4EQ7
 
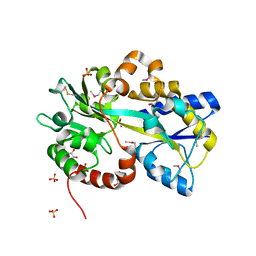 | | Structure of Atu4243-GABA receptor | | Descriptor: | ABC transporter, substrate binding protein (Polyamine), GLYCEROL, ... | | Authors: | Morera, S, Planamente, S. | | Deposit date: | 2012-04-18 | | Release date: | 2012-11-21 | | Last modified: | 2012-12-19 | | Method: | X-RAY DIFFRACTION (1.91 Å) | | Cite: | Structural basis for selective GABA binding in bacterial pathogens.
Mol.Microbiol., 86, 2012
|
|
4EA4
 
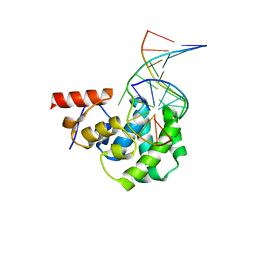 | | Structure of the glycosylase domain of MBD4 bound to 5hmU-containing DNA | | Descriptor: | DNA (5'-D(*CP*CP*AP*GP*CP*GP*(5HU)P*GP*CP*AP*GP*C)-3'), DNA (5'-D(*GP*CP*TP*GP*CP*GP*CP*GP*CP*TP*GP*G)-3'), Methyl-CpG-binding domain protein 4 | | Authors: | Morera, S, Vigouroux, A. | | Deposit date: | 2012-03-22 | | Release date: | 2012-08-08 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (2 Å) | | Cite: | Biochemical and structural characterization of the glycosylase domain of MBD4 bound to thymine and 5-hydroxymethyuracil-containing DNA.
Nucleic Acids Res., 40, 2012
|
|
8OF3
 
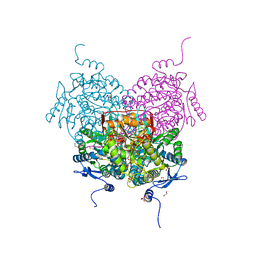 | | Structure of the apoform of ALDEHYDE DEHYDROGENASE 5F1 (ALDH5F1) from the moss Physcomitrium patens | | Descriptor: | 1,2-ETHANEDIOL, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, PENTAETHYLENE GLYCOL, ... | | Authors: | Morera, S, Kopecny, D, Vigouroux, A. | | Deposit date: | 2023-03-13 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.348 Å) | | Cite: | A study on abiotic stress responses of aldehyde dehydrogenase (ALDH) superfamilies in moss and barley focused on members linked to the GABA shunt pathway
To Be Published
|
|
8OFM
 
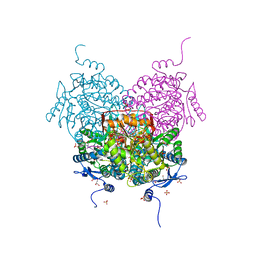 | |
8OF1
 
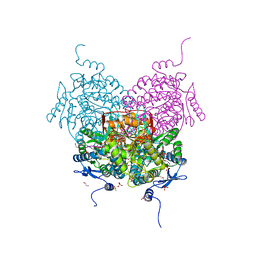 | | Structure of ALDH5F1 from moss Physcomitrium patens in complex with NAD+ in the contracted conformation | | Descriptor: | 1,2-ETHANEDIOL, 3,6,9,12,15,18-HEXAOXAICOSANE-1,20-DIOL, DI(HYDROXYETHYL)ETHER, ... | | Authors: | Morera, S, Kopecny, D, Vigouroux, A. | | Deposit date: | 2023-03-13 | | Release date: | 2024-03-27 | | Method: | X-RAY DIFFRACTION (1.81 Å) | | Cite: | A study on abiotic stress responses of aldehyde dehydrogenase (ALDH) superfamilies in moss and barley focused on members linked to the GABA shunt pathway
To Be Published
|
|
7Z8E
 
