3LSZ
 
 | | Crystal structure of glutathione s-transferase from Rhodobacter sphaeroides | | Descriptor: | 2-AMINO-2-HYDROXYMETHYL-PROPANE-1,3-DIOL, GLUTATHIONE, GLYCEROL, ... | | Authors: | Eswaramoorthy, S, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2010-02-14 | | Release date: | 2010-03-23 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal structure of glutathione s-transferase from Rhodobacter sphaeroides
To be Published
|
|
3LXZ
 
 | |
3LYK
 
 | |
3LYP
 
 | |
3LJR
 
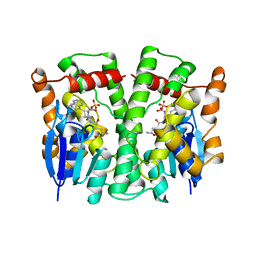 | | GLUTATHIONE TRANSFERASE (THETA CLASS) FROM HUMAN IN COMPLEX WITH THE GLUTATHIONE CONJUGATE OF 1-MENAPHTHYL SULFATE | | Descriptor: | 1-MENAPHTHYL GLUTATHIONE CONJUGATE, GLUTATHIONE S-TRANSFERASE, SULFATE ION | | Authors: | Rossjohn, J, Mckinstry, W.J, Oakley, A.J, Verger, D, Flanagan, J, Chelvanayagam, G, Tan, K.L, Board, P.G, Parker, M.W. | | Deposit date: | 1998-03-08 | | Release date: | 1999-03-23 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (3.3 Å) | | Cite: | Human theta class glutathione transferase: the crystal structure reveals a sulfate-binding pocket within a buried active site.
Structure, 6, 1998
|
|
3M3M
 
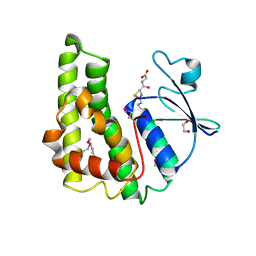 | | Crystal structure of glutathione S-transferase from Pseudomonas fluorescens [Pf-5] | | Descriptor: | 1,2-ETHANEDIOL, GLUTATHIONE, Glutathione S-transferase, ... | | Authors: | Bagaria, A, Burley, S.K, Swaminathan, S, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2010-03-09 | | Release date: | 2010-03-16 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | Crystal structure of glutathione S-transferase from Pseudomonas fluorescens [Pf-5]
To be Published
|
|
3MDK
 
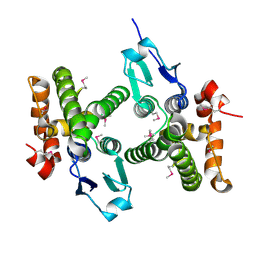 | |
3M8N
 
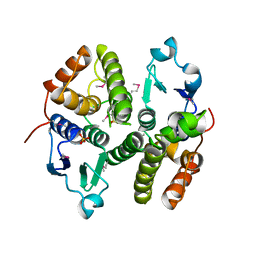 | |
3MAK
 
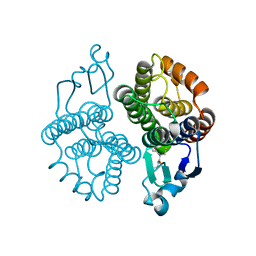 | |
2F8F
 
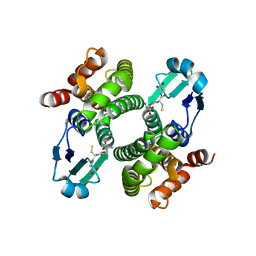 | | Crystal structure of the Y10F mutant of the gluathione s-transferase from schistosoma haematobium | | Descriptor: | GLUTATHIONE, Glutathione S-transferase 28 kDa | | Authors: | Gourlay, L.J, Baiocco, P, Angelucci, F, Miele, A.E, Bellelli, A, Brunori, M. | | Deposit date: | 2005-12-02 | | Release date: | 2006-07-04 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Probing the Mechanism of GSH Activation in Schistosoma haematobium Glutathione-S-transferase by Site-directed Mutagenesis and X-ray Crystallography.
J.Mol.Biol., 360, 2006
|
|
2F3M
 
