2Q9T
 
 | | High-resolution structure of the DING protein from Pseudomonas fluorescens | | Descriptor: | 1,2-ETHANEDIOL, ACETATE ION, DING, ... | | Authors: | Moniot, S, Ahn, S, Elias, M, Kim, D, Scott, K, Chabriere, E. | | Deposit date: | 2007-06-14 | | Release date: | 2008-06-17 | | Last modified: | 2023-08-30 | | Method: | X-RAY DIFFRACTION (1.43 Å) | | Cite: | Structure-function relationships in a bacterial DING protein.
Febs Lett., 581, 2007
|
|
6YII
 
 | |
6FFM
 
 | |
3CMC
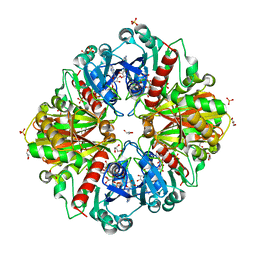 
 | | Thioacylenzyme intermediate of Bacillus stearothermophilus phosphorylating GAPDH | | Descriptor: | 1,2-ETHANEDIOL, GLYCERALDEHYDE-3-PHOSPHATE, GLYCEROL, ... | | Authors: | Moniot, S, Vonrhein, C, Bricogne, G, Didierjean, C, Corbier, C. | | Deposit date: | 2008-03-21 | | Release date: | 2008-06-17 | | Last modified: | 2011-07-13 | | Method: | X-RAY DIFFRACTION (1.77 Å) | | Cite: | Trapping of the Thioacylglyceraldehyde-3-phosphate Dehydrogenase Intermediate from Bacillus stearothermophilus: DIRECT EVIDENCE FOR A FLIP-FLOP MECHANISM
J.Biol.Chem., 283, 2008
|
|
5MAT
 
 | | Structure of human Sirtuin 2 in complex with a selective thienopyrimidinone based inhibitor | | Descriptor: | (7~{R})-7-[(3,5-dimethyl-1,2-oxazol-4-yl)methylamino]-3-[(4-methoxynaphthalen-1-yl)methyl]-5,6,7,8-tetrahydro-[1]benzothiolo[2,3-d]pyrimidin-4-one, HEXAETHYLENE GLYCOL, NAD-dependent protein deacetylase sirtuin-2, ... | | Authors: | Moniot, S, Steegborn, C. | | Deposit date: | 2016-11-04 | | Release date: | 2017-02-08 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (2.069 Å) | | Cite: | Thienopyrimidinone Based Sirtuin-2 (SIRT2)-Selective Inhibitors Bind in the Ligand Induced Selectivity Pocket.
J. Med. Chem., 60, 2017
|
|
5MAR
 
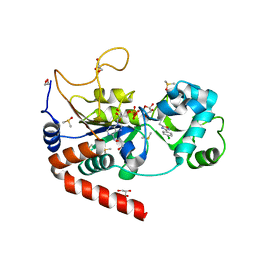 | | Structure of human SIRT2 in complex with 1,2,4-Oxadiazole inhibitor and ADP ribose. | | Descriptor: | 1,2-ETHANEDIOL, 3-[3-(4-chlorophenyl)-1,2,4-oxadiazol-5-yl]propan-1-ol, ACETATE ION, ... | | Authors: | Moniot, S, Steegborn, C. | | Deposit date: | 2016-11-04 | | Release date: | 2017-03-15 | | Last modified: | 2024-01-17 | | Method: | X-RAY DIFFRACTION (1.89 Å) | | Cite: | Development of 1,2,4-Oxadiazoles as Potent and Selective Inhibitors of the Human Deacetylase Sirtuin 2: Structure-Activity Relationship, X-ray Crystal Structure, and Anticancer Activity.
J. Med. Chem., 60, 2017
|
|
3ZGO
 
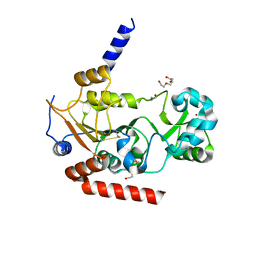 | |
3ZGV
 
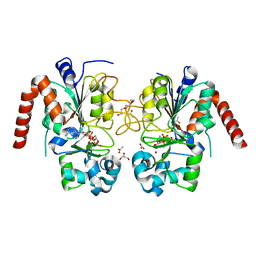 | | Structure of human SIRT2 in complex with ADP-ribose | | Descriptor: | ACETATE ION, GLYCEROL, NAD-DEPENDENT PROTEIN DEACETYLASE SIRTUIN-2, ... | | Authors: | Moniot, S, Steegborn, C. | | Deposit date: | 2012-12-19 | | Release date: | 2013-03-13 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.27 Å) | | Cite: | Crystal Structure Analysis of Human Sirt2 and its Adp-Ribose Complex
J.Struct.Biol., 182, 2013
|
|
2X5K
 