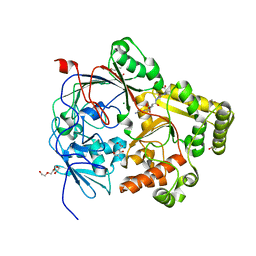 | | Crystal structure of the substrate-binding protein YejA from S. meliloti in complex with peptide fragment | | Descriptor: | 1,2-ETHANEDIOL, ABC transporter substrate-binding protein, GLY-SER-ASP-VAL-ALA, ... | | Authors: | Morera, S, Vigouroux, V, Travin, D.Y. | | Deposit date: | 2022-03-17 | | Release date: | 2023-02-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Dual-Uptake Mode of the Antibiotic Phazolicin Prevents Resistance Acquisition by Gram-Negative Bacteria.
Mbio, 14, 2023
|
|
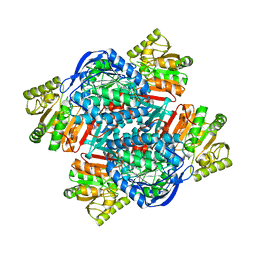
6QAK
 | |
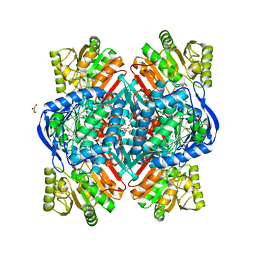
6QAP
 | |
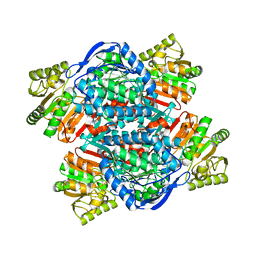
6QAO
 | |
3N2S
 
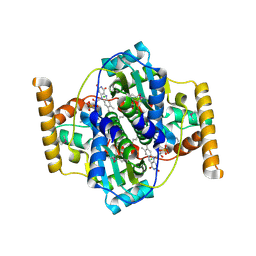 | | Structure of NfrA1 nitroreductase from B. subtilis | | Descriptor: | CHLORIDE ION, FLAVIN MONONUCLEOTIDE, NADPH-dependent nitro/flavin reductase | | Authors: | Morera, S, Gueguen-Chaignon, V, Meyer, P, Cortial, S, Ouazzani, J. | | Deposit date: | 2010-05-19 | | Release date: | 2010-09-15 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | NADH oxidase activity of Bacillus subtilis nitroreductase NfrA1: insight into its biological role.
Febs Lett., 584, 2010
|
|
7ZHC
 
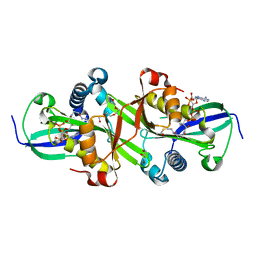 | | Moss spermine/spermidine acetyl transferase (PpSSAT) in complex with AcetylCoA and polyethylen glycol | | Descriptor: | ACETYL COENZYME *A, GLYCEROL, N-acetyltransferase domain-containing protein, ... | | Authors: | Morera, S, Kopecny, D, Vigouroux, A. | | Deposit date: | 2022-04-06 | | Release date: | 2023-03-15 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.819 Å) | | Cite: | Biochemical and structural basis of polyamine, lysine and ornithine acetylation catalyzed by spermine/spermidine N-acetyl transferase in moss and maize.
Plant J., 114, 2023
|
|
7ZKT
 
 | | Moss spermine/spermidine acetyl transferase (PpSSAT) in complex with CoA and lysine | | Descriptor: | 1,2-ETHANEDIOL, COENZYME A, LYSINE, ... | | Authors: | Morera, S, Kopecny, D, Vigouroux, A, Briozzo, P. | | Deposit date: | 2022-04-13 | | Release date: | 2023-03-22 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (2.06 Å) | | Cite: | Biochemical and structural basis of polyamine, lysine and ornithine acetylation catalyzed by spermine/spermidine N-acetyl transferase in moss and maize.
Plant J., 114, 2023
|
|
3FKB
 