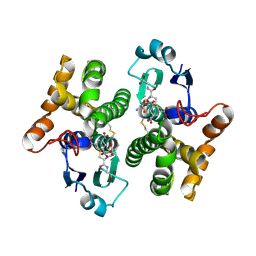 | | Structure of human GLUTATHIONE S-TRANSFERASE M1A-1A complexed with 1-(S-(GLUTATHIONYL)-2,4,6-TRINITROCYCLOHEXADIENATE ANION | | Descriptor: | 1-(S-GLUTATHIONYL)-2,4,6-TRINITROCYCLOHEXA-2,5-DIENE, Glutathione S-transferase Mu 1 | | Authors: | Patskovsky, Y, Patskovska, L, Almo, S.C, Listowsky, I. | | Deposit date: | 2005-11-21 | | Release date: | 2006-04-25 | | Last modified: | 2023-08-23 | | Method: | X-RAY DIFFRACTION (2.7 Å) | | Cite: | Transition state model and mechanism of nucleophilic aromatic substitution reactions catalyzed by human glutathione S-transferase M1a-1a.
Biochemistry, 45, 2006
|
|
3N5O
 
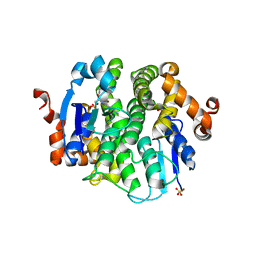 | |
2GDR
 
 | | Crystal structure of a bacterial glutathione transferase | | Descriptor: | GLUTATHIONE, glutathione S-transferase | | Authors: | Tocheva, E.I, Fortin, P.D, Eltis, L.D, Murphy, M.E.P. | | Deposit date: | 2006-03-16 | | Release date: | 2006-08-22 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | Structures of Ternary Complexes of BphK, a Bacterial Glutathione S-Transferase That Reductively Dechlorinates Polychlorinated Biphenyl Metabolites.
J.Biol.Chem., 281, 2006
|
|
5DQS
 
 | |
2GST
 
 | | STRUCTURE OF THE XENOBIOTIC SUBSTRATE BINDING SITE OF A GLUTATHIONE S-TRANSFERASE AS REVEALED BY X-RAY CRYSTALLOGRAPHIC ANALYSIS OF PRODUCT COMPLEXES WITH THE DIASTEREOMERS OF 9-(S-GLUTATHIONYL)-10-HYDROXY-9, 10-DIHYDROPHENANTHRENE | | Descriptor: | GLUTATHIONE S-TRANSFERASE, L-gamma-glutamyl-S-[(9S,10S)-10-hydroxy-9,10-dihydrophenanthren-9-yl]-L-cysteinylglycine, SULFATE ION | | Authors: | Ji, X, Armstrong, R.N, Gilliland, G.L. | | Deposit date: | 1993-06-07 | | Release date: | 1993-10-31 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.8 Å) | | Cite: | Structure and function of the xenobiotic substrate binding site of a glutathione S-transferase as revealed by X-ray crystallographic analysis of product complexes with the diastereomers of 9-(S-glutathionyl)-10-hydroxy-9,10-dihydrophenanthrene.
Biochemistry, 33, 1994
|
|
2GTU
 
 | | LIGAND-FREE HUMAN GLUTATHIONE S-TRANSFERASE M2-2 (E.C.2.5.1.18), MONOCLINIC CRYSTAL FORM | | Descriptor: | GLUTATHIONE S-TRANSFERASE | | Authors: | Patskovska, L.N, Fedorov, A.A, Patskovsky, Y.V, Almo, S.C, Listowsky, I. | | Deposit date: | 1998-05-26 | | Release date: | 1999-03-02 | | Last modified: | 2023-08-09 | | Method: | X-RAY DIFFRACTION (2.55 Å) | | Cite: | The enhanced affinity for thiolate anion and activation of enzyme-bound glutathione is governed by an arginine residue of human Mu class glutathione S-transferases.
J.Biol.Chem., 275, 2000
|
|
1K0D
 
 | | Ure2p in Complex with Glutathione | | Descriptor: | GLUTATHIONE, URE2 PROTEIN | | Authors: | Bousset, L, Belrhali, H, Melki, R, Morera, S. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Crystal structures of the yeast prion Ure2p functional region in complex with glutathione and related compounds.
Biochemistry, 40, 2001
|
|
4HJ2
 
 | | Crystal Structure Analysis of GSTA1-1 in complex with chlorambucil | | Descriptor: | Glutathione S-transferase A1, L-gamma-glutamyl-S-(2-{[4-(3-carboxypropyl)phenyl](2-chloroethyl)amino}ethyl)-L-cysteinylglycine | | Authors: | Karpusas, M, Chiniadis, L, Labrou, N.E. | | Deposit date: | 2012-10-12 | | Release date: | 2013-03-27 | | Last modified: | 2024-02-28 | | Method: | X-RAY DIFFRACTION (2.1 Å) | | Cite: | The interaction of the chemotherapeutic drug chlorambucil with human glutathione transferase A1-1: kinetic and structural analysis.
Plos One, 8, 2013
|
|
1JZR
 
