3KP9
 
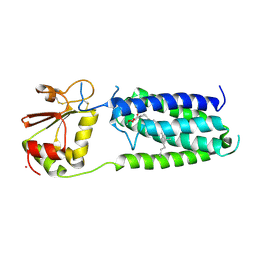 | | Structure of a bacterial homolog of vitamin K epoxide reductase | | Descriptor: | MERCURY (II) ION, UBIQUINONE-10, VKORC1/thioredoxin domain protein | | Authors: | Li, W, Schulman, S, Dutton, R.J, Boyd, D, Beckwith, J, Rapoport, T.A. | | Deposit date: | 2009-11-16 | | Release date: | 2010-02-09 | | Last modified: | 2021-10-13 | | Method: | X-RAY DIFFRACTION (3.6 Å) | | Cite: | Structure of a bacterial homologue of vitamin K epoxide reductase.
Nature, 463, 2010
|
|
2K73
 
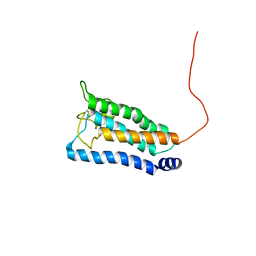 | | Solution NMR structure of integral membrane protein DsbB | | Descriptor: | Disulfide bond formation protein B | | Authors: | Zhou, Y, Cierpicki, T, Flores Jimenez, R.H, Lukasik, S.M, Ellena, J.F, Cafiso, D.S, Kadokura, H, Beckwith, J, Bushweller, J.H. | | Deposit date: | 2008-08-01 | | Release date: | 2008-10-07 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the integral membrane enzyme DsbB: functional insights into DsbB-catalyzed disulfide bond formation.
Mol.Cell, 31, 2008
|
|
2K74
 
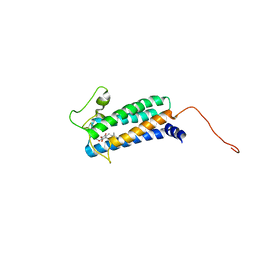 | | Solution NMR structure of DsbB-ubiquinone complex | | Descriptor: | Disulfide bond formation protein B, UBIQUINONE-2 | | Authors: | Zhou, Y, Cierpicki, T, Flores Jimenez, R.H, Lukasik, S.M, Ellena, J.F, Cafiso, D.S, Kadokura, H, Beckwith, J, Bushweller, J.H. | | Deposit date: | 2008-08-01 | | Release date: | 2008-10-07 | | Last modified: | 2021-10-20 | | Method: | SOLUTION NMR | | Cite: | NMR solution structure of the integral membrane enzyme DsbB: functional insights into DsbB-catalyzed disulfide bond formation.
Mol.Cell, 31, 2008
|
|
3KP8
 
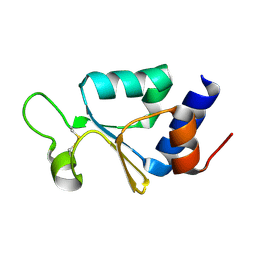 | | The thioredoxin-like domain of a VKOR homolog from Synechococcus sp. | | Descriptor: | VKORC1/thioredoxin domain protein | | Authors: | Li, W, Schulman, S, Dutton, R.J, Boyd, D, Beckwith, J, Rapoport, T.A. | | Deposit date: | 2009-11-15 | | Release date: | 2010-03-02 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.66 Å) | | Cite: | Structure of a bacterial homologue of vitamin K epoxide reductase.
Nature, 463, 2010
|
|
2PPT
 
 | |
1JZD
 
 | | DsbC-DsbDalpha complex | | Descriptor: | thiol:disulfide interchange protein dsbc, thiol:disulfide interchange protein dsbd | | Authors: | Haebel, P.W, Goldstone, D, Katzen, F, Beckwith, J, Metcalf, P. | | Deposit date: | 2001-09-15 | | Release date: | 2003-03-08 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | The Disulfide Bond Isomerase DsbC is Activated by an
Immunoglobulin-fold Thiol Oxidoreductase: Crystal structure of the
DsbC-DsbDalpha complex.
Embo J., 21, 2002
|
|
1JZO
 
 | | DsbC C101S | | Descriptor: | THIOL:DISULFIDE INTERCHANGE PROTEIN DSBC | | Authors: | Haebel, P.W, Goldstone, D, Katzen, F, Beckwith, J, Metcalf, P. | | Deposit date: | 2001-09-17 | | Release date: | 2003-03-08 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (1.92 Å) | | Cite: | The Disulfide Bond Isomerase DsbC is Activated by an
Immunoglobulin-fold Thiol Oxidoreductase: Crystal Structure of the
DsbC-DsbDalpha complex.
Embo J., 21, 2002
|
|
3HYP
 
 | |
3HXS
 
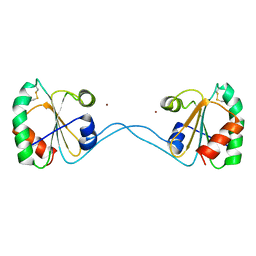 | | Crystal Structure of Bacteroides fragilis TrxP | | Descriptor: | Thioredoxin, ZINC ION | | Authors: | Shouldice, S.R. | | Deposit date: | 2009-06-22 | | Release date: | 2010-01-12 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.996 Å) | | Cite: | In vivo oxidative protein folding can be facilitated by oxidation-reduction cycling
Mol.Microbiol., 75, 2010
|
|
2YXR
 
 | |
2YXQ
 
 | |
1JPE
 
 | |
1G0T
 
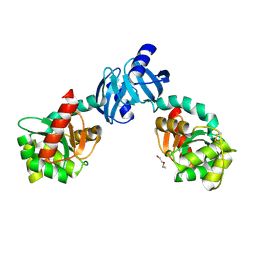 | | DSBC MUTANT C101S | | Descriptor: | DI(HYDROXYETHYL)ETHER, THIOL:DISULFIDE INTERCHANGE PROTEIN DSBC | | Authors: | Haebel, P.W, Metcalf, P. | | Deposit date: | 2000-10-09 | | Release date: | 2003-02-04 | | Last modified: | 2021-11-03 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Crystal structure of the protein disulfide bond isomerase, DsbC, from Escherichia coli
Embo J., 21, 2002
|
|
1A2J
 
 | | OXIDIZED DSBA CRYSTAL FORM II | | Descriptor: | DISULFIDE BOND FORMATION PROTEIN | | Authors: | Martin, J.L, Guddat, L.W. | | Deposit date: | 1998-01-06 | | Release date: | 1998-09-16 | | Last modified: | 2023-08-02 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Crystal structures of reduced and oxidized DsbA: investigation of domain motion and thiolate stabilization.
Structure, 6, 1998
|
|
1A2M
 
 | |
1A2L
 
 | | REDUCED DSBA AT 2.7 ANGSTROMS RESOLUTION | | Descriptor: | DISULFIDE BOND FORMATION PROTEIN | | Authors: | Martin, J.L, Guddat, L.W. | | Deposit date: | 1998-01-06 | | Release date: | 1998-07-08 | | Last modified: | 2024-05-22 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Crystal structures of reduced and oxidized DsbA: investigation of domain motion and thiolate stabilization.
Structure, 6, 1998
|
|
1DSB
 
 | |
1FVJ
 
 | |
1FVK
 
 | |
