1H15
 
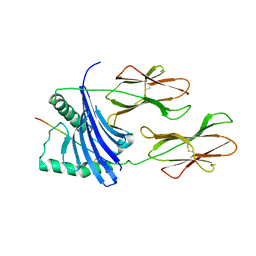 | | X-ray crystal structure of HLA-DRA1*0101/DRB5*0101 complexed with a peptide from Epstein Barr Virus DNA polymerase | | Descriptor: | 2-acetamido-2-deoxy-beta-D-glucopyranose, 2-acetamido-2-deoxy-beta-D-glucopyranose-(1-4)-2-acetamido-2-deoxy-beta-D-glucopyranose, DNA POLYMERASE, ... | | Authors: | Lang, H, Jacobsen, H, Ikemizu, S, Andersson, C, Harlos, K, Madsen, L, Hjorth, P, Sondergaard, L, Svejgaard, A, Wucherpfennig, K, Stuart, D.I, Bell, J.I, Jones, E.Y, Fugger, L. | | Deposit date: | 2002-07-02 | | Release date: | 2002-10-03 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | A Functional and Structural Basis for Tcr Cross-Reactivity in Multiple Sclerosis
Nat.Immunol., 3, 2002
|
|
4KAP
 
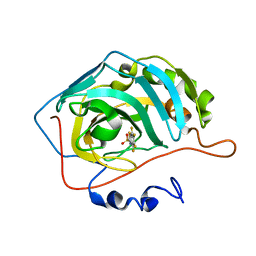 | | The Binding of Benzoarylsulfonamide Ligands to Human Carbonic Anhydrase is Insensitive to Formal Fluorination of the Ligand | | Descriptor: | 4,5,6,7-tetrafluoro-1,3-benzothiazole-2-sulfonamide, Carbonic anhydrase 2, ZINC ION | | Authors: | Lange, H, Lockett, M.R, Breiten, B, Heroux, A, Sherman, W, Rappoport, D, Yau, P.O, Snyder, P.W, Whitesides, G.M. | | Deposit date: | 2013-04-22 | | Release date: | 2013-07-10 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (1.45 Å) | | Cite: | The Binding of Benzoarylsulfonamide Ligands to Human Carbonic Anhydrase is Insensitive to Formal Fluorination of the Ligand.
Angew.Chem.Int.Ed.Engl., 52, 2013
|
|
9IOP
 
 | | Cryo-EM structure of cUA and MAFP bound CapE filament | | Descriptor: | 3'3'-cUMP-AMP, METHYL ARACHIDONYL FLUOROPHOSPHONATE, cUMP-AMP-activated phospholipase | | Authors: | Gao, A, Wang, J.G. | | Deposit date: | 2024-07-09 | | Release date: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.33 Å) | | Cite: | Cyclic-dinucleotide-induced filamentous assembly of phospholipases governs broad CBASS immunity.
Cell, 2025
|
|
9IOQ
 
 | | Crystal structure of CapE bound cUA | | Descriptor: | 3'3'-cUMP-AMP, cUMP-AMP-activated phospholipase | | Authors: | Gao, A, Wang, J.G. | | Deposit date: | 2024-07-09 | | Release date: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (1.999 Å) | | Cite: | Cyclic-dinucleotide-induced filamentous assembly of phospholipases governs broad CBASS immunity.
Cell, 2025
|
|
9IOM
 
 | | CapE apo form | | Descriptor: | cUMP-AMP-activated phospholipase | | Authors: | Gao, A, Wang, J.G. | | Deposit date: | 2024-07-09 | | Release date: | 2025-05-21 | | Method: | X-RAY DIFFRACTION (1.648 Å) | | Cite: | Cyclic-dinucleotide-induced filamentous assembly of phospholipases governs broad CBASS immunity.
Cell, 2025
|
|
9ION
 
 | | Cryo-EM structure of cUA bound CapE filament | | Descriptor: | 3'3'-cUMP-AMP, cUMP-AMP-activated phospholipase | | Authors: | Gao, A, Wang, J.G. | | Deposit date: | 2024-07-09 | | Release date: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.27 Å) | | Cite: | Cyclic-dinucleotide-induced filamentous assembly of phospholipases governs broad CBASS immunity.
Cell, 2025
|
|
7UVR
 
