullulanase from Aquifex aeolicus
Cryst.Growth Des., 24, 2024
|
|
9KYP
 
 | | GH57 family amylopullulanase from Aquifex aeolicus wild type complex with beta-cyclodextrin | | Descriptor: | 1,2-ETHANEDIOL, Cycloheptakis-(1-4)-(alpha-D-glucopyranose), DI(HYDROXYETHYL)ETHER, ... | | Authors: | Zhu, Z.M, Wang, W.W, Yu, F, Li, M.J, Xu, Q, Zhou, H, Huang, L.Q, Wang, Q.S. | | Deposit date: | 2024-12-09 | | Release date: | 2025-06-04 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | Mechanistic insights into cyclodextrins as substrates and inhibitors of GH57 family amylopullulanase from Aquifex aeolicus.
J.Struct.Biol., 217, 2025
|
|