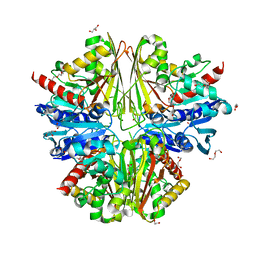 | | Structure of an active site mutant of the D-Erythrose-4-Phosphate Dehydrogenase from E. coli | | Descriptor: | 1,2-ETHANEDIOL, CHLORIDE ION, D-ERYTHROSE-4-PHOSPHATE DEHYDROGENASE, ... | | Authors: | Moniot, S, Didierjean, C, Boschi-Muller, S, Branlant, G, Corbier, C. | | Deposit date: | 2010-02-10 | | Release date: | 2011-02-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.37 Å) | | Cite: | Structural Characterization of Erythrose-4- Phosphate Dehydrogenase from Escherichia Coli: Peculiar Features When Compared to Phosphorylating Gapdhs
To be Published
|
|
2XF8
 
 | | Structure of the D-Erythrose-4-Phosphate Dehydrogenase from E. coli in complex with a NAD cofactor analog (3-Chloroacetyl adenine pyridine dinucleotide) and sulfate anion | | Descriptor: | 3-(CHLOROACETYL) PYRIDINE ADENINE DINUCLEOTIDE, D-ERYTHROSE-4-PHOSPHATE DEHYDROGENASE, SULFATE ION | | Authors: | Moniot, S, Didierjean, C, Boschi-Muller, S, Branlant, G, Corbier, C. | | Deposit date: | 2010-05-20 | | Release date: | 2011-06-08 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.95 Å) | | Cite: | Structural Characterization of Erythrose-4- Phosphate Dehydrogenase from Escherichia Coli: Peculiar Features When Compared to Phosphorylating Gapdhs
To be Published
|
|
2X5J
 
 | | Crystal structure of the Apoform of the D-Erythrose-4-phosphate dehydrogenase from E. coli | | Descriptor: | D-ERYTHROSE-4-PHOSPHATE DEHYDROGENASE, PHOSPHATE ION | | Authors: | Moniot, S, Didierjean, C, Boschi-Muller, S, Branlant, G, Corbier, C. | | Deposit date: | 2010-02-09 | | Release date: | 2011-02-16 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.3 Å) | | Cite: | Structural Characterization of Erythrose-4- Phosphate Dehydrogenase from Escherichia Coli: Peculiar Features When Compared to Phosphorylating Gapdhs
To be Published
|
|
4F1U
 
 | | Subatomic resolution structure of a high affinity periplasmic phosphate-binding protein (PfluDING) bound with phosphate at pH 4.5 | | Descriptor: | 1,2-ETHANEDIOL, HYDROGENPHOSPHATE ION, Putative alkaline phosphatase, ... | | Authors: | Liebschner, D, Elias, M, Tawfik, D.S, Moniot, S, Fournier, B, Scott, K, Jelsch, C, Guillot, B, Lecomte, C, Chabriere, E. | | Deposit date: | 2012-05-07 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (0.98 Å) | | Cite: | The molecular basis of phosphate discrimination in arsenate-rich environments.
Nature, 491, 2012
|
|
4F1V
 
 | | Subatomic resolution structure of a high affinity periplasmic phosphate-binding protein (PfluDING) bound with phosphate at pH 8.5 | | Descriptor: | HYDROGENPHOSPHATE ION, Putative alkaline phosphatase, SULFATE ION | | Authors: | Liebschner, D, Elias, M, Tawfik, D.S, Moniot, S, Fournier, B, Scott, K, Jelsch, C, Guillot, B, Lecomte, C, Chabriere, E. | | Deposit date: | 2012-05-07 | | Release date: | 2012-05-23 | | Last modified: | 2023-09-13 | | Method: | X-RAY DIFFRACTION (0.88 Å) | | Cite: | The molecular basis of phosphate discrimination in arsenate-rich environments.
Nature, 491, 2012
|
|
6HMJ
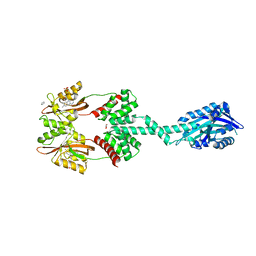 
 | | Structure of an RNA-binding Light-Oxygen-Voltage Receptor | | Descriptor: | 1,2-ETHANEDIOL, FLAVIN MONONUCLEOTIDE, GLYCEROL, ... | | Authors: | Ziegler, T, Moniot, S, Moeglich, A. | | Deposit date: | 2018-09-12 | | Release date: | 2019-08-28 | | Last modified: | 2020-09-09 | | Method: | X-RAY DIFFRACTION (2.51 Å) | | Cite: | A blue light receptor that mediates RNA binding and translational regulation.
Nat.Chem.Biol., 15, 2019
|
|
3ZSF
 