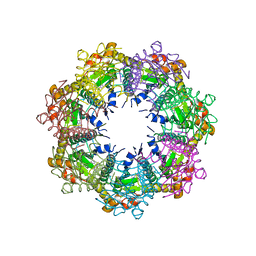 | | Crystal structure of human ClpP protease in complex with TR-65 | | Descriptor: | 3-{[(10R)-4-[(4-chlorophenyl)methyl]-5-oxo-1,2,4,5,8,9-hexahydroimidazo[1,2-a]pyrido[3,4-e]pyrimidin-7(6H)-yl]methyl}benzonitrile, ATP-dependent Clp protease proteolytic subunit, mitochondrial | | Authors: | Mabanglo, M.F, Houry, W.A. | | Deposit date: | 2022-05-02 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.86 Å) | | Cite: | Potent ClpP agonists with anticancer properties bind with improved structural complementarity and alter the mitochondrial N-terminome.
Structure, 31, 2023
|
|
7UW0
 
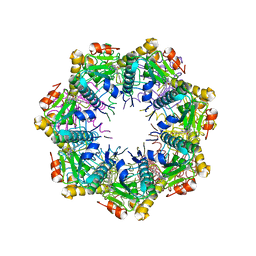 | | Crystal structure of human ClpP protease in complex with TR-133 | | Descriptor: | 3-({3-[(4-bromophenyl)methyl]-4-oxo-3,5,7,8-tetrahydropyrido[4,3-d]pyrimidin-6(4H)-yl}methyl)benzonitrile, ATP-dependent Clp protease proteolytic subunit, mitochondrial | | Authors: | Mabanglo, M.F, Houry, W.A. | | Deposit date: | 2022-05-02 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.8 Å) | | Cite: | Potent ClpP agonists with anticancer properties bind with improved structural complementarity and alter the mitochondrial N-terminome.
Structure, 31, 2023
|
|
7UVM
 | | Crystal structure of human ClpP protease in complex with TR-27 | | Descriptor: | (10R)-4-[(4-chlorophenyl)methyl]-7-[(3-ethynylphenyl)methyl]-2,4,6,7,8,9-hexahydroimidazo[1,2-a]pyrido[3,4-e]pyrimidin-5(1H)-one, ATP-dependent Clp protease proteolytic subunit, mitochondrial | | Authors: | Mabanglo, M.F, Houry, W.A. | | Deposit date: | 2022-05-02 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (2.19 Å) | | Cite: | Potent ClpP agonists with anticancer properties bind with improved structural complementarity and alter the mitochondrial N-terminome.
Structure, 31, 2023
|
|
7UVN
 
 | | Crystal structure of human ClpP protease in complex with TR-57 | | Descriptor: | 3-({3-[(4-chlorophenyl)methyl]-1-methyl-2,4-dioxo-1,3,4,5,7,8-hexahydropyrido[4,3-d]pyrimidin-6(2H)-yl}methyl)benzonitrile, ATP-dependent Clp protease proteolytic subunit, mitochondrial | | Authors: | Mabanglo, M.F, Houry, W.A. | | Deposit date: | 2022-05-02 | | Release date: | 2023-01-11 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (3.11 Å) | | Cite: | Potent ClpP agonists with anticancer properties bind with improved structural complementarity and alter the mitochondrial N-terminome.
Structure, 31, 2023
|
|
7UVU
 
 | | Crystal structure of human ClpP protease in complex with TR-107 | | Descriptor: | 3-({3-[(4-chlorophenyl)methyl]-4-oxo-3,5,7,8-tetrahydropyrido[4,3-d]pyrimidin-6(4H)-yl}methyl)benzonitrile, ATP-dependent Clp protease proteolytic subunit, mitochondrial | | Authors: | Mabanglo, M.F, Houry, W.A. | | Deposit date: | 2022-05-02 | | Release date: | 2023-01-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (3.24 Å) | | Cite: | Potent ClpP agonists with anticancer properties bind with improved structural complementarity and alter the mitochondrial N-terminome.
Structure, 31, 2023
|
|
6FZY
 
 | | PPAR mutant | | Descriptor: | Peroxisome proliferator-activated receptor gamma | | Authors: | Rochel, N. | | Deposit date: | 2018-03-15 | | Release date: | 2019-02-13 | | Last modified: | 2024-05-08 | | Method: | X-RAY DIFFRACTION (3.1 Å) | | Cite: | Recurrent activating mutations of PPAR gamma associated with luminal bladder tumors.
Nat Commun, 10, 2019
|
|
6FZG
 