 | | Ure2p in complex with glutathione | | Descriptor: | GLUTATHIONE, URE2 PROTEIN | | Authors: | Bousset, L, Belrhali, H, Melki, R, Morera, S. | | Deposit date: | 2001-09-17 | | Release date: | 2001-12-21 | | Last modified: | 2023-08-16 | | Method: | X-RAY DIFFRACTION (2.9 Å) | | Cite: | Crystal structures of the yeast prion Ure2p functional region in complex with glutathione and related compounds.
Biochemistry, 40, 2001
|
|
4HI7
 | | Crystal structure of glutathione transferase homolog from drosophilia mojavensis, TARGET EFI-501819, with bound glutathione | | Descriptor: | GI20122, GLUTATHIONE | | Authors: | Vetting, M.W, Toro, R, Bhosle, R, Al Obaidi, N.F, Morisco, L.L, Wasserman, S.R, Sojitra, S, Washington, E, Scott Glenn, A, Chowdhury, S, Evans, B, Hammonds, J, Hillerich, B, Love, J, Seidel, R.D, Imker, H.J, Armstrong, R.N, Gerlt, J.A, Almo, S.C, Enzyme Function Initiative (EFI) | | Deposit date: | 2012-10-11 | | Release date: | 2012-10-31 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.25 Å) | | Cite: | Crystal structure of glutathione transferase homolog from drosophilia mojavensis, TARGET EFI-501819, with bound glutathione
To be Published
|
|
1K0B
 
 | | Ure2p in Complex with Glutathione | | Descriptor: | GLUTATHIONE, URE2 PROTEIN | | Authors: | Bousset, L, Belrhali, H, Melki, R, Morera, S. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structures of the yeast prion Ure2p functional region in complex with glutathione and related compounds.
Biochemistry, 40, 2001
|
|
1JLV
 
 | | Anopheles dirus species B glutathione S-transferases 1-3 | | Descriptor: | GLUTATHIONE, glutathione transferase GST1-3 | | Authors: | Oakley, A.J, Harnnoi, T, Udomsinprasert, R, Jirajaroenrat, K, Ketterman, A.J, Wilce, M.C. | | Deposit date: | 2001-07-16 | | Release date: | 2002-07-16 | | Last modified: | 2024-04-03 | | Method: | X-RAY DIFFRACTION (1.75 Å) | | Cite: | The crystal structures of glutathione S-transferases isozymes 1-3 and 1-4 from Anopheles dirus species B.
Protein Sci., 10, 2001
|
|
1K0C
 
 | | Ure2p in complex with S-p-nitrobenzylglutathione | | Descriptor: | GLUTATHIONE, S-(P-NITROBENZYL)GLUTATHIONE, URE2 PROTEIN | | Authors: | Bousset, L, Belrhali, H, Melki, R, Morera, S. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structures of the yeast prion Ure2p functional region in complex with glutathione and related compounds.
Biochemistry, 40, 2001
|
|
4HOJ
 
 | | Crystal structure of glutathione transferase homolog from Neisseria Gonorrhoeae, target EFI-501841, with bound glutathione | | Descriptor: | ACETATE ION, CALCIUM ION, GLUTATHIONE, ... | | Authors: | Vetting, M.W, Toro, R, Bhosle, R, Al Obaidi, N.F, Morisco, L.L, Wasserman, S.R, Sojitra, S, Washington, E, Scott Glenn, A, Chowdhury, S, Evans, B, Hammonds, J, Hillerich, B, Love, J, Seidel, R.D, Imker, H.J, Armstrong, R.N, Gerlt, J.A, Almo, S.C, Enzyme Function Initiative (EFI) | | Deposit date: | 2012-10-22 | | Release date: | 2012-11-07 | | Last modified: | 2023-09-20 | | Method: | X-RAY DIFFRACTION (1.4 Å) | | Cite: | Crystal structure of glutathione transferase homolog from Neisseria Gonorrhoeae, target EFI-501841, with bound glutathione
To be Published
|
|
1K0A
 
 | | Ure2p in Complex with S-hexylglutathione | | Descriptor: | GLUTATHIONE, S-HEXYLGLUTATHIONE, URE2 PROTEIN | | Authors: | Bousset, L, Belrhali, H, Melki, R, Morera, S. | | Deposit date: | 2001-09-19 | | Release date: | 2001-12-21 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Crystal structures of the yeast prion Ure2p functional region in complex with glutathione and related compounds.
Biochemistry, 40, 2001
|
|