 | | Structure of NDPK H122G and tenofovir-diphosphate | | Descriptor: | 1,2-ETHANEDIOL, GLYCEROL, MAGNESIUM ION, ... | | Authors: | Morera, S, Chen, Y.X. | | Deposit date: | 2008-12-16 | | Release date: | 2009-09-29 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (1.65 Å) | | Cite: | Nucleoside diphosphate kinase and the activation of antiviral phosphonate analogs of nucleotides: binding mode and phosphorylation of tenofovir derivatives
Nucleosides Nucleotides Nucleic Acids, 28, 2009
|
|
7R4U
 | | Apoform of FtrA/P19 from Rubrivivax gelatinosus | | Descriptor: | FtrA-P19, GLYCEROL, SODIUM ION, ... | | Authors: | Morera, S, Vigouroux, A, Plancqueel, S. | | Deposit date: | 2022-02-09 | | Release date: | 2022-05-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.23 Å) | | Cite: | New insights into the mechanism of iron transport through the bacterial Ftr system present in pathogens.
Febs J., 289, 2022
|
|
3FJO
 
 | | Structure of chimeric YH CPR | | Descriptor: | FLAVIN MONONUCLEOTIDE, FLAVIN-ADENINE DINUCLEOTIDE, NADPH-cytochrome P450 reductase | | Authors: | Morera, S, Aigrain, L, Truan, G. | | Deposit date: | 2008-12-15 | | Release date: | 2009-06-16 | | Last modified: | 2023-11-01 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of the open conformation of a functional chimeric NADPH cytochrome P450 reductase
Embo Rep., 10, 2009
|
|
7R3P
 
 | |
7R4Z
 
 | |
7R5E
 | |
7R5P
 
 | |
7R4V
 
 | |
7R3S
 
 | | FtrA/P19 of Rubrivivax gelatinosus in complex with Ni | | Descriptor: | FtrA-P19 protein, GLYCEROL, NICKEL (II) ION, ... | | Authors: | Morera, S, Vigouroux, A. | | Deposit date: | 2022-02-07 | | Release date: | 2022-05-18 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.79 Å) | | Cite: | New insights into the mechanism of iron transport through the bacterial Ftr system present in pathogens.
Febs J., 289, 2022
|
|
7R5G
 
 | |
1C3J
 
 | | T4 PHAGE BETA-GLUCOSYLTRANSFERASE: SUBSTRATE BINDING AND PROPOSED CATALYTIC MECHANISM | | Descriptor: | BETA-GLUCOSYLTRANSFERASE, URIDINE-5'-DIPHOSPHATE | | Authors: | Morera, S, Imberty, A, Aschke-Sonnenborn, U, Ruger, W, Freemont, P.S. | | Deposit date: | 1999-07-28 | | Release date: | 1999-08-09 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | T4 phage beta-glucosyltransferase: substrate binding and proposed catalytic mechanism.
J.Mol.Biol., 292, 1999
|
|
6ZK1
 
 | | Plant nucleoside hydrolase - ZmNRh2b enzyme | | Descriptor: | 1,2-ETHANEDIOL, CALCIUM ION, CHLORIDE ION, ... | | Authors: | Morera, S, Vigouroux, A, Kopecny, D. | | Deposit date: | 2020-06-29 | | Release date: | 2022-01-12 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (1.99 Å) | | Cite: | Plant nucleoside N-ribohydrolases: riboside binding and role in nitrogen storage mobilization.
Plant J., 2023
|
|
6ZK2
 
 | | Plant nucleoside hydrolase - ZmNRh2b in complex with forodesine | | Descriptor: | 1,2-ETHANEDIOL, 1,4-DIDEOXY-4-AZA-1-(S)-(9-DEAZAHYPOXANTHIN-9-YL)-D-RIBITOL, CALCIUM ION, ... | | Authors: | Morera, S, Vigouroux, A, Kopecny, D. | | Deposit date: | 2020-06-29 | | Release date: | 2022-01-12 | | Last modified: | 2024-01-31 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Plant nucleoside N-ribohydrolases: riboside binding and role in nitrogen storage mobilization.
Plant J., 2023
|
|