 | | PPAR gamma mutant complex | | Descriptor: | (2~{S})-3-[4-[2-[methyl(pyridin-2-yl)amino]ethoxy]phenyl]-2-[[2-(phenylcarbonyl)phenyl]amino]propanoic acid, CHLORIDE ION, Peroxisome proliferator-activated receptor gamma | | Authors: | Rochel, N. | | Deposit date: | 2018-03-14 | | Release date: | 2019-02-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Recurrent activating mutations of PPAR gamma associated with luminal bladder tumors.
Nat Commun, 10, 2019
|
|
6FZP
 
 | | PPAR gamma complex. | | Descriptor: | (2~{S})-3-[4-[2-[methyl(pyridin-2-yl)amino]ethoxy]phenyl]-2-[[2-(phenylcarbonyl)phenyl]amino]propanoic acid, Peroxisome proliferator-activated receptor gamma, Peroxisome proliferator-activated receptor gamma coactivator 1-alpha | | Authors: | Rochel, N, Beji, S. | | Deposit date: | 2018-03-15 | | Release date: | 2019-02-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Recurrent activating mutations of PPAR gamma associated with luminal bladder tumors.
Nat Commun, 10, 2019
|
|
6FZF
 
 | | PPAR mutant complex | | Descriptor: | (2~{S})-3-[4-[2-[methyl(pyridin-2-yl)amino]ethoxy]phenyl]-2-[[2-(phenylcarbonyl)phenyl]amino]propanoic acid, Peroxisome proliferator-activated receptor gamma, Peroxisome proliferator-activated receptor gamma coactivator 1-alpha | | Authors: | Rochel, N. | | Deposit date: | 2018-03-14 | | Release date: | 2019-02-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | Recurrent activating mutations of PPAR gamma associated with luminal bladder tumors.
Nat Commun, 10, 2019
|
|
6FZJ
 
 | | PPAR gamma mutant complex | | Descriptor: | (2~{S})-3-[4-[2-[methyl(pyridin-2-yl)amino]ethoxy]phenyl]-2-[[2-(phenylcarbonyl)phenyl]amino]propanoic acid, CHLORIDE ION, IODIDE ION, ... | | Authors: | Rochel, N. | | Deposit date: | 2018-03-14 | | Release date: | 2019-02-13 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.011 Å) | | Cite: | Recurrent activating mutations of PPAR gamma associated with luminal bladder tumors.
Nat Commun, 10, 2019
|
|
6CYC
 
 | | PDE2 in complex with compound 5 | | Descriptor: | 1,2-ETHANEDIOL, 3-(hydroxymethyl)-1-{(1S)-1-[4-(trifluoromethyl)phenyl]ethyl}-1H-pyrazolo[3,4-d]pyrimidine-4,6(5H,7H)-dione, MAGNESIUM ION, ... | | Authors: | Lu, J. | | Deposit date: | 2018-04-05 | | Release date: | 2018-09-19 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.54 Å) | | Cite: | Structure-Guided Design and Procognitive Assessment of a Potent and Selective Phosphodiesterase 2A Inhibitor.
ACS Med Chem Lett, 9, 2018
|
|
8GK7
 
 | | MsbA bound to cerastecin C | | Descriptor: | 2-[(4-butylbenzene-1-sulfonyl)amino]-5-[(3-{4-[(4-butylbenzene-1-sulfonyl)amino]-3-carboxyanilino}-3-oxopropyl)carbamoyl]benzoic acid, Lipid A export ATP-binding/permease protein MsbA, PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER | | Authors: | Chen, Y, Klein, D. | | Deposit date: | 2023-03-17 | | Release date: | 2024-04-24 | | Last modified: | 2025-05-21 | | Method: | ELECTRON MICROSCOPY (3.32 Å) | | Cite: | Cerastecins inhibit membrane lipooligosaccharide transport in drug-resistant Acinetobacter baumannii.
Nat Microbiol, 9, 2024
|
|
6CYB
 