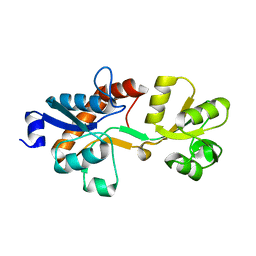 | | Crystal structure of the L-cystine solute receptor of Neisseria gonorrhoeae in the unliganded open conformation | | Descriptor: | ABC TRANSPORTER, PERIPLASMIC BINDING PROTEIN, AMINO ACID | | Authors: | Bulut, H, Moniot, S, Scheffel, F, Gathmann, S, Licht, A, Saenger, W, Schneider, E. | | Deposit date: | 2011-06-27 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.32 Å) | | Cite: | Crystal Structures of Two Solute Receptors for L-Cystine and L-Cysteine, Respectively, of the Human Pathogen Neisseria Gonorrhoeae.
J.Mol.Biol., 415, 2012
|
|
2VC5
 
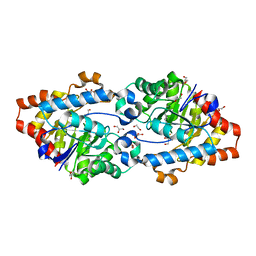 | | Structural basis for natural lactonase and promiscuous phosphotriesterase activities | | Descriptor: | 1,2-ETHANEDIOL, ARYLDIALKYLPHOSPHATASE, COBALT (II) ION, ... | | Authors: | Elias, M, Dupuy, J, Merone, L, Mandrich, L, Moniot, S, Lecomte, C, Rossi, M, Masson, P, Manco, G, Chabriere, E. | | Deposit date: | 2007-09-18 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.6 Å) | | Cite: | Structural Basis for Natural Lactonase and Promiscuous Phosphotriesterase Activities.
J.Mol.Biol., 379, 2008
|
|
2YJP
 
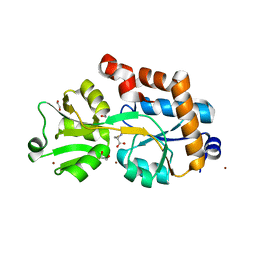 | | Crystal structure of the solute receptors for L-cysteine of Neisseria gonorrhoeae | | Descriptor: | 1,2-ETHANEDIOL, CYSTEINE, PUTATIVE ABC TRANSPORTER, ... | | Authors: | Bulut, H, Moniot, S, Scheffel, F, Gathmann, S, Licht, A, Saenger, W, Schneider, E. | | Deposit date: | 2011-05-23 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.26 Å) | | Cite: | Crystal Structures of Two Solute Receptors for L-Cystine and L-Cysteine, Respectively, of the Human Pathogen Neisseria Gonorrhoeae.
J.Mol.Biol., 415, 2012
|
|
2VC7
 
 | | Structural basis for natural lactonase and promiscuous phosphotriesterase activities | | Descriptor: | (4S)-4-(decanoylamino)-5-hydroxy-3,4-dihydro-2H-thiophenium, 1,2-ETHANEDIOL, ARYLDIALKYLPHOSPHATASE, ... | | Authors: | Elias, M, Dupuy, J, Merone, L, Mandrich, L, Moniot, S, Rochu, D, Lecomte, C, Rossi, M, Masson, P, Manco, G, Chabriere, E. | | Deposit date: | 2007-09-19 | | Release date: | 2008-04-15 | | Last modified: | 2023-12-13 | | Method: | X-RAY DIFFRACTION (2.05 Å) | | Cite: | Structural Basis for Natural Lactonase and Promiscuous Phosphotriesterase Activities.
J.Mol.Biol., 379, 2008
|
|
2YLN
 
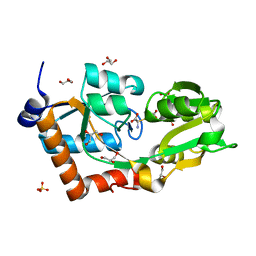 | | Crystal structure of the L-cystine solute receptor of Neisseria gonorrhoeae in the closed conformation | | Descriptor: | CYSTEINE, GLYCEROL, PUTATIVE ABC TRANSPORTER, ... | | Authors: | Bulut, H, Moniot, S, Scheffel, F, Gathmann, S, Licht, A, Saenger, W, Schneider, E. | | Deposit date: | 2011-06-03 | | Release date: | 2011-12-14 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.12 Å) | | Cite: | Crystal Structures of Two Solute Receptors for L-Cystine and L-Cysteine, Respectively, of the Human Pathogen Neisseria Gonorrhoeae.
J.Mol.Biol., 415, 2012
|
|
2YHY
 
