1JRR
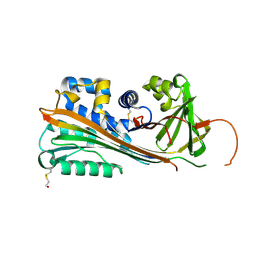 
 | | HUMAN PLASMINOGEN ACTIVATOR INHIBITOR-2.[LOOP (66-98) DELETIONMUTANT] COMPLEXED WITH PEPTIDE MIMIckING THE REACTIVE CENTER LOOP | | Descriptor: | BETA-MERCAPTOETHANOL, PLASMINOGEN ACTIVATOR INHIBITOR-2 | | Authors: | Jankova, L, Harrop, S.J, Saunders, D.N, Andrews, J.L, Bertram, K.C, Gould, A.R, Baker, M.S, Curmi, P.M.G. | | Deposit date: | 2001-08-15 | | Release date: | 2001-09-26 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.6 Å) | | Cite: | CRYSTAL STRUCTURE OF THE COMPLEX OF PLASMINOGEN ACTIVATOR INHIBITOR 2 WITH A PEPTIDE MIMICKING THE REACTIVE CENTER LOOP
J.Biol.Chem., 276, 2001
|
|
2ARQ
 
 | | Human plasminogen activator inhibitor-2.[loop (66-98) deletion mutant] complexed with peptide n-acetyl-teaaagdggvmtgr-oh | | Descriptor: | 14-mer from Plasminogen activator inhibitor-2, Plasminogen activator inhibitor-2 | | Authors: | Di Giusto, D.A, Sutherland, A.P, Jankova, L, Harrop, S.J, Curmi, P.M, King, G.C. | | Deposit date: | 2005-08-21 | | Release date: | 2006-07-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.85 Å) | | Cite: | Plasminogen activator inhibitor-2 is highly tolerant to P8 residue substitution--implications for serpin mechanistic model and prediction of nsSNP activities
J.Mol.Biol., 353, 2005
|
|
2ARR
 
 | | Human plasminogen activator inhibitor-2.[loop (66-98) deletion mutant] complexed with peptide n-acetyl-teaaagmggvmtgr-oh | | Descriptor: | 14-mer from Plasminogen activator inhibitor-2, Plasminogen activator inhibitor-2 | | Authors: | Di Giusto, D.A, Sutherland, A.P, Jankova, L, Harrop, S.J, Curmi, P.M, King, G.C. | | Deposit date: | 2005-08-21 | | Release date: | 2006-07-11 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.55 Å) | | Cite: | Plasminogen activator inhibitor-2 is highly tolerant to P8 residue substitution--implications for serpin mechanistic model and prediction of nsSNP activities
J.Mol.Biol., 353, 2005
|
|
1K0M
 
 | | Crystal structure of a soluble monomeric form of CLIC1 at 1.4 angstroms | | Descriptor: | CHLORIDE INTRACELLULAR CHANNEL PROTEIN 1 | | Authors: | Harrop, S.J, DeMaere, M.Z, Fairlie, W.D, Reztsova, T, Valenzuela, S.M, Mazzanti, M, Tonini, R, Qiu, M.R, Jankova, L, Warton, K, Bauskin, A.R, Wu, W.M, Pankhurst, S, Campbell, T.J, Breit, S.N, Curmi, P.M.G. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure of a soluble form of the intracellular chloride ion channel CLIC1 (NCC27) at 1.4-A resolution.
J.Biol.Chem., 276, 2001
|
|
1K0O
 
 | | Crystal structure of a soluble form of CLIC1. An intracellular chloride ion channel | | Descriptor: | CHLORIDE INTRACELLULAR CHANNEL PROTEIN 1 | | Authors: | Harrop, S.J, DeMaere, M.Z, Fairlie, W.D, Reztsova, T, Valenzuela, S.M, Mazzanti, M, Tonini, R, Qiu, M.R, Jankova, L, Warton, K, Bauskin, A.R, Wu, W.M, Pankhurst, S, Campbell, T.J, Breit, S.N, Curmi, P.M.G. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of a soluble form of the intracellular chloride ion channel CLIC1 (NCC27) at 1.4-A resolution.
J.Biol.Chem., 276, 2001
|
|
1K0N
 
