3D3E
 
 | | Crystal Structure of Human 11-beta-Hydroxysteroid Dehydrogenase (HSD1) in Complex with Benzamide Inhibitor | | Descriptor: | Corticosteroid 11-beta-dehydrogenase isozyme 1, N-cyclopropyl-N-(trans-4-pyridin-3-ylcyclohexyl)-4-[(1S)-2,2,2-trifluoro-1-hydroxy-1-methylethyl]benzamide, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Wang, Z, Sudom, A, Liu, J, Walker, N.P. | | Deposit date: | 2008-05-09 | | Release date: | 2008-07-01 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Discovery of Novel, Potent Benzamide Inhibitors of 11beta-Hydroxysteroid Dehydrogenase Type 1 (11beta-HSD1) Exhibiting Oral Activity in an Enzyme Inhibition ex Vivo Model
J.Med.Chem., 51, 2008
|
|
3DWF
 
 | | Crystal Structure of the Guinea Pig 11beta-Hydroxysteroid Dehydrogenase Type 1 Mutant F278E | | Descriptor: | 11-beta-hydroxysteroid dehydrogenase 1, GLYCEROL, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE, ... | | Authors: | Lawson, A.J, Ride, J.P, White, S.A, Walker, E.A. | | Deposit date: | 2008-07-22 | | Release date: | 2009-08-11 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.2 Å) | | Cite: | Mutations of key hydrophobic surface residues of 11 beta-hydroxysteroid dehydrogenase type 1 increase solubility and monodispersity in a bacterial expression system
Protein Sci., 18, 2009
|
|
3FRJ
 
 | | Crystal Structure of 11b-Hydroxysteroid Dehydrogenase-1 (11b-HSD1) in Complex with Piperidyl Benzamide Inhibitor | | Descriptor: | Corticosteroid 11-beta-Dehydrogenase, Isozyme 1, N-{1-[(1-carbamoylcyclopropyl)methyl]piperidin-4-yl}-N-cyclopropyl-4-[(1S)-2,2,2-trifluoro-1-hydroxy-1-methylethyl]benzamide, ... | | Authors: | Wang, Z, Sudom, A, Walker, N.P. | | Deposit date: | 2009-01-08 | | Release date: | 2009-06-16 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Discovery and optimization of piperidyl benzamide derivatives as a novel class of 11beta-HSD1 inhibitors.
Bioorg.Med.Chem.Lett., 19, 2009
|
|
3G1T
 
 | | CRYSTAL STRUCTURE OF short chain dehydrogenase from Salmonella enterica subsp. enterica serovar Typhi str. CT18 | | Descriptor: | MAGNESIUM ION, short chain dehydrogenase | | Authors: | Malashkevich, V.N, Toro, R, Sauder, J.M, Burley, S.K, Almo, S.C, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2009-01-30 | | Release date: | 2009-02-10 | | Last modified: | 2024-02-21 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | CRYSTAL STRUCTURE OF short chain dehydrogenase from Salmonella enterica subsp. enterica
serovar Typhi str. CT18
To be Published
|
|
3E03
 
 | | Crystal structure of a putative dehydrogenase from Xanthomonas campestris | | Descriptor: | CALCIUM ION, Short chain dehydrogenase | | Authors: | Sampathkumar, P, Wasserman, S, Rutter, M, Hu, S, Bain, K, Rodgers, L, Atwell, S, Sauder, J.M, Burley, S.K, New York SGX Research Center for Structural Genomics (NYSGXRC) | | Deposit date: | 2008-07-30 | | Release date: | 2008-09-16 | | Last modified: | 2021-02-10 | | Method: | X-RAY DIFFRACTION (1.69 Å) | | Cite: | Crystal structure of a putative dehydrogenase from Xanthomonas campestris
To be Published
|
|
3G49
 