 | | PDE2 in complex with compound 7 | | Descriptor: | 1,2-ETHANEDIOL, 3-(2,2,2-trifluoroethyl)-1-{(1S)-1-[4-(trifluoromethyl)phenyl]ethyl}-1H-pyrazolo[3,4-d]pyrimidine-4,6(5H,7H)-dione, MAGNESIUM ION, ... | | Authors: | Lu, J. | | Deposit date: | 2018-04-05 | | Release date: | 2018-09-19 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.62 Å) | | Cite: | Structure-Guided Design and Procognitive Assessment of a Potent and Selective Phosphodiesterase 2A Inhibitor.
ACS Med Chem Lett, 9, 2018
|
|
6CYD
 
 | | PDE2 in complex with compound 7 | | Descriptor: | 1,2-ETHANEDIOL, 3-(hydroxymethyl)-6-methyl-1-{(1S)-1-[4-(trifluoromethyl)phenyl]ethyl}-1,5-dihydro-4H-pyrazolo[3,4-d]pyrimidin-4-one, MAGNESIUM ION, ... | | Authors: | Lu, J. | | Deposit date: | 2018-04-05 | | Release date: | 2018-09-19 | | Last modified: | 2024-03-06 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Structure-Guided Design and Procognitive Assessment of a Potent and Selective Phosphodiesterase 2A Inhibitor.
ACS Med Chem Lett, 9, 2018
|
|
1C5B
 
 | | DECARBOXYLASE CATALYTIC ANTIBODY 21D8 UNLIGANDED FORM | | Descriptor: | CHIMERIC DECARBOXYLASE ANTIBODY 21D8 | | Authors: | Hotta, K, Wilson, I.A. | | Deposit date: | 1999-11-08 | | Release date: | 2000-10-11 | | Last modified: | 2024-11-20 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Catalysis of decarboxylation by a preorganized heterogeneous microenvironment: crystal structures of abzyme 21D8.
J.Mol.Biol., 302, 2000
|
|
1C5C
 
 | | DECARBOXYLASE CATALYTIC ANTIBODY 21D8-HAPTEN COMPLEX | | Descriptor: | 2-ACETYLAMINO-NAPTHALENE-1,5-DISULFONIC ACID, CHIMERIC DECARBOXYLASE ANTIBODY 21D8, GLYCEROL | | Authors: | Hotta, K, Wilson, I.A. | | Deposit date: | 1999-11-09 | | Release date: | 2000-10-11 | | Last modified: | 2024-10-30 | | Method: | X-RAY DIFFRACTION (1.61 Å) | | Cite: | Catalysis of decarboxylation by a preorganized heterogeneous microenvironment: crystal structures of abzyme 21D8.
J.Mol.Biol., 302, 2000
|
|
6ZTK
 
 | | Crystal structure of Mialostatin, a gut cystatin from the hard tick Ixodes ricinus | | Descriptor: | FRAGMENT OF TRITON X-100, Mialostatin, SULFATE ION | | Authors: | Busa, M, Rezacova, P, Mares, M. | | Deposit date: | 2020-07-20 | | Release date: | 2021-06-02 | | Last modified: | 2024-10-23 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Mialostatin, a Novel Midgut Cystatin from Ixodes ricinus Ticks: Crystal Structure and Regulation of Host Blood Digestion.
Int J Mol Sci, 22, 2021
|
|
7PMU
 | | Crystal structure of native Iripin-8 | | Descriptor: | DI(HYDROXYETHYL)ETHER, HEXAETHYLENE GLYCOL, Serpin-8, ... | | Authors: | Polderdijk, S, Kotal, J, Chmelar, J, Huntington, J.A. | | Deposit date: | 2021-09-02 | | Release date: | 2021-10-13 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Ixodes ricinus Salivary Serpin Iripin-8 Inhibits the Intrinsic Pathway of Coagulation and Complement.
Int J Mol Sci, 22, 2021
|
|
7QTZ
 
 | | Crystal structure of Iripin-1 serpin from tick Ixodes ricinus | | Descriptor: | MAGNESIUM ION, Putative salivary serpin | | Authors: | Kascakova, B, Kuta Smatanova, I, Chmelar, J, Prudnikova, T. | | Deposit date: | 2022-01-17 | | Release date: | 2023-01-25 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Iripin-1, a new anti-inflammatory tick serpin, inhibits leukocyte recruitment in vivo while altering the levels of chemokines and adhesion molecules.
Front Immunol, 14, 2023
|
|