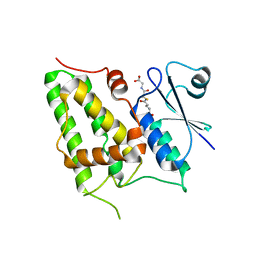 | | Chloride Intracellular Channel 1 (CLIC1) complexed with glutathione | | Descriptor: | CHLORIDE INTRACELLULAR CHANNEL PROTEIN 1, GLUTATHIONE | | Authors: | Harrop, S.J, DeMaere, M.Z, Fairlie, W.D, Reztsova, T, Valenzuela, S.M, Mazzanti, M, Tonini, R, Qiu, M.R, Jankova, L, Warton, K, Bauskin, A.R, Wu, W.M, Pankhurst, S, Campbell, T.J, Breit, S.N, Curmi, P.M.G. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-12 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Crystal structure of a soluble form of the intracellular chloride ion channel CLIC1 (NCC27) at 1.4-A resolution.
J.Biol.Chem., 276, 2001
|
|
1QGW
 
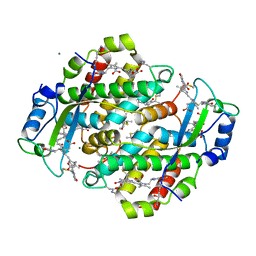 | | CRYSTAL STRUCTURE OF PHYCOERYTHRIN 545 FROM THE MARINE CRYPTOPHYTE RHODOMONAS CS24 | | Descriptor: | 15,16-DIHYDROBILIVERDIN, CHLORIDE ION, MAGNESIUM ION, ... | | Authors: | Harrop, S.J, Wilk, K.E, Hiller, R.G, Curmi, P.M.G. | | Deposit date: | 1999-05-10 | | Release date: | 1999-05-19 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.63 Å) | | Cite: | Evolution of a light-harvesting protein by addition of new subunits and rearrangement of conserved elements: crystal structure of a cryptophyte phycoerythrin at 1.63-A resolution.
Proc.Natl.Acad.Sci.USA, 96, 1999
|
|
1BY7
 
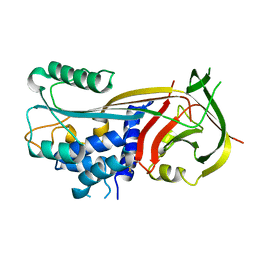 | |
1XF6
 
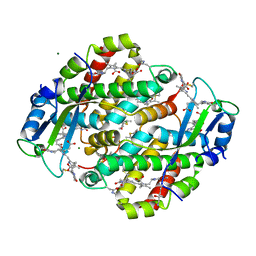 | | High resolution crystal structure of phycoerythrin 545 from the marine cryptophyte rhodomonas CS24 | | Descriptor: | 15,16-DIHYDROBILIVERDIN, B-phycoerythrin beta chain, CHLORIDE ION, ... | | Authors: | Doust, A.B, Marai, C.N.J, Harrop, S.J, Wilk, K.E, Curmi, P.M.G, Scholes, G.D. | | Deposit date: | 2004-09-14 | | Release date: | 2004-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (1.1 Å) | | Cite: | Developing a structure-function model for the cryptophyte phycoerythrin 545 using ultrahigh resolution crystallography and ultrafast laser spectroscopy
J.Mol.Biol., 344, 2004
|
|
1XG0
 
 | | High resolution crystal structure of phycoerythrin 545 from the marine cryptophyte rhodomonas CS24 | | Descriptor: | 15,16-DIHYDROBILIVERDIN, B-phycoerythrin beta chain, CHLORIDE ION, ... | | Authors: | Doust, A.B, Marai, C.N.J, Harrop, S.J, Wilk, K.E, Curmi, P.M.G, Scholes, G.D. | | Deposit date: | 2004-09-16 | | Release date: | 2004-11-30 | | Last modified: | 2023-10-25 | | Method: | X-RAY DIFFRACTION (0.97 Å) | | Cite: | Developing a structure-function model for the cryptophyte phycoerythrin 545 using ultrahigh resolution crystallography and ultrafast laser spectroscopy
J.Mol.Biol., 344, 2004
|
|