 | |
7ONU
 
 | | Structure of human mitochondrial RNase P in complex with mitochondrial pre-tRNA-Tyr | | Descriptor: | 3-hydroxyacyl-CoA dehydrogenase type-2, MAGNESIUM ION, Mitochondrial Precursor tRNA-Tyr, ... | | Authors: | Bhatta, A, Dienemann, C, Cramer, P, Hillen, H.S. | | Deposit date: | 2021-05-26 | | Release date: | 2021-08-11 | | Last modified: | 2021-09-29 | | Method: | ELECTRON MICROSCOPY (3 Å) | | Cite: | Structural basis of RNA processing by human mitochondrial RNase P.
Nat.Struct.Mol.Biol., 28, 2021
|
|
5L53
 
 | | Menthone neomenthol reductase from Mentha piperita in complex with NADP | | Descriptor: | (-)-menthone:(+)-neomenthol reductase, NADP NICOTINAMIDE-ADENINE-DINUCLEOTIDE PHOSPHATE | | Authors: | Karuppiah, V, Toogood, H.S, Leys, D, Scrutton, N.S. | | Deposit date: | 2016-05-27 | | Release date: | 2016-08-31 | | Last modified: | 2024-01-10 | | Method: | X-RAY DIFFRACTION (2.24 Å) | | Cite: | Pinpointing a Mechanistic Switch Between Ketoreduction and "Ene" Reduction in Short-Chain Dehydrogenases/Reductases.
Angew.Chem.Int.Ed.Engl., 55, 2016
|
|
5L51
 
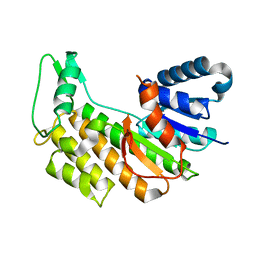 | |
2CDH
 
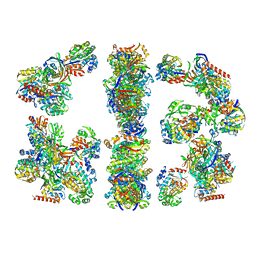 | | ARCHITECTURE OF THE THERMOMYCES LANUGINOSUS FUNGAL FATTY ACID SYNTHASE AT 5 ANGSTROM RESOLUTION. | | Descriptor: | DEHYDRATASE, ENOYL REDUCTASE, KETOACYL REDUCTASE, ... | | Authors: | Jenni, S, Leibundgut, M, Maier, T, Ban, N. | | Deposit date: | 2006-01-24 | | Release date: | 2006-03-07 | | Last modified: | 2024-02-14 | | Method: | X-RAY DIFFRACTION (4.2 Å) | | Cite: | Architecture of a Fungal Fatty Acid Synthase at 5 A Resolution.
Science, 311, 2006
|
|
2DKN
 
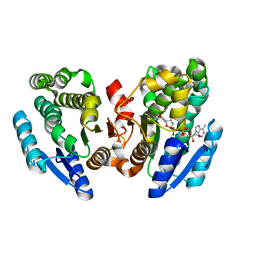 | |
2XOX
 
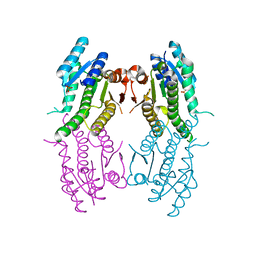 | | Crystal structure of pteridine reductase (PTR1) from Leishmania donovani | | Descriptor: | PTERIDINE REDUCTASE, SULFATE ION | | Authors: | Barrack, K.L, Tulloch, L.B, Burke, L.A, Fyfe, P.K, Hunter, W.N. | | Deposit date: | 2010-08-24 | | Release date: | 2011-01-12 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.5 Å) | | Cite: | Structure of Recombinant Leishmania Donovani Pteridine Reductase Reveals a Disordered Active Site.
Acta Crystallogr.,Sect.F, 67, 2011
|
|
1FJH
 