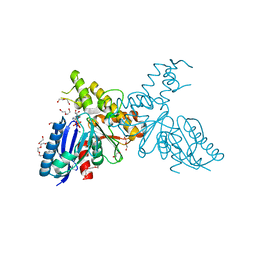 | | Structure of N-Acetylmannosamine kinase in complex with N- acetylmannosamine and ADP | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-alpha-D-mannopyranose, ADENOSINE-5'-DIPHOSPHATE, ... | | Authors: | Martinez, J, Nguyen, L.D, Tauberger, E, Hinderlich, S, Reutter, W, Fan, H, Saenger, W, Moniot, S. | | Deposit date: | 2011-05-10 | | Release date: | 2012-02-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (1.82 Å) | | Cite: | Crystal Structures of N-Acetylmannosamine Kinase Provide Insights Into Enzyme Specificity and Inhibition
J.Biol.Chem., 287, 2012
|
|
2YI1
 
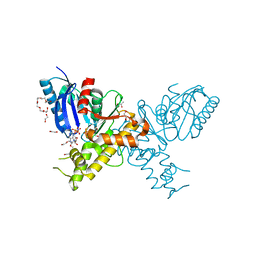 | | Crystal structure of N-Acetylmannosamine kinase in complex with N- acetyl mannosamine 6-phosphate and ADP. | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-6-O-phosphono-alpha-D-mannopyranose, 2-acetamido-2-deoxy-alpha-D-mannopyranose, ... | | Authors: | Martinez, J, Nguyen, L.D, Tauberger, E, Hinderlich, S, Reutter, W, Fan, H, Saenger, W, Moniot, S. | | Deposit date: | 2011-05-10 | | Release date: | 2012-02-29 | | Last modified: | 2023-12-20 | | Method: | X-RAY DIFFRACTION (2.15 Å) | | Cite: | Crystal Structures of N-Acetylmannosamine Kinase Provide Insights Into Enzyme Specificity and Inhibition
J.Biol.Chem., 287, 2012
|
|
2YHW
 
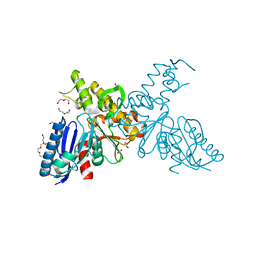 | | High-resolution crystal structures of N-Acetylmannosamine kinase: Insights about substrate specificity, activity and inhibitor modelling. | | Descriptor: | 1,2-ETHANEDIOL, 2-acetamido-2-deoxy-alpha-D-mannopyranose, ACETATE ION, ... | | Authors: | Martinez, J, Nguyen, L.D, Tauberger, E, Hinderlich, S, Zimmer, R, Tauberger, E, Reutter, W, Saenger, W, Fan, H, Moniot, S. | | Deposit date: | 2011-05-08 | | Release date: | 2012-02-29 | | Last modified: | 2023-11-15 | | Method: | X-RAY DIFFRACTION (1.64 Å) | | Cite: | Crystal Structures of N-Acetylmannosamine Kinase Provide Insights Into Enzyme Specificity and Inhibition
J.Biol.Chem., 287, 2012
|
|
3OG9
 
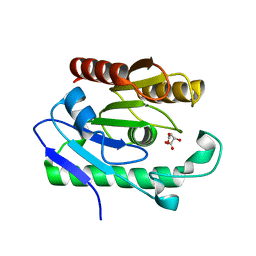 | | Structure of YahD with Malic acid | | Descriptor: | D-MALATE, protein yahD a copper inducible hydrolase | | Authors: | Martinez Font, J, Mancini, S, Tauberger, E, Moniot, S. | | Deposit date: | 2010-08-16 | | Release date: | 2010-12-15 | | Last modified: | 2023-09-06 | | Method: | X-RAY DIFFRACTION (1.88 Å) | | Cite: | Regulation and structure of YahD, a copper inducible alpha/beta hydrolase of Lactococcus lactis IL1403
Fems Microbiol.Lett., 314, 2011
|
|
4CLF
 
 | | Crystal structure of human soluble Adenylyl Cyclase (Apo form) | | Descriptor: | ACETATE ION, ADENYLATE CYCLASE TYPE 10, GLYCEROL | | Authors: | Kleinboelting, S, Moniot, S, Weyand, M, Steegborn, C. | | Deposit date: | 2014-01-14 | | Release date: | 2014-03-05 | | Last modified: | 2019-04-03 | | Method: | X-RAY DIFFRACTION (1.7 Å) | | Cite: | Crystal Structures of Human Soluble Adenylyl Cyclase Reveal Mechanisms of Catalysis and of its Activation Through Bicarbonate.
Proc.Natl.Acad.Sci.USA, 111, 2014
|
|