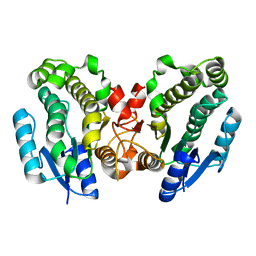 | | THE CRYSTAL STRUCTURE OF 3-ALPHA-HYDROXYSTEROID DEHYDROGENASE FROM COMAMONAS TESTOSTERONI, A MEMBER OF THE SHORT CHAIN DEHYDROGENASE/REDUCTASE FAMILY | | Descriptor: | 3ALPHA-HYDROXYSTEROID DEHYDROGENASE/CARBONYL REDUCTASE | | Authors: | Grimm, C, Ficner, R, Maser, E, Klebe, G, Reuter, K. | | Deposit date: | 2000-08-08 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.68 Å) | | Cite: | The crystal structure of 3alpha -hydroxysteroid dehydrogenase/carbonyl reductase from Comamonas testosteroni shows a novel oligomerization pattern within the short chain dehydrogenase/reductase family.
J.Biol.Chem., 275, 2000
|
|
1FK8
 
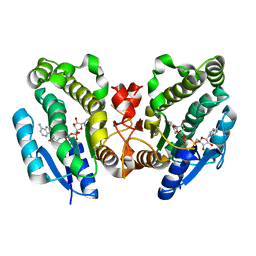 | | THE CRYSTAL STRUCTURE OF THE BINARY COMPLEX WITH NAD OF 3-ALPHA-HYDROXYSTEROID DEHYDROGENASE FROM COMAMONAS TESTOSTERONI, A MEMBER OF THE SHORT CHAIN DEHYDROGENASE/REDUCTASE FAMILY | | Descriptor: | 3ALPHA-HYDROXYSTEROID DEHYDROGENASE/CARBONYL REDUCTASE, NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Grimm, C, Ficner, R, Maser, E, Klebe, G, Reuter, K. | | Deposit date: | 2000-08-09 | | Release date: | 2001-01-17 | | Last modified: | 2024-02-07 | | Method: | X-RAY DIFFRACTION (1.95 Å) | | Cite: | The crystal structure of 3alpha -hydroxysteroid dehydrogenase/carbonyl reductase from Comamonas testosteroni shows a novel oligomerization pattern within the short chain dehydrogenase/reductase family.
J.Biol.Chem., 275, 2000
|
|
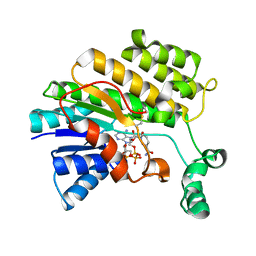
3O26
 
 | |
3OML
 
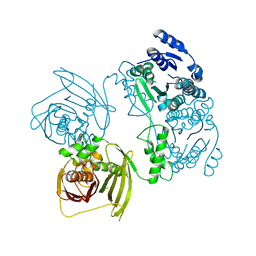 | | Structure of full-length peroxisomal multifunctional enzyme type 2 from Drosophila melanogaster | | Descriptor: | Peroxisomal Multifunctional Enzyme Type 2, CG3415 | | Authors: | Haataja, T.J.K, Koski, M.K, Glumoff, T, Hiltunen, J.K. | | Deposit date: | 2010-08-27 | | Release date: | 2011-03-09 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Peroxisomal multifunctional enzyme type 2 from the fruitfly: dehydrogenase and hydratase act as separate entities, as revealed by structure and kinetics.
Biochem.J., 435, 2011
|
|
3WYE
 
 | | Crystal Structure of chimeric engineered (2S,3S)-butanediol dehydrogenase complexed with NAD+ | | Descriptor: | Diacetyl reductase [(S)-acetoin forming],L-2,3-butanediol dehydrogenase,Diacetyl reductase [(S)-acetoin forming],L-2,3-butanediol dehydrogenase,Diacetyl reductase [(S)-acetoin forming],L-2,3-butanediol dehydrogenase,Diacetyl reductase [(S)-acetoin forming], NICOTINAMIDE-ADENINE-DINUCLEOTIDE | | Authors: | Shimegi, T, Oyama, T, Kusunoki, M, Ui, S. | | Deposit date: | 2014-08-26 | | Release date: | 2015-08-26 | | Last modified: | 2024-03-20 | | Method: | X-RAY DIFFRACTION (1.58 Å) | | Cite: | Crystal Structure of chimeric engineered (2S,3S)-butanediol dehydrogenase complexed with NAD+
To be Published
|
|
